Abstract
Purpose
This study aimed to investigate whether genetic polymorphisms in TGFB1 contribute to breast cancer (BC) susceptibility, and explore the mechanism of action.
Methods
A total of 7 tagging SNPs (tSNPs) were genotyped in 1161 BC cases and 1337 age-matched controls among Chinese Han population. Bioinformatics analysis was used to predict functional SNP closely linked to tSNPs. Luciferase gene reporter assay was performed to determine the effect of genetic variants on promoter activity. DNA pull-down assay and mass spectrometry were used to identify the differentially binding proteins to genetic variants.
Results
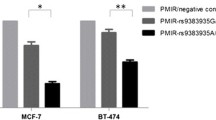
Genotyping analysis showed that rs1800469 (C>T) in the 5′ regulatory region of TGFB1 was associated with reduced BC risk. Bioinformatics analysis predicted that rs11466313 (-2389_-2391 Del/AGG) in the 5′ regulatory region of TGFB1, was closely linked to tSNP rs1800469 and could be functional. The genotyping of rs11466313 by PCR-SSCP showed that rs11466313 also conferred decreased BC risk. Luciferase assays demonstrated that rs11466313 minor allele reduced over ninefold of promoter activity compared with its major allele (p < 0.001). DNA pull-down assay and mass spectrometry revealed that rs11466313 minor allele lost the binding ability with FAM98B and HSP90B. Knocking down FAM98B but not HSP90B, the enhanced promoter activity driven by TGFB1 rs11466313 major allele was attenuated.
Conclusions
This study elucidates the impact of functional polymorphism rs11466313 in the regulatory region of TGFB1 on breast cancer susceptibility and gene expression, and could be helpful for future research to determine the value of this TGFB1 variant in the clinical setting.



Similar content being viewed by others
References
Steiner E, Klubert D, Knutson D (2008) Assessing breast cancer risk in women. Am Fam Physician 78(12):1361–1366
Scollen S, Luccarini C, Baynes C, Driver K, Humphreys MK, Garcia-Closas M, Figueroa J, Lissowska J, Pharoah PD, Easton DF, Hesketh R, Metcalfe JC, Dunning AM (2011) TGF-beta signaling pathway and breast cancer susceptibility. Cancer Epidemiol Biomarks Prev 20(6):1112–1119. https://doi.org/10.1158/1055-9965.EPI-11-0062
Hollestelle A, Wasielewski M, Martens JW, Schutte M (2010) Discovering moderate-risk breast cancer susceptibility genes. Curr Opin Genet Dev 20(3):268–276
Easton DF, Pooley KA, Dunning AM et al (2007) Genome-wide association study identifies novel breast cancer susceptibility loci. Nature 447(7148):1087–1093. https://doi.org/10.1038/nature05887
1000 Genomes Project Consortium, Abecasis GR, Altshuler D, Auton A, Brooks LD, Durbin RM, Gibbs RA, Hurles ME, McVean GA (2010) A map of human genome variation from population-scale sequencing. Nature 467(7319):1061–1073
Xu J, Yu X, Huang C, Qin R, Peng F, Lin J, Niu W (2014) Association of 5 well-defined polymorphisms in the gene encoding transforming growth factor-beta1 with coronary artery disease among Chinese patients with hypertension. Angiology. https://doi.org/10.1177/0003319714547946
Wall JD, Pritchard JK (2003) Haplotype blocks and linkage disequilibrium in the human genome. Nat Rev Genet 4(8):587–597. https://doi.org/10.1038/nrg1123
Clark AG (2004) The role of haplotypes in candidate gene studies. Genet Epidemiol 27(4):321–333
Johnson GC, Esposito L, Barratt BJ, Smith AN, Heward J, Di Genova G, Ueda H, Cordell HJ, Eaves IA, Dudbridge F, Twells RC, Payne F, Hughes W, Nutland S, Stevens H, Carr P, Tuomilehto-Wolf E, Tuomilehto J, Gough SC, Clayton DG, Todd JA (2001) Haplotype tagging for the identification of common disease genes. Nat Genet 29(2):233–237
Zu X, Zhang Q, Cao R, Liu J, Zhong J, Wen G, Cao D (2012) Transforming growth factor-beta signaling in tumor initiation, progression and therapy in breast cancer: an update. Cell Tissue Res 347(1):73–84
Zarzynska JM (2014) Two faces of TGF-beta1 in breast cancer. Mediat Inflamm 2014:141747. https://doi.org/10.1155/2014/141747
Saltzman BS, Yamamoto JF, Decker R, Yokochi L, Theriault AG, Vogt TM, Le Marchand L (2008) Association of genetic variation in the transforming growth factor beta-1 gene with serum levels and risk of colorectal neoplasia. Cancer Res 68(4):1236–1244. https://doi.org/10.1158/0008-5472.CAN-07-2144
Quan L, Gong Z, Yao S, Bandera EV, Zirpoli G, Hwang H, Roberts M, Ciupak G, Davis W, Sucheston L, Pawlish K, Bovbjerg DH, Jandorf L, Cabasag C, Coignet JG, Ambrosone CB, Hong CC (2014) Cytokine and cytokine receptor genes of the adaptive immune response are differentially associated with breast cancer risk in American women of African and European ancestry. Int J Cancer 134(6):1408–1421. https://doi.org/10.1002/ijc.28458
Akinyi MV, Dandara C, Gamieldien J, Heckmann JM (2012) Association of transforming growth factor beta-1 (TGFB1) regulatory region polymorphisms with myasthenia gravis-related ophthalmoparesis. J Neuroimmunol 246(1–2):96–99. https://doi.org/10.1016/j.jneuroim.2012.03.002
Ma X, Chen C, Xiong H, Li Y (2010) Transforming growth factorbeta1 L10P variant plays an active role on the breast cancer susceptibility in Caucasian: evidence from 10,392 cases and 11,697 controls. Breast Cancer Res Treat 124(2):453–457. https://doi.org/10.1007/s10549-010-0843-x
Broeks A, Schmidt MK, Sherman ME et al (2011) Low penetrance breast cancer susceptibility loci are associated with specific breast tumor subtypes: findings from the Breast Cancer Association Consortium. Hum Mol Genet 20(16):3289–3303. https://doi.org/10.1093/hmg/ddr228
Woo SU, Park KH, Woo OH, Yang DS, Kim AR, Lee ES, Lee JB, Kim YH, Kim JS, Seo JH (2010) Association of a TGF-beta1 gene -509 C/T polymorphism with breast cancer risk: a meta-analysis. Breast Cancer Res Treat 124(2):481–485. https://doi.org/10.1007/s10549-010-0871-6
Huang Y, Hao Y, Li B, Xie J, Qian J, Chao C, Yu L (2011) Lack of significant association between TGF-beta1-590C/T polymorphism and breast cancer risk: a meta-analysis. Med Oncol 28(2):424–428. https://doi.org/10.1007/s12032-010-9491-6
Shah R, Hurley CK, Posch PE (2006) A molecular mechanism for the differential regulation of TGF-beta1 expression due to the common SNP -509C-T (c. -1347C>T). Hum Genet 120(4):461–469. https://doi.org/10.1007/s00439-006-0194-1
Yang L, Wang YJ, Zheng LY, Jia YM, Chen YL, Chen L, Liu DG, Li XH, Guo HY, Sun YL, Tian XX, Fang WG (2016) Genetic polymorphisms of TGFB1, TGFBR1, SNAI1 and TWIST1 are associated with endometrial cancer susceptibility in Chinese Han Women. PLoS One 11(5):e0155270. https://doi.org/10.1371/journal.pone.0155270
Yamashita K, Tatebayashi T, Shinoda H, Okayasu I (1996) Simplified rapid non-radioactive PCR-SSCP method applied to K-ras mutation analysis. Pathol Int 46(10):801–804. https://doi.org/10.1111/j.1440-1827.1996.tb03553.x
Lewis CM (2002) Genetic association studies: design, analysis and interpretation. Brief Bioinform 3(2):146–153
Ruan Y, Song AP, Wang H, Xie YT, Han JY, Sajdik C, Tian XX, Fang WG (2011) Genetic polymorphisms in AURKA and BRCA1 are associated with breast cancer susceptibility in a Chinese Han population. J Pathol 225(4):535–543. https://doi.org/10.1002/path.2902
Ding SL, Yu JC, Chen ST, Hsu GC, Kuo SJ, Lin YH, Wu PE, Shen CY (2009) Genetic variants of BLM interact with RAD51 to increase breast cancer susceptibility. Carcinogenesis 30(1):43–49. https://doi.org/10.1093/carcin/bgn233
Han JY, Wang H, Xie YT, Li Y, Zheng LY, Ruan Y, Song AP, Tian XX, Fang WG (2012) Association of germline variation in CCNE1 and CDK2 with breast cancer risk, progression and survival among Chinese Han women. PLoS One 7(11):e49296. https://doi.org/10.1371/journal.pone.0049296
Soldner F, Stelzer Y, Shivalila CS, Abraham BJ, Latourelle JC, Barrasa MI, Goldmann J, Myers RH, Young RA, Jaenisch R (2016) Parkinson-associated risk variant in distal enhancer of alpha-synuclein modulates target gene expression. Nature 533(7601):95–99. https://doi.org/10.1038/nature17939
Healy J, Dionne J, Belanger H, Lariviere M, Beaulieu P, Labuda D, Sinnett D (2009) Functional impact of sequence variation in the promoter region of TGFB1. Int J Cancer 125(6):1483–1489. https://doi.org/10.1002/ijc.24526
Ostrovsky O, Grushchenko-Polaq AH, Beider K, Mayorov M, Canaani J, Shimoni A, Vlodavsky I, Nagler A (2018) Identification of strong intron enhancer in the heparanase gene: effect of functional rs4693608 variant on HPSE enhancer activity in hematological and solid malignancies. Oncogenesis 7(6):51. https://doi.org/10.1038/s41389-018-0060-8
Li WH, Qiu Y, Zhang HQ, Liu Y, You JF, Tian XX, Fang WG (2013) P2Y2 receptor promotes cell invasion and metastasis in prostate cancer cells. Br J Cancer 109(6):1666–1675. https://doi.org/10.1038/bjc.2013.484
Blobe GC, Schiemann WP, Lodish HF (2000) Role of transforming growth factor beta in human disease. N Engl J Med 342(18):1350–1358. https://doi.org/10.1056/NEJM200005043421807
Wang W, Koka V, Lan HY (2005) Transforming growth factor-beta and Smad signalling in kidney diseases. Nephrology (Carlton) 10(1):48–56. https://doi.org/10.1111/j.1440-1797.2005.00334.x
Grossberg AJ, Lei X, Xu T, Shaitelman SF, Hoffman KE, Bloom ES, Stauder MC, Tereffe W, Schlembach PJ, Woodward WA, Buchholz TA, Smith BD (2018) Association of transforming growth factor beta polymorphism C-509T with radiation-induced fibrosis among patients with early-stage breast cancer: a secondary analysis of a randomized clinical trial. JAMA Oncol 4(12):1751–1757. https://doi.org/10.1001/jamaoncol.2018.2583
Cox DG, Penney K, Guo Q, Hankinson SE, Hunter DJ (2007) TGFB1 and TGFBR1 polymorphisms and breast cancer risk in the Nurses’ Health Study. BMC Cancer 7:175. https://doi.org/10.1186/1471-2407-7-175
Dunning AM, Ellis PD, McBride S, Kirschenlohr HL, Healey CS, Kemp PR, Luben RN, Chang-Claude J, Mannermaa A, Kataja V, Pharoah PD, Easton DF, Ponder BA, Metcalfe JC (2003) A transforming growth factorbeta1 signal peptide variant increases secretion in vitro and is associated with increased incidence of invasive breast cancer. Cancer Res 63(10):2610–2615
Yang J, Zhong T, Xiao G, Chen Y, Liu J, Xia C, Du H, Kang X, Lin Y, Guan R, Yan P, Xiao J (2015) Polymorphisms and haplotypes of the TGF-beta1 gene are associated with risk of polycystic ovary syndrome in Chinese Han women. Eur J Obstet Gynecol Reprod Biol 186:1–7. https://doi.org/10.1016/j.ejogrb.2014.11.004
Healy J, Bourgey M, Richer C, Sinnett D, Roy-Gagnon MH (2010) Detection of fetomaternal genotype associations in early-onset disorders: evaluation of different methods and their application to childhood leukemia. J Biomed Biotechnol 2010:369534. https://doi.org/10.1155/2010/369534
Grainger DJ, Heathcote K, Chiano M, Snieder H, Kemp PR, Metcalfe JC, Carter ND, Spector TD (1999) Genetic control of the circulating concentration of transforming growth factor type beta1. Hum Mol Genet 8(1):93–97. https://doi.org/10.1093/hmg/8.1.93
Luedecking EK, DeKosky ST, Mehdi H, Ganguli M, Kamboh MI (2000) Analysis of genetic polymorphisms in the transforming growth factor-beta1 gene and the risk of Alzheimer’s disease. Hum Genet 106(5):565–569. https://doi.org/10.1007/s004390000313
Akter KA, Mansour MA, Hyodo T, Senga T (2017) FAM98A associates with DDX1-C14orf166-FAM98B in a novel complex involved in colorectal cancer progression. Int J Biochem Cell Biol 84:1–13. https://doi.org/10.1016/j.biocel.2016.12.013
Pazo A, Perez-Gonzalez A, Oliveros JC, Huarte M, Chavez JP, Nieto A (2019) hCLE/RTRAF-HSPC117-DDX1-FAM98B: a new cap-binding complex that activates mRNA translation. Front Physiol 10:92. https://doi.org/10.3389/fphys.2019.00092
Tanaka K, Ikeda N, Miyashita K, Nuriya H, Hara T (2018) DEAD box protein DDX1 promotes colorectal tumorigenesis through transcriptional activation of the LGR5 gene. Cancer Sci 109(8):2479–2489. https://doi.org/10.1111/cas.13661
Funding
This study was funded by the National Natural Science Foundation of China (Nos. 81872382, 81672790 and 81621063).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
All the authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in studies involving human participants were approved by the Ethics Committee of Peking University Health Science Center and were in accordance with the 1964 Helsinki declaration.
Informed consent
Informed consent was obtained from all individual participants included in the study. The Peking University IRB (reference No. IRB00001052-11029) approved this study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhou, YT., Zheng, LY., Wang, YJ. et al. Effect of functional variant rs11466313 on breast cancer susceptibility and TGFB1 promoter activity. Breast Cancer Res Treat 184, 237–248 (2020). https://doi.org/10.1007/s10549-020-05841-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-020-05841-w