Abstract
Brook trout (Salvelinus fontinalis) often exist as highly differentiated populations, even at small spatial scales, due either to natural or anthropogenic sources of isolation and low rates of dispersal. In this study, we used molecular approaches to describe the unique population structure of brook trout inhabiting the Shavers Fork watershed, located in eastern West Virginia, and contrast it to nearby populations in tributaries of the upper Greenbrier River and North Fork South Branch Potomac Rivers. Bayesian and maximum likelihood clustering methods identified minimal population structuring among 14 collections of brook trout from throughout the mainstem and tributaries of Shavers Fork, highlighting the role of the cold-water mainstem for connectivity and high rates of effective migration among tributaries. In contrast, the Potomac and Greenbrier River collections displayed distinct levels of population differentiation among tributaries, presumably resulting from tributary isolation by warm-water mainstems. Our results highlight the importance of protecting and restoring cold-water mainstem habitats as part of region-wide brook trout conservation efforts. In addition, our results from Shavers Fork provide a contrast to previous genetic studies that characterize Appalachian brook trout as fragmented isolates rather than well-mixed populations. Additional study is needed to determine whether the existence of brook trout as genetically similar populations among tributaries is truly unique and whether connectivity among brook trout populations can potentially be restored within other central Appalachian watersheds.




Similar content being viewed by others
References
Allendorf FW, Luikart G (2007) Conservation and the genetics of populations, 1st edn. Blackwell Scientific Publications, Malden
Barton NH, Slatkin M (1986) A quasi-equilibrium theory of the distribution of rare alleles in a subdivided population. Heredity 56:409–415
Cavalli-Sforza LL, Edwards AWF (1967) Phylogenetic analysis: models and estimation procedures. Evolution 21:550–570
Cornuet JM, Piry S, Luikart G, Estoup A, Solignac M (1999) New methods employing multilocus genotypes to select or exclude populations as origins of individuals. Genetics 153:1989–2000
Duchesne P, Turgeon J (2012) FLOCK provides reliable solutions to the “Number of populations” problem. J Hered. doi:10.1093/jhered/ess038
Duchesne P, Methot J, Turgeon J (2013) FLOCK 3.0 User Guide. www.bio.ulaval.ca/no_cache/en/department/professors/professors/professeur/11/13/. Accessed 1 June 2013
Earl DA, vonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4(2):359–361
EBTJV (Eastern Brook Trout Joint Venture) (2013) About the Eastern Brook Trout Joint Venture. www.easternbrooktrout.org. Accessed 1 June 2013
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14(8):2611–2620
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131:479–491
Fagan WF (2002) Connectivity, fragmentation, and extinction risk in dendritic metapopulations. Ecol 83:3243–3249
Fausch KD, Torgersen CE, Baxter CV, Li HW (2002) Landscapes to riverscapes: bridging the gap between research and conservation of stream fishes. Biosci 52(6):483–498
Flebbe PA, Roghair LD, Bruggink JL (2006) Spatial modeling to project southern Appalachian brook trout distribution in a warmer climate. Trans Am Fish Soc 135:1371–1382
Hammer O, Harper DAT, Ryan PD (2002) PAST—palaeontological statistics, version 0.82. Palaeontological Museum, University of Oslo, Oslo
Hedrick PW (2005) A standardized genetic differentiation measure. Evolution 59(8):1633–1638
Hudy M, Thieling TM, Gillespie N, Smith EP (2008) Distribution, status, and land use characteristics of subwatersheds within the native range of brook trout in the eastern United States. North Am J Fish Manage 28:1069–1085
Hudy M, Coombs JA, Nislow KH, Letcher BH (2010) Dispersal and within-stream spatial population structure of brook trout revealed by pedigree reconstruction analysis. Trans Am Fish Soc 139:1276–1287
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinforma 23(14):1801–1806
Kalinowski ST (2005) HP-Rare: a computer program for performing rarefaction on measures of allelic diversity. Mol Ecol Notes 5:187–189
Kanda N, Allendorf FW (2001) Genetic population structure of bull trout from the Flathead River basin as shown by microsatellites and mitochondrial DNA markers. Trans Am Fish Soc 130:92–106
Kanno Y, Vokoun JC, Letcher BH (2011) Fine-scale population structure and riverscape genetics of brook trout (Salvelinus fontinalis) distributed continuously along headwater channel networks. Mol Ecol 20:3711–3729
King TL, Lubinski BA, Burnham-Curtis MK, Stott W, Morgan ll RP (2012) Tools for the management and conservation of genetic diversity in brook trout (Salvelinus fontinalis): tri- and tetranucleotide microsatellite markers for the assessment of genetic diversity, phylogeography, and historical demographics. Conserv Genet Resour 4(3):539–543
Letcher BH, Nislow KH, Coombs JA, O’Donnell MJ, Dubreuil TL (2007) Population response to habitat fragmentation in a stream-dwelling brook trout population. PLoS ONE 2(11):e1139. doi:10.1371/journal.pone.0001139
McClurg SE, Petty JT, Mazik PM, Clayton JL (2007) Stream ecosystem response to limestone treatment in acid impacted watersheds of the Allegheny plateau. Ecol Appl 17:1087–1104
Meirmans PG (2006) Using the AMOVA framework to estimate a standardized genetic differentiation measure. Evolution 60:2399–2402
Neville HM, Dunham JB, Peacock MM (2006) Landscape attributes and life history shape genetic structure of trout populations in a stream network. Landsc Ecol 21:901–916
Nyce LG, Eby L, Clancy CG, Painter S, Leary RF (2013) Genetic population structure of bull trout in the East Fork Bitterroot River drainage, Montana. North Am J Fish Manage 33:432–445
Paetkau D, Calvert W, Sterling I, Strobeck C (1995) Microsatellite analysis of population structure in Canadian polar bears. Mol Ecol 4:347–354
Page RDM (2001) TreeView: Tree drawing software for Apple Macintosh and Windows. http://www.taxonomy.zoology.gla.ac.uk/rod/treeview.html. Accessed 1 June 2013
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research—an update. Bioinformatics 28:2537–2539
Petty JT, Merriam EP (2012) The Nature Education Knowledge Project: Brook trout restoration. http://www.nature.com/scitable/knowledge/library/brook-trout-restoration-83031062. Accessed 1 June 2013
Petty JT, Thorne D (2005) An ecologically based approach for identifying restoration priorities in acid impacted watersheds. Restor Ecol 13:348–357
Petty JT, Lamothe PJ, Mazik PM (2005) Spatial and seasonal dynamics of brook trout populations inhabiting a central Appalachian watershed. Trans Am Fish Soc 134:572–587
Petty JT, Hansbarger JL, Huntsman BM, Mazik PM (2012) Brook trout movement in response to temperature, flow, and thermal refugia within a complex Appalachian riverscape. Trans Am Fish Soc 141:1060–1073
Petty JT, Thorne D, Hunstman BM, Mazik PM (2014) The temperature-productivity squeeze: constraints on brook trout growth along an Appalachian river continuum. Hydrobiologia 727:151–166
Poplar-Jeffers IO, Petty JT, Anderson JT, Kite SJ, Strager MP, Fortney RH (2009) Culvert replacement and stream habitat restoration: implications from brook trout management in an Appalachian watershed, USA. Restor Ecol 17:404–413
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–949
Ramilo ST, Wang J (2012) The effect of close relatives on unsupervised Bayesian clustering algorithms in population genetic structure analysis. Mol Ecol Resour 12(3):873–884
Rannala B, Mountain JL (1997) Detecting immigration by using multilocus genotypes. Proc Natl Acad Sci USA 94:9197–9201
Raymond M, Rousset F (1995) An exact test for population differentiation. Evolution 49(6):1280–1283
Rice WR (1989) Analyzing tables of statistical tests. Evolution 43:223–225
Richards AL, King TL, Lubinski BA, Moore SE, Kulp M, Webb LS (2008) Characterization of the genetic structure among brook trout in LeConte Creek, Tennessee. Proc Annu Southeast Assoc Fish and Wildl Agencies 62:195–202
Rieman BE, Dunham JB (2000) Metapopulations and salmonids: a synthesis of life history patterns and empirical observations. Ecol Freshw Fish 9:51–64
Rosenberg NA (2004) DISTRUCT: a program for the graphical display of population structure. Mol Ecol Notes 4:137–138
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Swanberg TR (1997) Movements of and habitat use by fluvial bull trout in the Blackfoot River, Montana. Trans Am Fish Soc 126:735–746
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Vähä JP, Erkinaro J, Niemelä E, Primmer CR (2007) Life history and habitat features influence the within-river genetic structure of Atlantic salmon. Mol Ecol 16(13):2638–2654
van Oosterhout CV, Hutchinson WF, Wills DPM, Shipley P (2004) MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:535–538
Wang J, Santure AW (2009) Parentage and sibship inference from multilocus genotype data under polygamy. Genetics 181(4):1579–1594
Warnock WG, Rasmussen JB, Taylor EB (2010) Genetic clustering methods reveal bull trout (Salvelinus confluentus) fine-scale population structure as a spatially nested hierarchy. Conserv Genet 11:1421–1433
Whiteley AR, Coombs JA, Hudy M, Robinson Z, Colton AR, Nislow KH, Letcher BJ (2013) Fragmentation and patch size shape genetic structure of brook trout populations. Can J Fish Aquat Sci 70:678–688
Acknowledgments
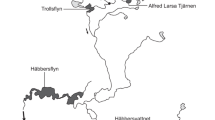
We thank the U.S. Geological Survey and the U.S. Fish and Wildlife Service for financial support for this study. We also thank Pete Lamothe, Jesse Bopp, Jason Clingerman and Jason Freund for their help with sample collection. Comments from Amy Welsh and three anonymous reviewers greatly improved earlier versions of this manuscript. Eric Merriam designed and drafted Fig. 1. We also thank Mike Shingleton and Steve Brown from the WVDNR for sharing their expertise and ideas. The use of trade names does not imply endorsement by the U.S. Government.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Aunins, A.W., Petty, J.T., King, T.L. et al. River mainstem thermal regimes influence population structuring within an appalachian brook trout population. Conserv Genet 16, 15–29 (2015). https://doi.org/10.1007/s10592-014-0636-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-014-0636-6