Abstract
Purpose
Successful identification of transcriptomic biomarkers within human IVF embryos may enhance implantation prediction and provide insights not available through conventional embryo biopsy genomic analysis. We demonstrate proof-of-concept for a methodology to assess overall embryo gene expression using qPCR with blastocoel fluid-conditioned media by examining the comparative presence of apoptotic genes.
Methods
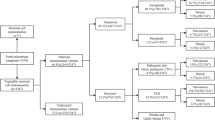
Blastocoel fluid-conditioned media were collected from 19 embryos (11 euploid) following trophectoderm biopsy of day-5 ICSI-IVF blastocysts. Media were assessed for apoptotic gene expression via qPCR. Statistical analysis of gene expression was conducted via Wilcoxon Signed-Ranks test (overall expression), multivariate ANOVA (functional gene groups), and chi-square test of independence (gene level).
Results
A significantly higher overall apoptotic gene expression within euploid versus aneuploid embryos (p = 0.001) was observed. There was significantly (p = 0.045) higher expression of pro-apoptotic genes between implanted and not implanted embryos. Pro- vs. anti-apoptotic gene expression from all euploid embryos approached significance (p = 0.053). The ploidy status-based claim is further substantiated at the gene level with significantly higher expression of BBC3 (p = 0.012) and BCL2L13 (p = 0.003) in euploid embryos compared to aneuploid embryos.
Conclusions
In this preliminary study, we demonstrate that (1) qualitative analysis of blastocoel fluid-conditioned media gene expression is possible, (2) global trends of expression are potentially related to clinical outcomes, and (3) gene-level expression trends exist and may be another viable metric for comparative expression between samples. The presence of statistical significance within analyses conducted with this sample size warrants a larger investigation of blastocoel fluid-conditioned media as an additional beneficial predictive tool for future IVF cases.

Similar content being viewed by others
References
Vidal F, Gimenez C, Rubio C, Simon C, Pellicer A, Santalo J, et al. FISH preimplantation diagnosis of chromosome aneuploidy in recurrent pregnancy wastage. J Assist Reprod Genet. 1998;15:310–3.
Franasiak JM, Scott RT Jr. Embryonic aneuploidy: overcoming molecular genetics challenges improves outcomes and changes practice patterns. Trends Mol Med. 2014;20:499–508.
Balaban B, Gardner DK. Morphological assessment of blastocyst stage embryos: types of grading systems and their reported outcomes. In: Gardner D, Sakkas D, Seli E, Wells D, editors. Human Gametes and Preimplantation Embryos. New York: Springer; 2013. p. 31–43.
Gonzalez XV, Odia R, Naja R, Serhal P, Saab W, Seshadri S, et al. Euploid blastocysts implant irrespective of their morphology after NGS-(PGT-A) testing in advanced maternal age patients. J Assist Reprod Genet. 2019;36:1–7.
Lal A, Roudebush WE, Chosed RJ. Embryo biopsy can offer more information than just ploidy status. Front in Cell and Dev Biol. 2020;8.
Capalbo A, Ubaldi FM, Cimadomo D, Noli L, Khalaf Y, Farcomeni A, et al. MicroRNAs in spent blastocyst culture medium are derived from trophectoderm cells and can be explored for human embryo reproductive competence assessment. Fertil Steril. 2016;105:225–35.
Palini S, Galluzzi L, De Stefani S, Bianchi M, Wells D, Magnani M, et al. Genomic DNA in human blastocoele fluid. Reprod Biomed Online. 2013;26:603–10.
Poli M, Ori A, Child T, Jaroudi S, Spath K, Beck M, et al. Characterization and quantification of proteins secreted by single human embryos prior to implantation. EMBO Mol Med. 2015;7:1465–79.
Tobler K, Zhao Y, Ross R, Benner A, Xu X, Du L, et al. Blastocoel fluid (bf) harbors embryonic DNA that may result from the marginalization of aneuploid cells during embryogenesis. Fertil Steril. 2014;102: e205.
Hammond ER, McGillivray BC, Wicker SM, Peek JC, Shelling AN, Stone P, et al. Characterizing nuclear and mitochondrial DNA in spent embryo culture media: genetic contamination identified. Fertil Steril. 2017;107:220-228.e5.
Fragouli E, Munne S, Wells D. The cytogenetic constitution of human blastocysts: insights from comprehensive chromosome screening strategies. Hum Reprod Update. 2019;25:15–33.
Brezina PR, Anchan R, Kearns WG. Preimplantation genetic testing for aneuploidy: what technology should you use and what are the differences? J Assist Reprod Genet. 2016;33:823–32.
Battaglia R, Palini S, Vento ME, La Ferlita A, Lo Faro MJ, Caroppo E, et al. Identification of extracellular vesicles and characterization of miRNA expression profiles in human blastocoel fluid. Sci Rep. 2019;9:84.
Hammond ER, Shelling AN, Cree LM. Nuclear and mitochondrial DNA in blastocoele fluid and embryo culture medium: evidence and potential clinical use. Hum Reprod. 2016;31:1653–61.
Zhang Y, Li N, Wang L, Sun H, Ma M, Wang H, et al. Molecular analysis of DNA in blastocoele fluid using next-generation sequencing. J Assist Reprod Genet. 2016;33:637–45.
Hardy K. Apoptosis in the human embryo. Rev Reprod. 1999;4:125–34.
Brill A, Torchinsky A, Carp H, Toder V. The role of apoptosis in normal and abnormal embryonic development. J Assist Reprod Genet. 1999;16:512–9.
Jurisicova A, Acton BM. Deadly decisions: the role of genes regulating programmed cell death in human preimplantation embryo development. Reproduction. 2004;128:281–91.
Jurisicova A, Varmuza S, Casper R. Programmed cell death and human embryo fragmentation. MHR Basic Sci Reprod Med. 1996;2:93–8.
Brison DR. Apoptosis in mammalian preimplantation embryos: regulation by survival factors. Hum Fertil. 2000;3:36–47.
Jacobs K, Van de Velde H, De Paepe C, Sermon K, Spits C. Mitotic spindle disruption in human preimplantation embryos activates the spindle assembly checkpoint but not apoptosis until Day 5 of development. Mol Hum Reprod. 2017;23:321–9.
Spanos S, Rice S, Karagiannis P, Taylor D, Becker D, Winston R, et al. Caspase activity and expression of cell death genes during development of human preimplantation embryos. Reproduction. 2002;124:353–63.
Rule K, Chosed RJ, Chang TA, Wininger JD, Roudebush WE. Relationship between blastocoel cell-free DNA and day-5 blastocyst morphology. J Assist Reprod Genet. 2018;35:1497–501.
Metcalfe AD, Hunter HR, Bloor DJ, Lieberman BA, Picton HM, Leese HJ, et al. Expression of 11 members of the BCL-2 family of apoptosis regulatory molecules during human preimplantation embryo development and fragmentation. Mol Reprod Dev Incorp Gamete Res. 2004;68:35–50.
Wells D, Bermudez M, Steuerwald N, Thornhill A, Walker D, Malter H, et al. Expression of genes regulating chromosome segregation, the cell cycle and apoptosis during human preimplantation development. Hum Reprod. 2005;20:1339–48.
Bolton H, Graham SJ, Van der Aa N, Kumar P, Theunis K, Fernandez Gallardo E, et al. Mouse model of chromosome mosaicism reveals lineage-specific depletion of aneuploid cells and normal developmental potential. Nat Commun. 2016;7:11165.
Capalbo A, Romanelli V, Patassini C, Poli M, Girardi L, Giancani A, Stoppa M, Cimadomo D, Ubaldi FM, Rienzi L. Diagnostic efficacy of blastocoel fluid and spent media as sources of DNA for preimplantation genetic testing in standard clinical conditions. Fertil Steril. 2018;110(5):870-879.e5. Erratum in: Fertil Steril. 2019;111(1):194 https://doi.org/10.1016/j.fertnstert.2018.05.031.
Huang L, Bogale B, Tang Y, Lu S, Xie XS, Racowsky C. Noninvasive preimplantation genetic testing for aneuploidy in spent medium may be more reliable than trophectoderm biopsy. Proc Natl Acad Sci USA. 2019;116(28):14105–12.
Lal A, Roudebush WE, Mainigi M, Chosed RJ. Fluorescent-dependent comparative Ct method for qPCR gene expression analysis in IVF clinical pre-implantation embryonic testing. Biol Methods Protoc. 2021;6:bpab001. https://doi.org/10.1093/biomethods/bpab001.
Yang M, Rito T, Metzger J, Naftaly J, Soman R, Hu J, et al. Depletion of aneuploid cells in human embryos and gastruloids. Nat Cell Biol. 2021;23(4):314–21.
Ntostis P, Kokkali G, Iles D, Huntriss J, Tzetis M, Picton H, et al. Can trophectoderm RNA analysis predict human blastocyst competency? Syst Biol in Reprod Med. 2019;65:312–25.
Licciardi F, Lhakhang T, Kramer YG, Zhang Y, Heguy A, Tsirigos A. Human blastocysts of normal and abnormal karyotypes display distinct transcriptome profiles. Sci Rep. 2018;8:14906.
Vera-Rodriguez M, Chavez SL, Rubio C, Reijo Pera RA, Simon C. Prediction model for aneuploidy in early human embryo development revealed by single-cell analysis. Nat Commun. 2015;6:7601.
Parks JC, McCallie BR, Janesch AM, Schoolcraft WB, Katz-Jaffe MG. Blastocyst gene expression correlates with implantation potential. Fertil Steril. 2011;95:1367–72.
Kirkegaard K, Villesen P, Jensen JM, Hindkjaer JJ, Kolvraa S, Ingerslev HJ, et al. Distinct differences in global gene expression profiles in non-implanted blastocysts and blastocysts resulting in live birth. Gene. 2015;571:212–20.
McCallie BR, Parks JC, Griffin DK, Schoolcraft WB, Katz-Jaffe MG. Infertility diagnosis has a significant impact on the transcriptome of developing blastocysts. Mol Hum Reprod. 2017;23:549–56.
McCallie BR, Parks JC, Trahan GD, Jones KL, Coate BD, Griffin DK, et al. Compromised global embryonic transcriptome associated with advanced maternal age. J Assist Reprod Genet. 2019;36:915–24.
Acknowledgements
We thank Alex Ewing for his statistical analysis recommendations for this dataset.
Funding
University of South Carolina Magellan Scholar funds supported JB and offset some costs for experiments. University of South Carolina School of Medicine Greenville development funds (to RJC and WER) offset experimental costs.
Author information
Authors and Affiliations
Contributions
R. J. C. and W. E. R. designed the experiments. S. Z., T. A. C., R. D. R., and J. D. W. carried out sample collection, pooled patient data records, and contributed to the revision of the manuscript. A. L., A. K., J. B., D. A., A. B, and A. B. carried out the cDNA synthesis and real-time experiments and aided in data analysis. A. L. led figure construction and played an integral role in manuscript writing. L. A. F. performed the statistical analysis on the dataset. A. L. and R. J. C. led the data analysis, interpretation of results, and manuscript writing. All authors contributed to the manuscript drafting process.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lal, A., Kranyak, A., Blalock, J. et al. Apoptotic qPCR gene expression array analysis demonstrates proof-of-concept for rapid blastocoel fluid-conditioned media molecular prediction. J Assist Reprod Genet 39, 1515–1522 (2022). https://doi.org/10.1007/s10815-022-02510-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10815-022-02510-3