Abstract
Background
Homeobox B9 (HOXB9) is one of the HOX family of transcription factors that are essential for cancer development and embryonic growth. However, the clinical importance and biological involvement of HOXB9 in colon cancer (CC) are not adequately understood.
Aims
To investigate whether HOXB9 participates in the proliferation, invasion, and migration of CC.
Methods
This study investigated the function and clinical significance of HOXB9 mRNA and protein expression in CC. Furthermore, overexpression and knockdown experiments of HOXB9 were developed to explore their effects on CC cell transwell and proliferation. Moreover, a molecular mechanism of HOXB9 regulate serine/arginine-rich splicing factor 3 (SRSF3) was explored.
Results
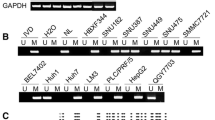
HOXB9 expression was higher in CC cells and tissues at both the mRNA and protein levels. Poor survival in CC patients was significantly connected with high HOXB9 expression, which was also strongly associated with the TNM stage and lymph node metastases. Furthermore, in vitro CC cell proliferation, transwell were markedly aided by HOXB9 overexpression. Contrarily, HOXB9 knockdown had the reverse result and inhibited the formation of xenograft tumors in naked mice. Gene set enrichment analysis (GSEA) revealed a correlation between high HOXB9 expression and spliceosomes. JASPAR and GEPIA2.0, in addition to CHIP and dual-luciferase reporting assays, confirmed that HOXB9 targets the promoter of SRSF3 to enhance its expression. We also found that SRSF3 knockdown eliminated HOXB9 from cell proliferation and transwell.
Conclusion
We characterized the function and mechanism of HOXB9 in regulating colon cancer growth, suggesting a novel molecular approach for colon cancer-targeted therapy.










Similar content being viewed by others
References
Arbyn M, Weiderpass E, Bruni L et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 2018;68:394–424.
Sung H, Ferlay J, Siegel RL et al. Cancer statistics, 2021. CA Cancer J Clin 2021;71:7–33.
Singh D, Vaccarella S, Gini A, De Paula SN, Steliarova-Foucher E, Bray F. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 2021;71:209–249.
Markowitz SD, Bertagnolli MM. Molecular origins of cancer: molecular basis of colorectal cancer. N Engl J Med 2009;361:2449–2460.
Dekker E, Tanis PJ, Vleugels JLA, Kasi PM, Wallace MB. Colorectal cancer. Lancet 2019;394:1467–1480.
Tauriello DV, Palomo-Ponce S, Stork D et al. TGFβ drives immune evasion in genetically reconstituted colon cancer metastasis. Nature 2018;554:538–543.
Araghi M, Arnold M, Rutherford MJ et al. Colon and rectal cancer survival in seven high-income countries 2010–2014: variation by age and stage at diagnosis (the ICBP SURVMARK-2 project). Gut 2021;70:114–126.
Garcia-Fernàndez J. The genesis and evolution of homeobox gene clusters. Nat Rev Genet 2005;6:881–892.
Grier DG, Thompson A, Kwasniewska A, McGonigle GJ, Halliday HL, Lappin TR. The pathophysiology of HOX genes and their role in cancer. J Pathol 2005;205:154–171.
Wang T, Guo H, Li Q et al. The AMPK-HOXB9-KRAS axis regulates lung adenocarcinoma growth in response to cellular energy alterations. Cell Rep 2022;40:111210.
Hayashida T, Takahashi F, Chiba N et al. HOXB9, a gene overexpressed in breast cancer, promotes tumorigenicity and lung metastasis. Proc Natl Acad Sci U S A 2010;107:1100–1105.
Yao Y, Liu C, Wang B et al. HOXB9 blocks cell cycle progression to inhibit pancreatic cancer cell proliferation through the DNMT1/RBL2/c-Myc axis. Cancer Lett 2022;533:215595.
Zhang L, Wu Q, He C et al. HOXB9 inhibits proliferation in gastric carcinoma cells via suppression of phosphorylated-Akt and NF-κB-dependent Snail expression. Dig Liver Dis 2019;51:157–165.
Nguyen DX, Chiang AC, Zhang XH et al. WNT/TCF signaling through LEF1 and HOXB9 mediates lung adenocarcinoma metastasis. Cell 2009;138:51–62.
Wan J, Xu W, Zhan J et al. PCAF-mediated acetylation of transcriptional factor HOXB9 suppresses lung adenocarcinoma progression by targeting oncogenic protein JMJD6. Nucleic Acids Res 2016;44:10662–10675.
Song J, Wang T, Xu W et al. HOXB9 acetylation at K27 is responsible for its suppression of colon cancer progression. Cancer Lett 2018;426:63–72.
Wan J, Liu H, Feng Q, Liu J, Ming L. HOXB9 promotes endometrial cancer progression by targeting E2F3. Cell Death Dis 2018;9:509.
Chiba N, Comaills V, Shiotani B et al. Homeobox B9 induces epithelial-to-mesenchymal transition-associated radioresistance by accelerating DNA damage responses. Proc Natl Acad Sci U S A 2012;109:2760–2765.
Huang K, Yuan R, Wang K et al. Overexpression of HOXB9 promotes metastasis and indicates poor prognosis in colon cancer. Chin J Cancer Res 2014;26:72–80.
Mure F, Corbin A, Benbahouche NE, Bertrand E, Manet E, Gruffat H. The splicing factor SRSF3 is functionally connected to the nuclear RNA exosome for intronless mRNA decay. Sci Rep 2018;8:12901.
Jumaa H, Nielsen PJ. The splicing factor SRp20 modifies splicing of its own mRNA and ASF/SF2 antagonizes this regulation. Embo j 1997;16:5077–5085.
Zheng X, Peng Q, Wang L et al. Serine/arginine-rich splicing factors: the bridge linking alternative splicing and cancer. Int J Biol Sci 2020;16:2442–2453.
Long JC, Caceres JF. The SR protein family of splicing factors: master regulators of gene expression. Biochem J 2009;417:15–27.
Do DV, Strauss B, Cukuroglu E et al. SRSF3 maintains transcriptome integrity in oocytes by regulation of alternative splicing and transposable elements. Cell Discov 2018;4:33.
Jia R, Li C, McCoy JP, Deng CX. Zheng ZM SRp20 is a proto-oncogene critical for cell proliferation and tumor induction and maintenance. Int J Biol Sci 2010;6:806–826.
Song X, Wan X, Huang T et al. SRSF3-regulated RNA alternative splicing promotes glioblastoma tumorigenicity by affecting multiple cellular processes. Cancer Res 2019;79:5288–5301.
Fuentes-Fayos AC, Vázquez-Borrego MC, Jimenez-Vacas JM et al. Splicing machinery dysregulation drives glioblastoma development/aggressiveness: oncogenic role of SRSF3. Brain 2020;143:3273–3293.
Kurokawa K, Akaike Y, Masuda K et al. Downregulation of serine/arginine-rich splicing factor 3 induces G1 cell cycle arrest and apoptosis in colon cancer cells. Oncogene 2014;33:1407–1417.
Chen Y, Yang M, Meng F et al. SRSF3 promotes angiogenesis in colorectal cancer by splicing SRF. Front Oncol 2022;12:810610.
Franken NA, Rodermond HM, Stap J, Haveman J, Van Bree C. Clonogenic assay of cells in vitro. Nat Protoc 2006;1:2315–2319.
Wittekind C, Compton CC, Greene FL, Sobin LH. TNM residual tumor classification revisited. Cancer 2002;94:2511–2516.
Castro-Mondragon JA et al. JASPAR 2022: the 9th release of the open-access database of transcription factor binding profiles. Nucleic Acids Res 2022;50:D165–D173.
Sandelin A, Alkema W, Engström P, Wasserman WW, Lenhard B. JASPAR: an open-access database for eukaryotic transcription factor binding profiles. Nucleic Acids Res 2004;32:91–94.
Sobecki M, Mrouj K, Camasses A et al. The cell proliferation antigen Ki-67 organises heterochromatin. Elife 2016;5:e13722.
Sobecki M, Mrouj K, Colinge J et al. Cell-cycle regulation accounts for variability in Ki-67 expression levels. Cancer Res 2017;77:2722–2734.
Corbo C, Orrù S, Gemei M et al. Protein cross-talk in CD133+ colon cancer cells indicates activation of the Wnt pathway and upregulation of SRp20 that is potentially involved in tumorigenicity. Proteomics 2012;12:2045–2059.
Gonçalves V, Matos P, Jordan P. Antagonistic SR proteins regulate alternative splicing of tumor-related Rac1b downstream of the PI3-kinase and Wnt pathways. Hum Mol Genet 2009;18:3696–3707.
Torres S, García-Palmero I, Marín-Vicente C et al. Proteomic characterization of transcription and splicing factors associated with a metastatic phenotype in colorectal cancer. J Proteome Res 2018;17:252–264.
Luo L, Yang X, Takihara Y, Knoetgen H, Kessel M. The cell-cycle regulator geminin inhibits Hox function through direct and polycomb-mediated interactions. Nature 2004;427:749–753.
Abate-Shen C. Deregulated homeobox gene expression in cancer: cause or consequence? Nat Rev Cancer 2002;2:777–785.
Xiong H, Xiao H, Luo C et al. GRP78 activates the Wnt/HOXB9 pathway to promote invasion and metastasis of hepatocellular carcinoma by chaperoning LRP6. Exp Cell Res 2019;383:111493.
Lin J, Zhang D, Fan Y et al. Regulation of cancer stem cell self-renewal by HOXB9 antagonizes endoplasmic reticulum stress-induced melanoma cell apoptosis via the miR-765-FOXA2 axis. J Invest Dermatol 2018;138:1609–1619.
Francis JC, Gardiner JR, Renaud Y et al. HOX genes promote cell proliferation and are potential therapeutic targets in adrenocortical tumours. Br J Cancer 2021;124:805–816.
Martinou E, Moller-Levet C, Karamanis D, Bagwan I, Angelidi AM. HOXB9 overexpression promotes colorectal cancer progression and is associated with worse survival in liver resection patients for colorectal liver metastases. Int J Mol Sci 2022;23:2281.
Martinou E, Falgari G, Bagwan I, Angelidi AM. A systematic review on hox genes as potential biomarkers in colorectal cancer: an emerging role of HOXB9. Int J Mol Sci 2021;22:13429.
Kuranaga Y, Sugito N, Shinohara H et al. SRSF3, a splicer of the PKM gene, regulates cell growth and maintenance of cancer-specific energy metabolism in colon cancer cells. Int J Mol Sci 2018;19:3012.
Wang JL, Guo CR, Sun TT et al. SRSF3 functions as an oncogene in colorectal cancer by regulating the expression of ArhGAP30. Cancer Cell Int 2020;20:120.
Zhang Y, Wang M, Meng F et al. A novel SRSF3 inhibitor, SFI003, exerts anticancer activity against colorectal cancer by modulating the SRSF3/DHCR24/ROS axis. Cell Death Discov 2022;8:238.
Zaborowski AM, Murphy B, Creavin B et al. Clinicopathological features and oncological outcomes of patients with young-onset rectal cancer. Br J Surg 2020;107:606–612.
Saraste D, Järås J, Martling A. Population-based analysis of outcomes with early-age colorectal cancer. Br J Surg 2020;107:301–309.
Acknowledgment
The authors thank the contributors to GEPIA 2.0 for contributing shared sequencing datasets for open access. Also, the GSEA database has provided us with new ideas to search for signaling pathway mechanisms, and furthermore, we are grateful to JASPAR for providing us with convenient information on transcription factor-related data.
Funding
This research was supported by the National Natural Science Foundation of China, No .81860432 (1) and the Natural Science Foundation of Jiangxi Province (20171BAB205064).
Author information
Authors and Affiliations
Contributions
WS conducted the study design. YL drafted the manuscript, CF participated in the coordination of the study and acted as technical advisor, WZ performed the analysis and collected the samples, LX performed the statistical analysis of the data, with equal contributions from YL and CF. CF, WZ and XD revised the manuscript. All authors reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
This study was approved by Ethics Committee of the Second Affiliated Hospital of Nanchang University (Nanchang, China).
Consent to participate
Each patient had signed the informed consent.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yuan, L., Cheng, F., Wu, Z. et al. Homeobox B9 Promotes Colon Cancer Progression by Targeting SRSF3. Dig Dis Sci 68, 3324–3340 (2023). https://doi.org/10.1007/s10620-023-07977-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10620-023-07977-3