Abstract
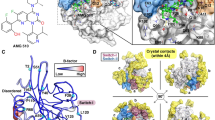
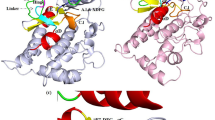
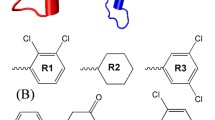
The mutant KRAS was considered as an “undruggable” target for decades, especially KRASG12D. It is a great challenge to develop the inhibitors for KRASG12D which lacks the thiol group for covalently binding ligands. The discovery of MRTX1133 solved the dilemma. Interestingly, MRTX1133 can bind to both the inactive and active states of KRASG12D. The binding mechanism of MRTX1133 with KRASG12D, especially how MRTX1133 could bind the active state KRASG12D without triggering the active function of KRASG12D, has not been fully understood. Here, we used a combination of all-atom molecular dynamics simulations and Markov state model (MSM) to understand the inhibition mechanism of MRTX1133 and its analogs. The stationary probabilities derived from MSM show that MRTX1133 and its analogs can stabilize the inactive or active states of KRASG12D into different conformations. More remarkably, by scrutinizing the conformational differences, MRTX1133 and its analogs were hydrogen bonded to Gly60 to stabilize the switch II region and left switch I region in a dynamically inactive conformation, thus achieving an inhibitory effect. Our simulation and analysis provide detailed inhibition mechanism of KRASG12D induced by MRTX1133 and its analogs. This study will provide guidance for future design of novel small molecule inhibitors of KRASG12D.





Similar content being viewed by others
Data availability
Data are freely available upon request of the corresponding author.
Abbreviations
- KRAS:
-
Kirsten rat sarcoma virus oncogene
- GDP:
-
Guanosine diphosphate
- GTP:
-
Guanosine triphosphate
- GppNHp:
-
Guanosine 5′-(β-γ-imido) triphosphate
- RMSD:
-
Root mean squared deviation
- MD:
-
Molecular dynamics
- TIP3P:
-
Transferable interatomic potential with three points
- MSM:
-
Markov state model
References
Bos JL (1989) Ras oncogenes in human cancer: a review. Can Res 49(17):4682–4689
Simanshu DK, Nissley DV, McCormick F (2017) RAS proteins and their regulators in human disease. Cell 170(1):17–33. https://doi.org/10.1016/j.cell.2017.06.009
Mo SP, Coulson JM, Prior IA (2018) RAS variant signalling. Biochem Soc Trans 46(5):1325. https://doi.org/10.1042/BST20180173
Pantsar T (2020) The current understanding of KRAS protein structure and dynamics. Comput Struct Biotechnol J 18:189–198. https://doi.org/10.1016/j.csbj.2019.12.004
Prior IA, Lewis PD, Mattos C (2012) A comprehensive survey of Ras mutations in cancer. Can Res 72(10):2457–2467. https://doi.org/10.1158/0008-5472.CAN-11-2612
Bourne HR, Sanders DA, McCormick F (1991) The GTPase superfamily: conserved structure and molecular mechanism. Nature 349(6305):117–127. https://doi.org/10.1038/349117a0
Wittinghofer A, Scheffzek K, Ahmadian MR (1997) The interaction of Ras with GTPase-activating proteins. FEBS Lett 410(1):63–67. https://doi.org/10.1016/S0014-5793(97)00321-9
Gideon P, John J, Frech M et al (1992) Mutational and kinetic analyses of the GTPase-activating protein (GAP)-p21 interaction: the C-terminal domain of GAP is not sufficient for full activity. Mol Cell Biol 12(5):2050–2056. https://doi.org/10.1128/mcb.12.5.2050-2056.1992
Eccleston JF, Moore K, Morgan L et al (1993) Kinetics of interaction between normal and proline 12 Ras and the GTPase-activating proteins, p120-GAP and neurofibromin: the significance of the intrinsic GTPase rate in determining the transforming ability of ras. J Biol Chem 268(36):27012–27019. https://doi.org/10.1016/S0021-9258(19)74211-2
Vetter IR, Wittinghofer A (2001) The guanine nucleotide-binding switch in three dimensions. Science 294(5545):1299–1304. https://doi.org/10.1126/science.1062023
Pacold ME, Suire S, Perisic O et al (2000) Crystal structure and functional analysis of Ras binding to its effector phosphoinositide 3-kinase γ. Cell 103(6):931–944. https://doi.org/10.1016/S0092-8674(00)00196-3
Nassar N, Horn G, Herrmann CA et al (1995) The 2.2 Å crystal structure of the Ras-binding domain of the serine/threonine kinase c-Raf1 in complex with RaplA and a GTP analogue. Nature 375(6532):554–560. https://doi.org/10.1038/375554a0
Tuttle RL, Gill NS, Pugh W et al (2001) Regulation of pancreatic β-cell growth and survival by the serine/threonine protein kinase Akt1/PKBα. Nat Med 7(10):1133–1137. https://doi.org/10.1038/nm1001-1133
Drosten M, Dhawahir A, Sum EY et al (2010) Genetic analysis of Ras signalling pathways in cell proliferation, migration and survival. EMBO J 29(6):1091–1104. https://doi.org/10.1038/emboj.2010.7
Crespo P, Leon J (2000) Ras proteins in the control of the cell cycle and cell differentiation. Cell Mol Life Sci CMLS 57(11):1613–1636. https://doi.org/10.1007/PL00000645
Marshall C (1995) Specificity of receptor tyrosine kinase signaling: transient versus sustained extracellular signal-regulated kinase activation. Cell 80(2):179–185. https://doi.org/10.1016/0092-8674(95)90401-8
Cully M, You H, Levine AJ, Mak TW (2006) Beyond PTEN mutations: the PI3K pathway as an integrator of multiple inputs during tumorigenesis. Nat Rev Cancer 6(3):184–192. https://doi.org/10.1038/nrc1819
Kessler D, Gmachl M, Mantoulidis A et al (2019) Drugging an undruggable pocket on KRAS. Proc Natl Acad Sci 116(32):15823–15829. https://doi.org/10.1073/pnas.1904529116
Zhu K, Li C, Wu KY et al (2022) Modeling receptor flexibility in the structure-based design of KRASG12C inhibitors. J Comput Aided Mol Des 36(8):591–604. https://doi.org/10.1007/s10822-022-00467-0
Canon J, Rex K, Saiki AY et al (2019) The clinical KRAS (G12C) inhibitor AMG 510 drives anti-tumour immunity. Nature 575(7781):217–223. https://doi.org/10.1038/s41586-019-1694-1
Nakajima EC, Drezner N, Li X et al (2022) FDA approval summary: sotorasib for KRAS G12C-mutated metastatic NSCLC. Clin Cancer Res 28(8):1482–1486. https://doi.org/10.1158/1078-0432.CCR-21-3074
Fell JB, Fischer JP, Baer BR et al (2020) Identification of the clinical development candidate MRTX849, a covalent KRASG12C inhibitor for the treatment of cancer. J Med Chem 63(13):6679–6693. https://doi.org/10.1021/acs.jmedchem.9b02052
Hallin J, Engstrom LD, Hargis L et al (2020) The KRASG12C inhibitor MRTX849 provides insight toward therapeutic susceptibility of KRAS-mutant cancers in mouse models and patients therapeutic insight from the KRASG12C inhibitor MRTX849. Cancer Discov 10(1):54–71. https://doi.org/10.1158/2159-8290.CD-19-1167
Prior IA, Hood FE, Hartley JL (2020) The frequency of ras mutations in cancer ras cancer statistics. Can Res 80(14):2969–2974. https://doi.org/10.1158/0008-5472.CAN-19-3682
Eser S, Schnieke A, Schneider G, Saur D (2014) Oncogenic KRAS signalling in pancreatic cancer. Br J Cancer 111(5):817–822. https://doi.org/10.1158/10.1038/bjc.2014.215
Stephen AG, Esposito D, Bagni RK, McCormick F (2014) Dragging ras back in the ring. Cancer Cell 25(3):272–281. https://doi.org/10.1016/j.ccr.2014.02.017
McCormick F (2015) KRAS as a therapeutic target. Clin Cancer Res 21(8):1797–1801. https://doi.org/10.1158/1078-0432.CCR-14-2662
Wang X, Allen S, Blake JF et al (2021) Identification of MRTX1133, a noncovalent, potent, and selective KRASG12D inhibitor. J Med Chem 65(4):3123–3133. https://doi.org/10.1021/acs.jmedchem.1c01688
Hallin J, Bowcut V, Calinisan A et al (2022) Anti-tumor efficacy of a potent and selective non-covalent KRASG12D inhibitor. Nat Med 10:2171–2182. https://doi.org/10.1038/s41591-022-02007-7
Issahaku AR, Mukelabai N, Agoni C et al (2022) Characterization of the binding of MRTX1133 as an avenue for the discovery of potential KRASG12D inhibitors for cancer therapy. Sci Rep 12(1):17796. https://doi.org/10.1038/s41598-022-22668-1
Vasta JD, Peacock DM, Zheng Q et al (2022) KRAS is vulnerable to reversible switch-II pocket engagement in cells. Nat Chem Biol 18(6):596–604. https://doi.org/10.1038/s41589-022-00985-w
Zheng Q, Peacock DM, Shokat KM (2022) Drugging the next undruggable KRAS allele-Gly12Asp. J Med Chem 65(4):3119–3122. https://doi.org/10.1021/acs.jmedchem.2c00099
Hallin J, Bowcut V, Calinisan A et al (2022) Anti-tumor efficacy of a potent and selective non-covalent KRASG12D inhibitor. Nat Med 28(10):2171–2182. https://doi.org/10.1038/s41591-022-02007-7
Mccammon JA, Gelin BR, Karplus M (1977) Dynamics of folded proteins. Nature 267(5612):585–590. https://doi.org/10.1038/267585a0
Piana S, Lindorff-Larsen K, Shaw DE (2012) Protein folding kinetics and thermodynamics from atomistic simulation. Proc Natl Acad Sci 109(44):17845–17850. https://doi.org/10.1073/pnas.1201811109
Hu J, Sun X, Kang Z, Cheng J (2023) Computational investigation of functional water molecules in GPCRs bound to G protein or arrestin. J Comput Aided Mol Des 37(2):91–105. https://doi.org/10.1007/s10822-022-00492-z
Rudrapal M, Issahaku AR, Agoni C et al (2022) In silico screening of phytopolyphenolics for the identification of bioactive compounds as novel protease inhibitors effective against SARS-CoV-2. J Biomol Struct Dyn 40(20):10437–10453. https://doi.org/10.1080/07391102.2021.1944909
Pande VS, Beauchamp K, Bowman GR (2010) Everything you wanted to know about Markov state models but were afraid to ask. Methods 52(1):99–105. https://doi.org/10.1016/j.ymeth.2010.06.002
Chodera JD, Noé F (2014) Markov state models of biomolecular conformational dynamics. Curr Opin Struct Biol 25:135–144. https://doi.org/10.1016/j.sbi.2014.04.002
Schrödinger Release 2015-3 (2015) Maestro. Schrödinger, LLC, New York
Schrödinger Release 2015-3 (2015) Desmond molecular dynamics system. Schrödinger, LLC, New York
Harder E, Damm W, Maple J et al (2016) OPLS3: a force field providing broad coverage of drug-like small molecules and proteins. J Chem Theory Comput 12(1):281–296. https://doi.org/10.1021/acs.jctc.5b00864
Martyna GJ, Klein ML, Tuckerman M (1992) Nosé-Hoover chains: the canonical ensemble via continuous dynamics. J Chem Phys 97(4):2635–2643. https://doi.org/10.1063/1.463940
Martyna GJ, Tobias DJ, Klein ML (1994) Constant pressure molecular dynamics algorithms. J Chem Phys 101(5):4177–4189. https://doi.org/10.1063/1.467468
Pedregosa F, Varoquaux G, Gramfort A et al (2011) Scikit-learn: machine learning in Python. J Mach Learn Res 12:2825–2830. https://doi.org/10.48550/arXiv.1201.0490
Gowers RJ, Linke M, Barnoud J, et al (2016) MDAnalysis: a Python package for the rapid analysis of molecular dynamics simulations. In: Proceedings of the 15th python in science conference. SciPy Austin, p 105. https://doi.org/10.25080/Majora-629e541a-00e
Michaud-Agrawal N, Denning EJ, Woolf TB, Beckstein O (2011) MDAnalysis: a toolkit for the analysis of molecular dynamics simulations. J Comput Chem 32(10):2319–2327. https://doi.org/10.1002/jcc.21787
Sun X, Singh S, Blumer KJ, Bowman GR (2018) Simulation of spontaneous G protein activation reveals a new intermediate driving GDP unbinding. Elife 7:e38465. https://doi.org/10.7554/eLife.38465.001
Norris JR (1998) Markov chains. Cambridge University Press, Cambridge
Lu S, Jang H, Nussinov R, Zhang J (2016) The structural basis of oncogenic mutations G12, G13 and Q61 in small GTPase K-Ras4B. Sci Rep 6(1):1–15. https://doi.org/10.1038/srep21949
Shima F, Ijiri Y, Muraoka S et al (2010) Structural basis for conformational dynamics of GTP-bound Ras protein. J Biol Chem 285(29):22696–22705. https://doi.org/10.1074/jbc.M110.125161
Araki M, Shima F, Yoshikawa Y et al (2011) Solution structure of the state 1 conformer of GTP-bound H-Ras protein and distinct dynamic properties between the state 1 and state 2 conformers. J Biol Chem 286(45):39644–39653. https://doi.org/10.1074/jbc.M111.227074
Lu S, Jang H, Muratcioglu S et al (2016) Ras conformational ensembles, allostery, and signaling. Chem Rev 116(11):6607–6665. https://doi.org/10.1021/acs.chemrev.5b00542
Pagba CV, Abebe AG, Gilbertson SR, Dilsha K (2021) Synthesis of substituted 7-(piperazin-1-yl)pyrazolo[1,5-a]pyrimidine analogs as inhibitors of KRAS. The Board of Regents of the University of Texas System, Assignee. Patent WO2021119343A1
Mao Z, Xiao H, Shen P et al (2022) KRAS (G12D) can be targeted by potent inhibitors via formation of salt bridge. Cell Discov 8(1):1–14. https://doi.org/10.1038/s41421-021-00368-w
Acknowledgements
F.L. would like to thank Zhenchao Wei for his help with the analysis script.
Funding
This work was supported by the National Natural Science Foundation of China (Grant 22063004), the Youth project of Jiangxi Provincial Department of Education (Grant GJJ170696), the University Doctoral Fund (Grant 2017BSQD016), the Open Fund of Provincial Research Platform (Grant KFGJ19014), and the horizontal research funding (Grant H20210713181758000003).
Author information
Authors and Affiliations
Contributions
FL carried out the molecular dynamics simulations. FL and XS designed the study and analyzed the data. JC was responsible for the project. FL, XS, ZK, XD, HH, and JC contributed to writing and commenting on the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing financial interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liang, F., Kang, Z., Sun, X. et al. Inhibition mechanism of MRTX1133 on KRASG12D: a molecular dynamics simulation and Markov state model study. J Comput Aided Mol Des 37, 157–166 (2023). https://doi.org/10.1007/s10822-023-00498-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-023-00498-1