Abstract
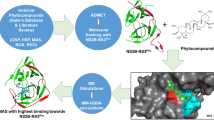
Viruses are microscopic biological entities that can quickly invade and multiply in a living organism. Each year, over 36,000 people die and nearly 400 million are infected with the dengue virus (DENV). Despite dengue being an endemic disease, no targeted and effective antiviral peptide resource is available against the dengue species. Antiviral peptides (AVPs) have shown tremendous ability to fight against different viruses. Accelerating antiviral drug discovery is crucial, particularly for RNA viruses. DDX3X, a vital cell component, supports viral translation and interacts with TRPV4, regulating viral RNA metabolism and infectivity. Its diverse signaling pathway makes it a potential therapeutic target. Our study focuses on inhibiting viral RNA translation by blocking the activity of the target gene and the TRPV4-mediated Ca2+ cation channel. Six major proteins from camel milk were first extracted and split with the enzyme pepsin. The antiviral properties were then analyzed using online bioinformatics programs, including AVPpred, Meta-iAVP, AMPfun, and ENNAVIA. The stability of the complex was assessed using MD simulation, MM/GBSA, and principal component analysis. Cytotoxicity evaluations were conducted using COPid and ToxinPred. The top ten AVPs, determined by optimal scores, were selected and saved for docking studies with the GalaxyPepDock tools. Bioinformatics analyses revealed that the peptides had very short hydrogen bond distances (1.8 to 3.6 Å) near the active site of the target protein. Approximately 76% of the peptide residues were 5–11 amino acids long. Additionally, the identified peptide candidates exhibited desirable properties for potential therapeutic agents, including a net positive charge, moderate toxicity, hydrophilicity, and selectivity. In conclusion, this computational study provides promising insights for discovering peptide-based therapeutic agents against DENV.








Similar content being viewed by others
References
Altayb HN (2022) Fludarabine, a potential DNA-dependent RNA polymerase inhibitor, as a prospective drug against Monkeypox virus: a computational approach. Pharmaceuticals (Basel, Switzerland) 15(9). https://doi.org/10.3390/PH15091129
Ariumi Y, Kuroki M, Abe K, Dansako H, Ikeda M, Wakita T, Kato N (2007) DDX3 DEAD-box RNA helicase is required for hepatitis C virus RNA replication. J Virol 81(24):13922–13926. https://doi.org/10.1128/JVI.01517-07
Awal MA, Nur SM, Al Khalaf AK, Rehan M, Ahmad A, Hosawi SBI, Choudhry H, Khan MI (2022) Structural-guided identification of small molecule inhibitor of UHRF1 methyltransferase activity. Front Genet 13:1750. https://doi.org/10.3389/FGENE.2022.928884/BIBTEX
Ayyash M, Al-Dhaheri AS, Al Mahadin S, Kizhakkayil J, Abushelaibi A (2018) In vitro investigation of anticancer, antihypertensive, antidiabetic, and antioxidant activities of camel milk fermented with camel milk probiotic: a comparative study with fermented bovine milk. J Dairy Sci 101(2):900–911. https://doi.org/10.3168/JDS.2017-13400
Bakan A, Meireles LM, Bahar I (2011) ProDy: protein dynamics inferred from theory and experiments. Bioinformatics 27(11):1575–1577. https://doi.org/10.1093/BIOINFORMATICS/BTR168
Behrouz S, Saadat S, Memarzia A, Sarir H, Folkerts G, Boskabady MH (2022) The antioxidant, anti-inflammatory and immunomodulatory effects of camel milk. Front Immunol 13. https://doi.org/10.3389/fimmu.2022.855342
Bhatt S, Gething PW, Brady OJ, Messina JP, Farlow AW, Moyes CL, Drake JM, Brownstein JS, Hoen AG, Sankoh O, Myers MF, George DB, Jaenisch T, William Wint GR, Simmons CP, Scott TW, Farrar JJ, Hay SI (2013) The global distribution and burden of dengue. Nature 496(7446):504–507. https://doi.org/10.1038/NATURE12060
Bolinger C, Boris-Lawrie K (2009) Mechanisms employed by retroviruses to exploit host factors for translational control of a complicated proteome. Retrovirology 6. https://doi.org/10.1186/1742-4690-6-8
Bulet P, Stöcklin R, Menin L (2004) Anti-microbial peptides: from invertebrates to vertebrates. Immunol Rev 198:169–184. https://doi.org/10.1111/J.0105-2896.2004.0124.X
Chang KY, Yang JR (2013) Analysis and prediction of highly effective antiviral peptides based on random forests. PLoS One 8(8):e70166. https://doi.org/10.1371/JOURNAL.PONE.0070166
Chen H, Mollstedt O, Tsai M-H, Kreider R (2014) Potential clinical applications of multi-functional milk proteins and peptides in cancer management. Curr Med Chem 21(21):2424–2437. https://doi.org/10.2174/0929867321666140205135739
Chianese A, Zannella C, Monti A, De Filippis A, Doti N, Franci G, Galdiero M (2022) The broad-spectrum antiviral potential of the amphibian peptide AR-23. Int J Mol Sci 23(2). https://doi.org/10.3390/IJMS23020883
Chung CR, Kuo TR, Wu LC, Lee TY, Horng JT (2020) Characterization and identification of antimicrobial peptides with different functional activities. Brief Bioinform 21(3):1098–1114. https://doi.org/10.1093/BIB/BBZ043
Dimitrov I, Bangov I, Flower DR, Doytchinova I (2014a) AllerTOP v.2--a server for in silico prediction of allergens. J Mol Model 20(6). https://doi.org/10.1007/S00894-014-2278-5
Dimitrov I, Naneva L, Doytchinova I, Bangov I (2014b) AllergenFP: allergenicity prediction by descriptor fingerprints. Bioinformatics 30(6):846–851. https://doi.org/10.1093/BIOINFORMATICS/BTT619
Doñate-Macián P, Jungfleisch J, Pérez-Vilaró G, Rubio-Moscardo F, Perálvarez-Marín A, Diez J, Valverde MA (2018) The TRPV4 channel links calcium influx to DDX3X activity and viral infectivity. Nat Commun 9(1):1–13. https://doi.org/10.1038/s41467-018-04776-7
El Agamy ESI, Ruppanner R, Ismail A, Champagne CP, Assaf R (1992) Antibacterial and antiviral activity of camel milk protective proteins. J Dairy Res 59(2):169–175. https://doi.org/10.1017/S0022029900030417
Fu T, Zheng G, Tu G, Yang F, Chen Y, Yao X, Li X, Xue W, Zhu F (2018) Exploring the binding mechanism of metabotropic glutamate receptor 5 negative allosteric modulators in clinical trials by molecular dynamics simulations. ACS Chem Neurosci 9(6):1492–1502. https://doi.org/10.1021/ACSCHEMNEURO.8B00059
Gasteiger E, Hoogland C, Gattiker A, Duvaud S, Wilkins MR, Appel RD, Bairoch A (2005) Protein identification and analysis tools on the ExPASy server. The Proteomics Protocols Handbook, pp 571–607. https://doi.org/10.1385/1-59259-890-0:571
Gould E, Solomon T (2008) Pathogenic flaviviruses. Lancet (London, England) 371(9611):500–509. https://doi.org/10.1016/S0140-6736(08)60238-X
Grant BJ, Rodrigues APC, ElSawy KM, McCammon JA, Caves LSD (2006) Bio3d: an R package for the comparative analysis of protein structures. Bioinformatics 22(21):2695–2696. https://doi.org/10.1093/BIOINFORMATICS/BTL461
Hancock REW (1997) Peptide antibiotics. Lancet (London, England) 349(9049):418–422. https://doi.org/10.1016/S0140-6736(97)80051-7
Heller S, O’Neil RG (2007) Molecular mechanisms of TRPV4 Gating. 113–124. https://doi.org/10.1201/9781420005844.ch8
Hernández-Díaz T, Valiente-Echeverría F, Soto-Rifo R (2021) RNA helicase DDX3: a double-edged sword for viral replication and immune signaling. Microorganisms 9(6). https://doi.org/10.3390/MICROORGANISMS9061206
Ibrahim HR, Isono H, Miyata T (2018) Potential antioxidant bioactive peptides from camel milk proteins. Anim Nutr 4(3):273. https://doi.org/10.1016/J.ANINU.2018.05.004
Islam MR, Awal MA, Khames A, Abourehab MAS, Samad A, Hassan WMI, Alam R, Osman OI, Nur SM, Molla MHR, Abdulrahman AO, Rajia S, Ahammad F, Hasan MN, Qadri I, Kim B (2022) Computational identification of Druggable bioactive compounds from Catharanthus roseus and Avicennia marina against colorectal Cancer by targeting thymidylate synthase. Molecules 27(7):2089. https://doi.org/10.3390/MOLECULES27072089
Islam MR, Osman OI, Hassan WMI (2023) Identifying novel therapeutic inhibitors to target FMS-like tyrosine kinase-3 (FLT3) against acute myeloid leukemia: a molecular docking, molecular dynamics, and DFT study. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2023.2192798/SUPPL_FILE/TBSD_A_2192798_SM6396.DOCX
Kappeler SR, Ackermann M, Farah Z, Puhan Z (1999) Sequence analysis of camel (Camelus dromedarius) lactoferrin. Int Dairy J 9(7):481–486. https://doi.org/10.1016/S0958-6946(99)00117-X
Korashy HM, Maayah ZH, Abd-Allah AR, El-Kadi AOS, Alhaider AA (2012) Camel milk triggers apoptotic signaling pathways in human hepatoma HepG2 and breast cancer MCF7 cell lines through transcriptional mechanism. J Biomed Biotechnol 2012. https://doi.org/10.1155/2012/593195
Kumar R, Singh N, Abdin MZ, Patel AH, Medigeshi GR (2018) Dengue virus capsid interacts with DDX3X-A potential mechanism for suppression of antiviral functions in dengue infection. Front Cell Infect Microbiol 7(JAN):542. https://doi.org/10.3389/FCIMB.2017.00542/FULL
Kumar N, Sood D, Van Der Spek PJ, Sharma HS, Chandra R (2020) Molecular binding mechanism and pharmacology comparative analysis of noscapine for repurposing against SARS-CoV-2 protease. J Proteome Res 19(11):4678–4689. https://doi.org/10.1021/ACS.JPROTEOME.0C00367
Mata J, Marguerat S, Bähler J (2005) Post-transcriptional control of gene expression: a genome-wide perspective. Trends Biochem Sci 30(9):506–514. https://doi.org/10.1016/J.TIBS.2005.07.005
Mirmiran P, Ejtahed HS, Angoorani P, Eslami F, Azizi F (2017) Camel milk has beneficial effects on diabetes mellitus: a systematic review. Int J Endocrinol Metab 15(2). https://doi.org/10.5812/IJEM.42150
Mookherjee N, Hancock REW (2007) Cationic host defence peptides: innate immune regulatory peptides as a novel approach for treating infections. Cell Mol Life Sci: CMLS 64(7–8):922–933. https://doi.org/10.1007/S00018-007-6475-6
Otvos L, Bokonyi K, Varga I, Ertl HCJ, Hoffmann R, Bulet P, Otvos BI, Wade JD, Mcmanus AM, Craik DJ (2008) Insect peptides with improved protease-resistance protect mice against bacterial infection. Protein Sci 9(4):742–749. https://doi.org/10.1110/PS.9.4.742
Pascut D, Bedogni G, Tiribelli C (2015) Silencing efficacy prediction: a retrospective study on target mRNA features. Biosci Rep 35(2). https://doi.org/10.1042/BSR20140147
Puri B, Nelson WM, Henchal EA, Hoke CH, Eckels KH, Dubois DR, Porter KR, Hayes CG (1997) Molecular analysis of dengue virus attenuation after serial passage in primary dog kidney cells. J Gen Virol 78(9):2287–2291. https://doi.org/10.1099/0022-1317-78-9-2287
Qureshi A, Kaur G, Kumar M (2017) AVCpred: an integrated web server for prediction and design of antiviral compounds. Chem Biol Drug Des 89(1):74. https://doi.org/10.1111/CBDD.12834
Rajan S, Schremmer C, Weber J, Alt P, Geiger F, Dietrich A (2021) Ca2+ signaling by TRPV4 channels in respiratory function and disease. Cells 10(4). https://doi.org/10.3390/CELLS10040822
Recio C, Maione F, Iqbal AJ, Mascolo N, De Feo V (2017) The Potential Therapeutic Application of Peptides and Peptidomimetics in Cardiovascular Disease. Front Pharmacol 7(JAN). https://doi.org/10.3389/FPHAR.2016.00526
Rodenhuis-Zybert IA, Wilschut J, Smit JM (2010) Dengue virus life cycle: viral and host factors modulating infectivity. Cell Mol Life Sci 67(16):2773–2786. https://doi.org/10.1007/S00018-010-0357-Z
Schaduangrat N, Nantasenamat C, Prachayasittikul V, Shoombuatong W (2019) Meta-iAVP: a sequence-based Meta-predictor for improving the prediction of antiviral peptides using effective feature representation. Int J Mol Sci 20(22). https://doi.org/10.3390/IJMS20225743
Shai Y (2002) Mode of action of membrane active antimicrobial peptides. Biopolymers - Peptide Sci Sect 66(4):236–248. https://doi.org/10.1002/bip.10260
Shin W-H, Lee GR, Heo L, Lee H, Seok C (2014) Prediction of protein structure and interaction by GALAXY protein modeling programs. BioDesign 2(1):1–11 https://www.bdjn.org/journal/view.html?spage=1&volume=2&number=1&vmd=A. Accessed 05 Oct 2023
Silva AF, Bastos EL, Torres MDT, Costa-Da-Silva AL, Ioshino RS, Capurro ML, Alves FL, Miranda A, De Freitas Fischer Vieira R, Oliveira VX (2014) Antiplasmodial activity study of angiotensin II via ala scan analogs. J Pept Sci 20(8):640–648. https://doi.org/10.1002/PSC.2641
Soliman MM, Hassan MY, Abdel-Hafiz Mostafa S, Abdel-Maksoud Ali H, Saleh OM (2015) Protective effects of camel milk against pathogenicity induced by Escherichia coli and Staphylococcus aureus in Wistar rats. Mol Med Rep 12(6):8306–8312. https://doi.org/10.3892/MMR.2015.4486
Sterri SH, Fonnum F (2009) Role of carboxylesterases in therapeutic intervention of nerve gas poisoning. Handbook Toxicol Chem Warfare Agents:1033–1040. https://doi.org/10.1016/B978-012374484-5.00068-7
Thakur N, Qureshi A, Kumar M (2012) AVPpred: collection and prediction of highly effective antiviral peptides. Nucleic Acids Res 40(W1):W199–W204. https://doi.org/10.1093/NAR/GKS450
Timmons PB, Hewage CM (2021) ENNAVIA is a novel method which employs neural networks for antiviral and anti-coronavirus activity prediction for therapeutic peptides. Brief Bioinform 22(6):1–17. https://doi.org/10.1093/BIB/BBAB258
Valiente-Echeverría F, Hermoso MA, Soto-Rifo R (2015) RNA helicase DDX3: at the crossroad of viral replication and antiviral immunity. Rev Med Virol 25(5):286–299. https://doi.org/10.1002/RMV.1845
Wang CK, Shih LY, Chang KY (2017) Large-scale analysis of antimicrobial activities in relation to Amphipathicity and charge reveals novel characterization of antimicrobial peptides. Molecules 22(11):2037. https://doi.org/10.3390/MOLECULES22112037
Webb-Robertson BJM (2009) Support vector machines for improved peptide identification from tandem mass spectrometry database search. Methods Mol Biol (Clifton, NJ) 492:453–460. https://doi.org/10.1007/978-1-59745-493-3_28
World Health Orgnization (2018) Dengue and severe dengue. www.who.int/en/news-room/fact-sheets/detail/dengueand-severe-dengue. Accessed 23 Nov 2023
Wu W, Bai Z, Zhou H, Tu Z, Fang M, Tang B, Liiu J, Liu L, Liu J, Chen W (2011) Molecular epidemiology of dengue viruses in southern China from 1978 to 2006. Virol J 8. https://doi.org/10.1186/1743-422X-8-322
Yedavalli VSRK, Neuveut C, Chi YH, Kleiman L, Jeang KT (2004) Requirement of DDX3 DEAD box RNA helicase for HIV-1 Rev-RRE export function. Cell 119(3):381–392. https://doi.org/10.1016/j.cell.2004.09.029
Yin LM, Edwards MA, Li J, Yip CM, Deber CM (2012) Roles of hydrophobicity and charge distribution of cationic antimicrobial peptides in peptide-membrane interactions. J Biol Chem 287(10):7738. https://doi.org/10.1074/JBC.M111.303602
Zannella C, Chianese A, Palomba L, Marcocci ME, Bellavita R, Merlino F, Grieco P, Folliero V, De Filippis A, Mangoni M, Nencioni L, Franci G, Galdiero M (2022) Broad-spectrum antiviral activity of the amphibian antimicrobial peptide Temporin L and its analogs. Int J Mol Sci 23(4):2060. https://doi.org/10.3390/IJMS23042060/S1
Zhou Y, Frey TK, Yang JJ (2009) Viral calciomics: interplays between Ca2+ and virus. Cell Calcium 46(1):1–17. https://doi.org/10.1016/J.CECA.2009.05.005
Acknowledgments
This project was funded by the Deanship of Scientific Research (D.S.R.), King Abdulaziz University, Jeddah, Saudi Arabia, under Grant no. (G: 441-130-1443). The authors, therefore, acknowledge with thanks to D.S.R. and the Advanced Biological Invention Centre, Bioinventics (https://www.bioinventics.com/) technical and financial support.
Funding
This project was funded by the Deanship of Scientific Research (D.S.R.), King Abdulaziz University, Jeddah, Saudi Arabia, under Grant no. (G: 441–130-1443). The authors, therefore, acknowledge with thanks to D.S.R. technical and financial support.
Author information
Authors and Affiliations
Contributions
A.H.A. and H.N.A.: conceptualization, supervision, methodology, investigation, and funding acquisition; M.R.I: writing review and editing, software, bioinformatics analysis, data curation, validation, resources, visualization, and original draft preparation. R.M.A: writing review and editing, software, bioinformatics analysis. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(DOCX 293 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Asseri, A.H., Islam, M.R., Alghamdi, R.M. et al. Identification of natural antimicrobial peptides mimetic to inhibit Ca2+ influx DDX3X activity for blocking dengue viral infectivity. J Bioenerg Biomembr 56, 125–139 (2024). https://doi.org/10.1007/s10863-023-09996-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10863-023-09996-1