Abstract
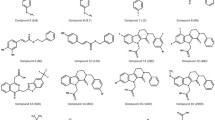
Vascular endothelial growth factor (VEGF) and its receptor tyrosine kinase VEGFR-2 or kinase insert domain receptor (KDR) have been identified as promising targets for novel anticancer agents. To achieve new potent inhibitors of KDR, we conducted molecular fragment replacement (MFR) studies for the understanding of 3D-QSAR modeling and the docking investigation of arylphthalazines and 2-((1H-Azol-1-yl)methyl)-N-arylbenzamides-based KDR inhibitors. Two favorable 3D-QSAR models (CoMFA with q 2, 0.671; r 2, 0.969; CoMSIA with q 2, 0.608; r 2, 0.936) have been developed to predict the biological activity of new compounds. The new molecular database generated by MFR was virtually screened using Glide (docking) and further evaluated with CoMFA prediction, protein–ligand interaction fingerprint (PLIF) and ADMET analysis. 44 N-(pyridin-4-ylmethyl)aniline derivatives as novel potential KDR inhibitors were finally obtained. In this paper, the work flow developed could be applied to de novo drug design and virtual screening potential KDR inhibitors, and use hit compounds to further optimize and design new potential KDR inhibitors.
Similar content being viewed by others
References
Ivy SP, Wick JY, Kaufman BM (2009) An overview of small-molecule inhibitors of VEGFR signaling. Nat Rev Clin Oncol 6: 569–579. doi:10.1038/nrclinonc.2009.130
Teicher BA (2011) Antiangiogenic agents and targets: a perspective. Biochem Pharmacol 81: 6–12. doi:10.1016/j.bcp.2010.09.023
Ferrara N, Kerbel RS (2005) Angiogenesis as a therapeutic target. Nature 438: 967–974. doi:10.1038/nature04483
Ferrara N, Hillan KJ, Gerber HP, Novotny W (2004) Discovery and development of bevacizumab, an anti-VEGF antibody for treating cancer. Nat Rev Drug Discov 3: 391–400. doi:10.1038/nrd1381
Ahmad T, Eisen T (2004) Kinase inhibition with BAY 43-9006 in renal cell carcinoma. Clin Cancer Res 10: 6388S–6392S. doi:10.1158/1078-0432.ccr-040028
Sakamoto KM (2004) Su-11248 Sugen. Curr Opin Investig Drugs 5: 1329–1339
Cabebe E, Wakelee H (2006) Sunitinib: a newly approved small-molecule inhibitor of angiogenesis. Drugs Today (Barc) 42: 387–398. doi:10.1358/dot.2006.42.6.985633
Podar K, Catley LP, Tai YT, Shringarpure R, Carvalho P, Hayashi T, Burger R, Schlossman RL, Richardson PG, Pandite LN, Kumar R, Hideshima T, Chauhan D, Anderson KC (2004) GW654652, the pan-inhibitor of VEGF receptors, blocks the growth and migration of multiple myeloma cells in the bone marrow microenvironment. Blood 103: 3474–3479. doi:10.1182/blood-2003-10-3527
Tyagi P (2005) Vatalanib (PTK787/ZK 222584) in combination with FOLFOX4 versus FOLFOX4 alone as first-line treatment for colorectal cancer: preliminary results from the CONFIRM-1 trial. Clin Colorectal Cancer 5: 24–26. doi:10.1016/S1533-0028(11)70162-1
ISCHEMIA CL (2012) Drug approved to treat advanced disease. J Vasc Surg 55: 371–380
Morabito A, De Maio E, Di Maio M, Normanno N, Perrone F (2006) Tyrosine kinase inhibitors of vascular endothelial growth factor receptors in clinical trials: current status and future directions. Oncologist 11: 753–764. doi:10.1634/theoncologist.11-7-753
Goodman VL, Rock EP, Dagher R, Ramchandani RP, Abraham S, Gobburu JV, Booth BP, Verbois SL, Morse DE, Liang CY, Chidambaram N, Jiang JX, Tang S, Mahjoob K, Justice R, Pazdur R (2007) Approval summary: sunitinib for the treatment of imatinib refractory or intolerant gastrointestinal stromal tumors and advanced renal cell carcinoma. Clin Cancer Res 13: 1367–1373. doi:10.1158/1078-0432.ccr-06-2328
Wood JM, Bold G, Buchdunger E, Cozens R, Ferrari S, Frei J, Hofmann F, Mestan J, Mett H, O’Reilly T, Persohn E, Rosel J, Schnell C, Stover D, Theuer A, Towbin H, Wenger F, Woods-Cook K, Menrad A, Siemeister G, Schirner M, Thierauch KH, Schneider MR, Drevs J, Martiny-Baron G, Totzke F (2000) PTK787/ZK 222584, a novel and potent inhibitor of vascular endothelial growth factor receptor tyrosine kinases, impairs vascular endothelial growth factor-induced responses and tumor growth after oral administration. Cancer Res 60: 2178–2189
Li R, Stafford JA (2009) Kinase inhibitor drugs. Wiley, Hoboken
Duncton MA, Piatnitski Chekler EL, Katoch-Rouse R, Sherman D, Wong WC, Smith LM, Kawakami JK, Kiselyov AS, Milligan DL, Balagtas C, Hadari YR, Wang Y, Patel SN, Rolster RL, Tonra JR, Surguladze D, Mitelman S, Kussie P, Bohlen P, Doody JF (2009) Arylphthalazines as potent, and orally bioavailable inhibitors of VEGFR-2. Bioorg Med Chem 17: 731–740. doi:10.1016/j.bmc.2008.11.049
Kiselyov AS, Semenova M, Semenov VV, Piatnitski E (2006) 2-((1H-Azol-1-yl)methyl)-N-arylbenzamides: novel dual inhibitors of VEGFR-1/2 kinases. Bioorg Med Chem Lett 16: 1726–1730. doi:10.1016/j.bmcl.2005.11.105
Selassie C (2003) History of quantitative structure-activity relationships. In: Burger’s medicinal chemistry and drug discovery, 6th edn. Wiley, New York
Cramer RD, Patterson DE, Bunce JD (1988) Comparative molecular field analysis (CoMFA). 1. Effect of shape on binding of steroids to carrier proteins. J Am Chem Soc 110: 5959–5967. doi:10.1021/ja00226a005
Cramer RD, Bunce JD, Patterson DE, Frank IE (1988) Crossvalidation, bootstrapping, and partial least squares compared with multiple regression in conventional QSAR studies. Quant Struct-Act Relat 7: 18–25. doi:10.1002/qsar.19880070105
Klebe G, Abraham U, Mietzner T (1994) Molecular similarity indices in a comparative analysis (CoMSIA) of drug molecules to correlate and predict their biological activity. J Med Chem 37: 4130–4146. doi:10.1021/jm00050a010
Neaz M, Pasha F, Muddassar M, Lee SH, Sim T, Hah JM, Cho SJ (2009) Pharmacophore based 3D-QSAR study of VEGFR-2 inhibitors. Med Chem Res 18: 127–142. doi:10.1007/s00044-008-9113-4
Lu X, Chen Y, You Q (2009) Pharmacophore guided 3D-QSAR CoMFA analysis of amino substituted nitrogen heterocycle ureas as KDR inhibitors. QSAR Comb Sci 28: 1524–1536. doi:10.1002/qsar.200960032
Zeng H, Zhang H (2010) Combined 3D-QSAR modeling and molecular docking study on 1,4-dihydroindeno[1,2-c]pyrazoles as VEGFR-2 kinase inhibitors. J Mol Graph Model 29: 54–71. doi:10.1016/j.jmgm.2010.04.004
Wu X, Wu S, Chen WH (2012) Molecular docking and 3D-QSAR study on 4-(1H-indazol-4-yl) phenylamino and aminopyrazolopyridine urea derivatives as kinase insert domain receptor (KDR) inhibitors. J Mol Model 18: 1207–1218. doi:10.1007/s00894-011-1146-9
Munoz C, Adasme F, Alzate-Morales JH, Vergara-Jaque A, Kniess T, Caballero J (2012) Study of differences in the VEGFR2 inhibitory activities between semaxanib and SU5205 using 3D-QSAR, docking, and molecular dynamics simulations. J Mol Graph Model 32: 39–48. doi:10.1016/j.jmgm.2011.10.005
Golbraikh A, Tropsha A (2002) Predictive QSAR modeling based on diversity sampling of experimental datasets for the training and test set selection. J Comput Aided Mol Des 16: 357–369. doi:10.1023/A:1020869118689
St. Louis M (1999) Sybyl version6.9. Tripos Associates, St. Louis
Myint KZ, Xie XQ (2010) Recent advances in fragment-based QSAR and multi-dimensional QSAR methods. Int J Mol Sci 11: 3846–3866. doi:10.3390/ijms11103846
Kontoyianni M, McClellan LM, Sokol GS (2004) Evaluation of docking performance: comparative data on docking algorithms. J Med Chem 47: 558–565. doi:10.1021/jm0302997
Kaminski GA, Friesner RA, Tirado-Rives J, Jorgensen WL (2001) Evaluation and reparametrization of the OPLS-AA force field for proteins via comparison with accurate quantum chemical calculations on peptides. J Phys Chem B 105: 6474–6487. doi:10.1021/jp003919d
Geladi P (1988) Notes on the history and nature of partial least squares (PLS) modelling. J Chemometr 2: 231–246. doi:10.1002/cem.1180020403
Geladi P, Kowalski BR (1986) Partial least-squares regression: a tutorial. Anal chim Acta 185: 1–17. doi:10.1016/0003-2670(86)80028-9
Bergmann R, Linusson A, Zamora I (2007) SHOP: scaffold HOPping by GRID-based similarity searches. J Med Chem 50: 2708–2717. doi:10.1021/jm061259g
Labute P (2009) Protonate3D: assignment of ionization states and hydrogen coordinates to macromolecular structures. Proteins 75: 187–205. doi:10.1002/prot.22234
Wildman SA, Crippen GM (1999) Prediction of physicochemical parameters by atomic contributions. J Chem Inf Comput Sci 39: 868–873. doi:10.1021/ci990307l
Ertl P, Rohde B, Selzer P (2000) Fast calculation of molecular polar surface area as a sum of fragment-based contributions and its application to the prediction of drug transport properties. J Med Chem 43: 3714–3717. doi:10.1021/jm000942e
Lipkus AH (1999) A proof of the triangle inequality for the Tanimoto distance. J Math Chem 26: 263–265. doi:10.1023/A:1019154432472
Mpamhanga CP, Chen B, McLay IM, Willett P (2006) Knowledge-based interaction fingerprint scoring: a simple method for improving the effectiveness of fast scoring functions. J Chem Inf Model 46: 686–698. doi:10.1021/ci050420d
Marcou G, Rognan D (2007) Optimizing fragment and scaffold docking by use of molecular interaction fingerprints. J Chem Inf Model 47: 195–207. doi:10.1021/ci600342e
Dogra SK (2007) Tanimoto_Coefficient. QSAR World:1–4
Author information
Authors and Affiliations
Corresponding authors
Electronic Supplementary Material
The Below is the Electronic Supplementary Material.
Rights and permissions
About this article
Cite this article
Zhang, Y., Liu, H., Jiao, Y. et al. De novo design of N-(pyridin-4-ylmethyl)aniline derivatives as KDR inhibitors: 3D-QSAR, molecular fragment replacement, protein-ligand interaction fingerprint, and ADMET prediction. Mol Divers 16, 787–802 (2012). https://doi.org/10.1007/s11030-012-9405-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11030-012-9405-y