Abstract
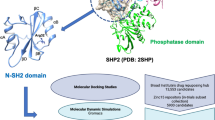
Discovery and development of a new drug to the market is a highly challenging and resource consuming process. Although, modern drug discovery technologies have enabled the rapid identification of lead compounds, translation of the lead compounds into successful clinical candidates remains a big challenge. In recent years, the availability of massive structural and biological data of diverse small molecules and macromolecules has helped the researchers to deep mine the multidimensional data with the help of artificial intelligence-based predictive tools to draw useful insights on the structural features of biological or therapeutic significance. The aim of this study was to utilize the available data on small molecule (SH2)-containing protein tyrosine phosphatase 2 (SHP2) inhibitors to build and develop machine learning (ML) models that can predict the SHP2 inhibitory potential of new compounds. The dataset contained 2739 unique small molecule SHP2 inhibitors obtained from the BindingDB, ChEMBL and recent literature. After curation of the data, the predictive models such as XGBoost, K nearest neighbours, neural networks were developed and validated through a tenfold cross-validation testing procedure. Out of the seven models developed, the XGBoost model showed an excellent performance with ROC AUC score of 0.96 and accuracy of 0.97 on the test data. Moreover, the Shapley Additive Explanations method was applied to assess a more in-depth understanding of the influence of variables on the model’s predictions. In summary, the XGBoost model developed in this study can be useful in the identification of novel SHP2 inhibitors and therefore, can accelerate the discovery of novel therapeutics for cancer therapy.
Graphical abstract



















Similar content being viewed by others
References
Feng GS, Shen R, Heng HH et al (1994) Receptor-binding, tyrosine phosphorylation and chromosome localization of the mouse SH2-containing phosphotyrosine phosphatase Syp. Oncogene 9:1545–1550
Feng GS, Hui CC, Pawson T (1993) Sh2-containing phosphotyrosine phosphatase as a target of protein-tyrosine kinases. Science 259:1607–1611. https://doi.org/10.1126/SCIENCE.8096088
Feng G-S, Pawson T (1994) Phosphotyrosine phosphatases with SH2 domains: regulators of signal transduction. Trends Genet 10:54–58. https://doi.org/10.1016/0168-9525(94)90149-X
Hof P, Pluskey S, Dhe-Paganon S et al (1998) Crystal structure of the tyrosine phosphatase SHP-2. Cell 92:441–450. https://doi.org/10.1016/S0092-8674(00)80938-1
Bennett AM, Tang TL, Sugimoto S et al (1994) Protein-tyrosine-phosphatase SHPTP2 couples platelet-derived growth factor receptor beta to Ras. Proc Natl Acad Sci USA 91:7335–7339. https://doi.org/10.1073/pnas.91.15.7335
Chan RJ, Feng G-S (2006) PTPN11 is the first identified proto-oncogene that encodes a tyrosine phosphatase. Blood 109:862–867. https://doi.org/10.1182/blood-2006-07-028829
Holgado-Madruga M, Emlet DR, Moscatello DK, Godwin AK, Wong AJ (1996) A Grb2-associated docking protein in EGF- and insulin-receptor signalling. Nature 379(6565):560–564. https://doi.org/10.1038/379560a0
Kouhara H, Hadari YR, Spivak-Kroizman T, Schilling J, Bar-Sagi D, Lax I, Schlessinger J (1997) A lipid-anchored Grb2-binding protein that links FGF-receptor activation to the Ras/MAPK signaling pathway. Cell 89(5):693–702
Xu D, Wang S, Yu WM et al (2010) A germline gain-of-function mutation in Ptpn11 (Shp-2) phosphatase induces myeloproliferative disease by aberrant activation of hematopoietic stem cells. Blood 116:3611–3621. https://doi.org/10.1182/BLOOD-2010-01-265652
Mohi MG, Neel BG (2007) The role of Shp2 (PTPN11) in cancer. Curr Opin Genet Dev 17(1):23–30
Bard-Chapeau EA, Li S, Ding J, Zhang SS, Zhu HH, Princen F, Fang DD, Han T, Bailly-Maitre B, Poli V, Varki NM, Wang H, Feng GS (2011) Ptpn11/Shp2 acts as a tumor suppressor in hepatocellular carcinogenesis. Cancer Cell 19(5):629–639. https://doi.org/10.1016/j.ccr.2011.03.023
Yang W, Wang J, Moore DC, Liang H, Dooner M, Wu Q, Terek R, Chen Q, Ehrlich MG, Quesenberry PJ, Neel BG (2013) Ptpn11 deletion in a novel progenitor causes metachondromatosis by inducing hedgehog signalling. Nature 499(7459):491–495. https://doi.org/10.1038/nature12396
Selvaraj C, Chandra I, Singh SK (2022) Artificial intelligence and machine learning approaches for drug design: challenges and opportunities for the pharmaceutical industries. Mol Divers 26:1893–1913. https://doi.org/10.1007/s11030-021-10326-z
Mao J, Akhtar J, Zhang X et al (2021) Comprehensive strategies of machine-learning-based quantitative structure-activity relationship models. iScience 24(9):103052. https://doi.org/10.1016/J.ISCI.2021.103052
Daina A, Michielin O, Zoete V (2017) SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci Rep 7:1–13. https://doi.org/10.1038/srep42717
Lavecchia A, Di Giovanni C (2013) Virtual screening strategies in drug discovery: a critical review. Curr Med Chem 20(23):2839–2860. https://doi.org/10.2174/09298673113209990001
Tripathi RKP, Ayyannan SR (2021) Emerging chemical scaffolds with potential SHP2 phosphatase inhibitory capabilities – A comprehensive review. Chem Biol Drug Des 97:721–773. https://doi.org/10.1111/cbdd.13807
Mitra R, Ayyannan SR (2021) Small-molecule inhibitors of Shp2 phosphatase as potential chemotherapeutic agents for glioblastoma: a minireview. ChemMedChem 16:777–787. https://doi.org/10.1002/cmdc.202000706
Yuan X, Bu H, Zhou J et al (2020) Recent advances of SHP2 inhibitors in cancer therapy: current development and clinical application. J Med Chem 63:11368–11396. https://doi.org/10.1021/acs.jmedchem.0c00249
Wang W-L, Chen X-Y, Gao Y et al (2017) Benzo[c][1,2,5]thiadiazole derivatives: a new class of potent Src homology-2 domain containing protein tyrosine phosphatase-2 (SHP2) inhibitors. Bioorg Med Chem Lett 27:5154–5157. https://doi.org/10.1016/j.bmcl.2017.10.059
Lawrence HR, Pireddu R, Chen L et al (2008) Inhibitors of Src homology-2 domain containing protein tyrosine phosphatase-2 (Shp2) based on oxindole scaffolds. J Med Chem 51:4948–4956. https://doi.org/10.1021/jm8002526
Satheeshkumar R, Zhu R, Feng B et al (2020) Synthesis and biological evaluation of heterocyclic bis-aryl amides as novel Src homology 2 domain containing protein tyrosine phosphatase-2 (SHP2) inhibitors. Bioorg Med Chem Lett 30:7170. https://doi.org/10.1016/j.bmcl.2020.127170
Geronikaki A, Eleftheriou P, Vicini P et al (2008) 2-Thiazolylimino/heteroarylimino-5-arylidene-4-thiazolidinones as new agents with SHP-2 inhibitory action. J Med Chem 51:5221–5228. https://doi.org/10.1021/jm8004306
Xie J, Si X, Gu S et al (2017) Allosteric inhibitors of SHP2 with therapeutic potential for cancer treatment. J Med Chem 60:10205–10219. https://doi.org/10.1021/acs.jmedchem.7b01520
Bagdanoff JT, Chen Z, Acker M et al (2019) Optimization of fused bicyclic allosteric SHP2 inhibitors. J Med Chem 62:1781–1792. https://doi.org/10.1021/acs.jmedchem.8b01725
Wu J, Li W, Zheng Z et al (2021) Design, synthesis, biological evaluation, common feature pharmacophore model and molecular dynamics simulation studies of ethyl 4-(phenoxymethyl)-2-phenylthiazole-5-carboxylate as Src homology-2 domain containing protein tyrosine phosphatase-2 (SHP2) inhibitors. J Biomol Struct Dyn 39:1174–1188. https://doi.org/10.1080/07391102.2020.1726817
Yu Z-H, Chen L, Wu L et al (2011) Small molecule inhibitors of SHP2 tyrosine phosphatase discovered by virtual screening. Bioorg Med Chem Lett 21:4238–4242. https://doi.org/10.1016/j.bmcl.2011.05.078
Zhang X, He Y, Liu S et al (2010) Salicylic acid based small molecule inhibitor for the oncogenic SRC homology-2 domain containing protein tyrosine phosphatase-2 (SHP2). J Med Chem 53:2482–2493. https://doi.org/10.1021/jm901645u
Landrum G (2016) RDKit: Open-Source Cheminformatics Software. https://www.rdkit.org/docs/. Accessed 1 Apr 2023
Li S, Ding Y, Chen M et al (2021) HDAC3i-finder: a machine learning-based computational tool to screen for HDAC3 inhibitors. Mol Inf 40:2000105. https://doi.org/10.1002/minf.202000105
Albadr MA, Tiun S, Ayob M, AL-Dhief F (2020) Genetic algorithm based on natural selection theory for optimization problems. Symmetry 12(11):1758. https://doi.org/10.3390/sym12111758
Leardi R, Boggia R, Terrile M (1992) Genetic algorithms as a strategy for feature selection. J Chemom 6:267–281. https://doi.org/10.1002/CEM.1180060506
Frazier, P.I. (2018). A Tutorial on Bayesian Optimization, arXiv:1807.02811. https://doi.org/10.48550/arXiv.1807.02811
Refaeilzadeh P, Tang L, Liu H (2009) Cross-Validation. Encyclopedia of Database Systems 532–538. https://doi.org/10.1007/978-0-387-39940-9_565
Furcht CM, Buonato JM, Skuli N et al (2014) Multivariate signaling regulation by SHP2 differentially controls proliferation and therapeutic response in glioma cells. J Cell Sci 127:3555–3567. https://doi.org/10.1242/JCS.150862/259148/AM/MULTIVARIATE-SIGNALING-REGULATION-BY-SHP2
Yang Z, Li Y, Yin F, Chan RJ (2008) Activating PTPN11 mutants promote hematopoietic progenitor cell-cycle progression and survival. Exp Hematol 36(10):1285–1296. https://doi.org/10.1016/j.exphem.2008.04.016
van der Maaten L, Hinton G (2008) Visualizing data using t-SNE. J Mach Learn Res 9:2579–2605
Nagai J, Imamura M, Sakagami H, Uesawa Y (2019) QSAR prediction model to search for compounds with selective cytotoxicity against oral cell cancer. Medicines 6:45. https://doi.org/10.3390/medicines6020045
Gasteiger J, Marsili M (1980) Iterative partial equalization of orbital electronegativity—a rapid access to atomic charges. Tetrahedron 36(22):3219–3228. https://doi.org/10.1016/0040-4020(80)80168-2
Wildman SA, Crippen GM (1999) Prediction of physicochemical parameters by atomic contributions. J Chem Inf Comput Sci 39(5):868–873. https://doi.org/10.1021/ci990307l
Riniker S, Landrum GA (2013) Similarity maps—a visualization strategy for molecular fingerprints and machine-learning methods. Journal of Cheminformatics 5:43. https://doi.org/10.1186/1758-2946-5-43
Pearlman RS, Smith KM (1998) Novel software tools for chemical diversity. Perspect Drug Discov Des 9:339–353. https://doi.org/10.1023/A:1027232610247
Acknowledgements
NA is thankful to the Indian Institute of Technology (Banaras Hindu University), Varanasi, for providing a teaching assistantship. The authors are thankful to Mr. R Mitra and Mr. J Lavudi for their support in the data collection. The authors also gratefully acknowledge the Centre for Computing and Information Services (CCIS), Indian Institute of Technology (Banaras Hindu University), Varanasi, for their outstanding computational assistance.
Author information
Authors and Affiliations
Contributions
Conceptualization, review and editing: Senthil Raja Ayyannan; Investigation and original draft preparation: Nilanjan Adhikari; Data analysis: Nilanjan Adhikari and Senthil Raja Ayyannan.
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Adhikari, N., Ayyannan, S.R. Development and validation of machine learning models for the prediction of SH-2 containing protein tyrosine phosphatase 2 inhibitors. Mol Divers (2023). https://doi.org/10.1007/s11030-023-10710-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11030-023-10710-x