Abstract
Purpose
The present study aimed to identify the molecular etiology of non-syndromic congenital cataract (CC) using whole-exome sequencing (WES) analysis.
Methods
In the present study, ophthalmologic results and pedigree analysis of the families of 12 patients with non-syndromic CC were evaluated. WES analysis was conducted after DNA was isolated from peripheral blood samples obtained from the patients.
Results
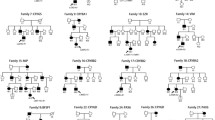
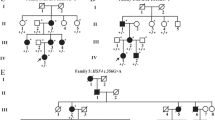
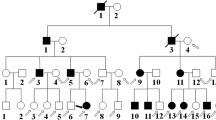
Twelve non-syndromic probands (10 males and 2 females) with bilateral CC were included in the study. Patient age ranged between 1 and 11 months. WES analysis showed pathogenic/likely pathogenic variant in 7 (58%) of the 12 families and variant of unknown significance (VUS) in 5 (42%) of them. All the 13 different variants detected in 9 different CC-related genes were co-segregated with the disease. Autosomal dominant inheritance was found in 7 (58%) of the families and autosomal recessive inheritance was found in 5 (42%) of them.
Conclusion
To the best of our knowledge, the present research is one of the limited numbers of studies in the Turkish population in which genetically heterogeneous non-syndromic CC was investigated using WES analysis. Novel variants that we identified in DNMBP, LSS, and WFS1 genes, which are rarely associated with the CC phenotype, have contributed to the mutation spectrum of this disease. Identifying the relevant molecular genetic etiology allows accurate genetic counseling to be provided to the families.


Similar content being viewed by others
Data availability
The authors declare that materials described in the manuscript, including all relevant raw data, will be freely available to any scientist wishing to use them for noncommercial purposes, without breaching participant confidentiality. Moreover, the authors ensure that their datasets are presented in the main manuscript.
References
Sheeladevi S, Lawrenson JG, Fielder AR, Suttle CM (2016) Global prevalence of childhood cataract: a systematic review. Eye (Lond) 30:1160–1169
Churchill A, Graw J (2011) Clinical and experimental advances in congenital and paediatric cataracts. Philos Trans R Soc Lond B Biol Sci 366:1234–1249
Berry V, Ionides A, Pontikos N et al (2020) The genetic landscape of crystallins in congenital cataract. Orphanet J Rare Dis 15:333
Messina-Baas O, Cuevas-Covarrubias SA (2017) Inherited congenital cataract: a guide to suspect the genetic etiology in the cataract genesis. Mol Syndromol 8:58–78
Fernández-Alcalde C, Nieves-Moreno M, Noval S et al (2021) Molecular and genetic mechanism of non-syndromic congenital cataracts. Mutat Screen Span Fam Genes (Basel) 12:580
Li H, Durbin R (2010) Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26:589–595
Kopanos C, Tsiolkas V, Kouris A et al (2019) VarSome: the human genomic variant search engine. Bioinformatics 35:1978–1980
Richards S, Aziz N, Bale S et al (2015) ACMG laboratory quality assurance committee. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American college of medical genetics and genomics and the association for molecular pathology. Genet Med 17:405–424
Venselaar H, Te Beek TA, Kuipers RK, Hekkelman ML, Vriend G (2010) Protein structure analysis of mutations causing inheritable diseases. An e-science approach with life scientist friendly interfaces. BMC Bioinform 11:548
Gillespie RL, O’Sullivan J, Ashworth J et al (2014) Personalized diagnosis and management of congenital cataract by next-generation sequencing. Ophthalmology 121:2124–2137
Lenassi E, Clayton-Smith J, Douzgou S et al (2020) Clinical utility of genetic testing in 201 preschool children with inherited eye disorders. Genet Med 22:745–751
Ma AS, Grigg JR, Ho G et al (2016) Sporadic and familial congenital cataracts: mutational spectrum and new diagnoses using next-generation sequencing. Hum Mutat 37:371–384
Javadiyan S, Craig JE, Souzeau E et al (2017) High-throughput genetic screening of 51 Pediatric cataract genes ıdentifies causative mutations in ınherited pediatric cataract in South Eastern Australia. G3 (Bethesda) 7:3257–3268
Ma A, Grigg JR, Flaherty M et al (2021) Genome sequencing in congenital cataracts improves diagnostic yield. Hum Mutat 42:1173–1183
Jin HS, Kim J, Kwak W et al (2017) Identification of a novel mutation in BRD4 that causes autosomal dominant syndromic congenital Cataracts associated with other neuro-skeletal anomalies. PLoS ONE 12:e0169226
Choquet H, Melles RB, Anand D et al (2021) A large multiethnic GWAS meta-analysis of cataract identifies new risk loci and sex-specific effects. Nat Commun 12(1):3595
Berry V, Pontikos N, Dudakova L et al (2020) A novel missense mutation in LIM2 causing isolated autosomal dominant congenital cataract. Ophthalmic Genet 41:131–134
Chen P, Chen H, Pan XJ, Tang SZ, Xia YJ, Zhang H (2018) Novel mutations in CRYBB1/CRYBB2 identified by targeted exome sequencing in Chinese families with congenital cataract. Int J Ophthalmol 11:1577–1582
Bell S, Malka S, Lloyd IC, Moosajee M (2021) Clinical spectrum and genetic diagnosis of 54 consecutive patients aged 0–25 with bilateral cataracts. Genes (Basel) 12(2):131
Ye Y, Wu M, Qiao Y, Xie T, Yu Y, Yao K (2019) Identification and preliminary functional analysis of two novel congenital cataract associated mutations of Cx46 and Cx50. Ophthalmic Genet 40:428–435
Li B, Liu Y, Liu Y, Guo H, Hu Z, Xia K, Jin X (2016) Identification of a GJA3 mutation in a large family with bilateral congenital cataract. DNA Cell Biol 35:135–139
Ansar M, Chung HL, Taylor RL et al (2018) Bi-allelic loss-of-function variants in DNMBP cause infantile cataracts. Am J Hum Genet 103:568–578
Berry V, Gregory-Evans C, Emmett W, Waseem N, Raby J, Prescott D, Moore AT, Bhattacharya SS (2013) Wolfram gene (WFS1) mutation causes autosomal dominant congenital nuclear cataract in humans. Eur J Hum Genet 21(12):1356–1360
Rechsteiner D, Issler L, Koller S et al (2021) Genetic analysis in a swiss cohort of bilateral congenital cataract. JAMA Ophthalmol 139:691–700
Sloan-Heggen CM, Bierer AO, Shearer AE, Kolbe DL, Nishimura CJ, Frees KL, Ephraim SS, Shibata SB, Booth KT, Campbell CA, Ranum PT, Weaver AE, Black-Ziegelbein EA, Wang D, Azaiez H, Smith RJH (2016) Comprehensive genetic testing in the clinical evaluation of 1119 patients with hearing loss. Hum Genet 135(4):441–450
Pankiv S, Alemu EA, Brech A et al (2010) FYCO1 is a Rab7 effector that binds to LC3 and PI3P to mediate microtubule plus end-directed vesicle transport. J Cell Biol 188:253–269
Capalbo A, Valero RA, Jimenez-Almazan J, Pardo PM, Fabiani M, Jiménez D, Simon C, Rodriguez JM (2019) Optimizing clinical exome design and parallel gene-testing for recessive genetic conditions in preconception carrier screening: translational research genomic data from 14,125 exomes. PLoS Genet 15(10):e1008409
Borck G, Kakar N, Hoch J et al (2012) An Alu repeat-mediated genomic GCNT2 deletion underlies congenital cataracts and adult i blood group. Hum Genet 131:209–216
Zhao L, Chen XJ, Zhu J et al (2015) Lanosterol reverses protein aggregation in cataracts. Nature 523:607–611
Funding
The authors did not receive support from any organization for the submitted work.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by AT, ATK, SÖY, SGS, SS. Genetic analysis was performed by AT. The first draft of the manuscript was written by AT and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of ınterest
All the authors declare that they have no conflict of interest and no financial disclosure.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. The study was approved by the Ethics Committee of the Kartal Dr. Lütfi Kırdar City Hospital, Istanbul, Turkey (decision date: 28 April 2021, No. 2021/514/200/3).
Consent to participate
Informed consent was obtained from legal guardians.
Consent to publish
Additional informed consent was obtained from all legal guardians for whom identifying information is included in this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Türkyılmaz, A., Kaplan, A.T., Öskan Yalçın, S. et al. Identification of novel variants in Turkish families with non-syndromic congenital cataracts using whole-exome sequencing. Int Ophthalmol 43, 4573–4583 (2023). https://doi.org/10.1007/s10792-023-02857-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10792-023-02857-1