Abstract
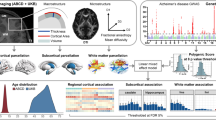
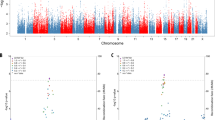
Genetic risk for Late Onset Alzheimer Disease (AD) has been associated with lower cognition and smaller hippocampal volume in healthy young adults. However, whether these and other associations are present during childhood remains unclear. Using data from 5556 genomically-confirmed European ancestry youth who completed the baseline session of the ongoing the Adolescent Brain Cognitive DevelopmentSM Study (ABCD Study®), our phenome-wide association study estimating associations between four indices of genetic risk for late-onset AD (i.e., AD polygenic risk scores (PRS), APOE rs429358 genotype, AD PRS with the APOE region removed (ADPRS-APOE), and an interaction between ADPRS-APOE and APOE genotype) and 1687 psychosocial, behavioral, and neural phenotypes revealed no significant associations after correction for multiple testing (all ps > 0.0002; all pfdr > 0.07). These data suggest that AD genetic risk may not phenotypically manifest during middle-childhood or that effects are smaller than this sample is powered to detect.

Similar content being viewed by others
Data availability
All ABCD data used in this study are available through the National Institute of Mental Health Data Archive (NDA), which may be accessed here: https://nda.nih.gov/.
Code availability
References
Albert MS, DeKosky ST, Dickson D et al (2011) The diagnosis of mild cognitive impairment due to Alzheimer’s disease: recommendations from the National Institute on Aging-Alzheimer’s Association workgroups on diagnostic guidelines for Alzheimer’s disease. Alzheimers Dement 7:270–279. https://doi.org/10.1016/j.jalz.2011.03.008
Alexander S, Kerr ME, Kim Y et al (2007) Apolipoprotein E4 allele presence and functional outcome after severe traumatic brain injury. J Neurotrauma 24:790–797. https://doi.org/10.1089/neu.2006.0133
Austin PC (2010) Estimating multilevel logistic regression models when the number of clusters is low: a comparison of different statistical software procedures. Int J Biostat 6:16. https://doi.org/10.2202/1557-4679.1195
Bartoń K (2009) MuMIn: multi-model inference
Basser PJ, Mattiello J, Lebihan D (1994) Estimation of the effective self-diffusion tensor from the NMR spin echo. J Magn Reson Ser B 103:247–254. https://doi.org/10.1006/jmrb.1994.1037
Bates D, Mächler M, Bolker B, Walker S (2015) Fitting linear mixed-effects models using lme4. J Stat Softw 67:1–48. https://doi.org/10.18637/jss.v067.i01
Baurley JW, Edlund CK, Pardamean CI et al (2016) Smokescreen: a targeted genotyping array for addiction research. BMC Genomics 17:145. https://doi.org/10.1186/s12864-016-2495-7
Bekris LM, Yu C-E, Bird TD, Tsuang DW (2010) Genetics of Alzheimer disease. J Geriatr Psychiatry Neurol 23:213–227. https://doi.org/10.1177/0891988710383571
Bellenguez C, Küçükali F, Jansen IE et al (2022) New insights into the genetic etiology of Alzheimer’s disease and related dementias. Nat Genet 54:412–436. https://doi.org/10.1038/s41588-022-01024-z
Cruchaga C, Kauwe JSK, Nowotny P et al (2012) Cerebrospinal fluid APOE levels: an endophenotype for genetic studies for Alzheimer’s disease. Hum Mol Genet 21:4558–4571. https://doi.org/10.1093/hmg/dds296
Dale AM, Fischl B, Sereno MI (1999) Cortical surface-based analysis I. Segmentation and surface reconstruction. Neuroimage 9:179–194. https://doi.org/10.1006/nimg.1998.0395
Dean DC, Jerskey BA, Chen K et al (2014) Brain differences in infants at differential genetic risk for late-onset Alzheimer disease: a cross-sectional imaging study. JAMA Neurol 71:11–22. https://doi.org/10.1001/jamaneurol.2013.4544
Desikan RS, Ségonne F, Fischl B et al (2006) An automated labeling system for subdividing the human cerebral cortex on MRI scans into gyral based regions of interest. Neuroimage 31:968–980. https://doi.org/10.1016/j.neuroimage.2006.01.021
Dondu A, Sevincoka L, Akyol A, Tataroglu C (2015) Is obsessive-compulsive symptomatology a risk factor for Alzheimer-type dementia? Psychiatry Res 225:381–386. https://doi.org/10.1016/j.psychres.2014.12.010
Elliott ML, Knodt AR, Ireland D et al (2020) What is the test-retest reliability of common task-functional MRI measures? New empirical evidence and a meta-analysis. Psychol Sci 31:792–806. https://doi.org/10.1177/0956797620916786
Escott-Price V, Hardy J (2022) Genome-wide association studies for Alzheimer’s disease: bigger is not always better. Brain Commun 4:fcac125. https://doi.org/10.1093/braincomms/fcac125
Evans SL, Dowell NG, Prowse F et al (2020) Mid age APOE ε4 carriers show memory-related functional differences and disrupted structure-function relationships in hippocampal regions. Sci Rep 10:3110. https://doi.org/10.1038/s41598-020-59272-0
Filippini N, MacIntosh BJ, Hough MG et al (2009) Distinct patterns of brain activity in young carriers of the APOE-epsilon4 allele. Proc Natl Acad Sci USA 106:7209–7214. https://doi.org/10.1073/pnas.0811879106
Fleisher A, Grundman M, Jack CR Jr et al (2005) Sex, apolipoprotein E ε4 status, and hippocampal volume in mild cognitive impairment. Arch Neurol 62:953–957. https://doi.org/10.1001/archneur.62.6.953
GBD 2019 Dementia Forecasting Collaborators (2022) Estimation of the global prevalence of dementia in 2019 and forecasted prevalence in 2050: an analysis for the Global Burden of Disease Study 2019. Lancet Public Health 7:e105–e125.https://doi.org/10.1016/S2468-2667(21)00249-8
Ge T, Chen C-Y, Ni Y et al (2019) Polygenic prediction via Bayesian regression and continuous shrinkage priors. Nat Commun 10:1776. https://doi.org/10.1038/s41467-019-09718-5
Gellersen HM, Guell X, Sami S (2021) Differential vulnerability of the cerebellum in healthy ageing and Alzheimer’s disease. NeuroImage 30:102605. https://doi.org/10.1016/j.nicl.2021.102605
Ghassabian A, Sundaram R, Bell E et al (2016) Gross motor milestones and subsequent development. Pediatrics 138:e20154372. https://doi.org/10.1542/peds.2015-4372
Gordon EM, Laumann TO, Adeyemo B et al (2016) Generation and evaluation of a cortical area parcellation from resting-state correlations. Cereb Cortex 26:288–303. https://doi.org/10.1093/cercor/bhu239
Grabher BJ (2018) Effects of Alzheimer disease on patients and their family. J Nucl Med Technol 46:335–340. https://doi.org/10.2967/jnmt.118.218057
Graham A, Livingston G, Purnell L, Huntley J (2022) Mild traumatic brain injuries and future risk of developing Alzheimer’s disease: systematic review and meta-analysis. J Alzheimers Dis 87:969–979. https://doi.org/10.3233/JAD-220069
Green P, MacLeod CJ (2016) SIMR: an R package for power analysis of generalized linear mixed models by simulation. Methods Ecol Evol 7:493–498. https://doi.org/10.1111/2041-210X.12504
Hendriks S, Peetoom K, Bakker C et al (2021) Global prevalence of young-onset dementia: a systematic review and meta-analysis. JAMA Neurol 78:1080–1090. https://doi.org/10.1001/jamaneurol.2021.2161
Hong EP, Park JW (2012) Sample size and statistical power calculation in genetic association studies. Genomics Inf 10:117–122. https://doi.org/10.5808/GI.2012.10.2.117
Jacobs HIL, Hopkins DA, Mayrhofer HC et al (2018) The cerebellum in Alzheimer’s disease: evaluating its role in cognitive decline. Brain J Neurol 141:37–47. https://doi.org/10.1093/brain/awx194
Johnson EC, Demontis D, Thorgeirsson TE et al (2020) A large-scale genome-wide association study meta-analysis of cannabis use disorder. Lancet Psychiatry 7:1032–1045. https://doi.org/10.1016/S2215-0366(20)30339-4
Joo YY, Moon S-Y, Wang H-H et al (2022) Association of genome-wide polygenic scores for multiple psychiatric and common traits in preadolescent youths at risk of suicide. JAMA Network Open 5:e2148585. https://doi.org/10.1001/jamanetworkopen.2021.48585
Korologou-Linden R, Bhatta L, Brumpton BM et al (2022) The causes and consequences of Alzheimer’s disease: phenome-wide evidence from Mendelian randomization. Nat Commun 13:4726. https://doi.org/10.1038/s41467-022-32183-6
Karcher NR, Paul SE, Johnson EC et al (2022) Psychotic-like experiences and polygenic liability in the adolescent brain cognitive development study. Biol Psychiatry Cognit Neurosci Neuroimaging 7:45–55. https://doi.org/10.1016/j.bpsc.2021.06.012
Kunkle BW, Grenier-Boley B, Sims R et al (2019) Genetic meta-analysis of diagnosed Alzheimer’s disease identifies new risk loci and implicates Aβ, tau, immunity and lipid processing. Nat Genet 51:414–430. https://doi.org/10.1038/s41588-019-0358-2
Kunkle BW, Schmidt M, Klein H-U et al (2021) Novel Alzheimer disease risk loci and pathways in African American individuals using the African genome resources panel: a meta-analysis. JAMA Neurol 78:102–113. https://doi.org/10.1001/jamaneurol.2020.3536
Ladouceur CD, Kerestes R, Schlund MW et al (2019) Neural systems underlying reward cue processing in early adolescence: the role of puberty and pubertal hormones. Psychoneuroendocrinology 102:281–291. https://doi.org/10.1016/j.psyneuen.2018.12.016
Lam M, Awasthi S, Watson HJ et al (2020) RICOPILI: rapid imputation for COnsortias PIpeLIne. Bioinformatics 36:930–933. https://doi.org/10.1093/bioinformatics/btz633
Liu JZ, Erlich Y, Pickrell JK (2017) Case-control association mapping by proxy using family history of disease. Nat Genet 49:325–331. https://doi.org/10.1038/ng.3766
Livingston G, Huntley J, Sommerlad A et al (2020) Dementia prevention, intervention, and care: 2020 report of the Lancet Commission. Lancet 396:413–446. https://doi.org/10.1016/S0140-6736(20)30367-6
Martin AR, Kanai M, Kamatani Y et al (2019) Clinical use of current polygenic risk scores may exacerbate health disparities. Nat Genet 51:584–591. https://doi.org/10.1038/s41588-019-0379-x
Mol MO, van der Lee SJ, Hulsman M et al (2022) Mapping the genetic landscape of early-onset Alzheimer’s disease in a cohort of 36 families. Alzheimers Res Ther 14:77. https://doi.org/10.1186/s13195-022-01018-3
Muir AM, Ching C, Santhalingam V et al (2021) The relationship between APOE genotype and subcortical volume: a UK Biobank study (N=36,920). Alzheimer’s Dementia 17:e055650. https://doi.org/10.1002/alz.055650
Murray GK, Jones PB, Kuh D, Richards M (2007) Infant developmental milestones and subsequent cognitive function. Ann Neurol 62:128–136. https://doi.org/10.1002/ana.21120
Murray AN, Chandler HL, Lancaster TM (2021) Multimodal hippocampal and amygdala subfield volumetry in polygenic risk for Alzheimer’s disease. Neurobiol Aging 98:33–41. https://doi.org/10.1016/j.neurobiolaging.2020.08.022
Nagelkerke NJD Miscellanea A note on a general definition of the coefficient of determination
Nichols E, Szoeke CEI, Vollset SE et al (2019) Global, regional, and national burden of Alzheimer’s disease and other dementias, 1990–2016: a systematic analysis for the Global Burden of Disease Study 2016. Lancet Neurol 18:88–106. https://doi.org/10.1016/S1474-4422(18)30403-4
Nie X, Sun Y, Wan S et al (2017) Subregional structural alterations in hippocampus and nucleus accumbens correlate with the clinical impairment in patients with Alzheimer’s disease clinical spectrum: parallel combining volume and vertex-based approach. Front Neurol 8:399. https://doi.org/10.3389/fneur.2017.00399
O’Dwyer L, Lamberton F, Matura S et al (2012) Reduced hippocampal volume in healthy young ApoE4 carriers: an MRI study. PLoS ONE 7:e48895. https://doi.org/10.1371/journal.pone.0048895
Ohi K, Ochi R, Noda Y et al (2021) Polygenic risk scores for major psychiatric and neurodevelopmental disorders contribute to sleep disturbance in childhood: Adolescent Brain Cognitive Development (ABCD) Study. Transl Psychiatry 11:1–11. https://doi.org/10.1038/s41398-021-01308-8
Paul SE, Hatoum AS, Fine JD et al (2021) Associations between prenatal cannabis exposure and childhood outcomes: results from the ABCD study. JAMA Psychiatry 78:64–76. https://doi.org/10.1001/jamapsychiatry.2020.2902
Reitz C, Rogaeva E, Beecham GW (2020) Late-onset vs nonmendelian early-onset Alzheimer disease: a distinction without a difference? Neurol Genet 6:e512. https://doi.org/10.1212/NXG.0000000000000512
Sakai J (2020) Core Concept: How synaptic pruning shapes neural wiring during development and possibly, in disease. Proc Natl Acad Sci USA 117:16096–16099. https://doi.org/10.1073/pnas.2010281117
Sims R, Hill M, Williams J (2020) The multiplex model of the genetics of Alzheimer’s disease. Nat Neurosci 23:311–322. https://doi.org/10.1038/s41593-020-0599-5
Stafford J, Chung WT, Sommerlad A et al (2022) Psychiatric disorders and risk of subsequent dementia: systematic review and meta-analysis of longitudinal studies. Int J Geriatr Psychiatry 37:5711. https://doi.org/10.1002/gps.5711
Taliun D, Harris DN, Kessler MD et al (2021) Sequencing of 53,831 diverse genomes from the NHLBI TOPMed Program. Nature 590:290–299. https://doi.org/10.1038/s41586-021-03205-y
Tiemeier H, Lenroot RK, Greenstein DK et al (2010) Cerebellum development during childhood and adolescence: a longitudinal morphometric MRI study. Neuroimage 49:63–70. https://doi.org/10.1016/j.neuroimage.2009.08.016
Volkow ND, Koob GF, Croyle RT et al (2018) The conception of the ABCD study: from substance use to a broad NIH collaboration. Dev Cogn Neurosci 32:4–7. https://doi.org/10.1016/j.dcn.2017.10.002
Walhovd KB, Fjell AM, Sørensen Ø et al (2020) Genetic risk for Alzheimer disease predicts hippocampal volume through the human lifespan. Neurol Genet 6:e506. https://doi.org/10.1212/NXG.0000000000000506
Wang H, Abbas KM, Abbasifard M et al (2020) Global age-sex-specific fertility, mortality, healthy life expectancy (HALE), and population estimates in 204 countries and territories, 1950–2019: a comprehensive demographic analysis for the Global Burden of Disease Study 2019. The Lancet 396:1160–1203. https://doi.org/10.1016/S0140-6736(20)30977-6
Wightman DP, Jansen IE, Savage JE et al (2021) A genome-wide association study with 1,126,563 individuals identifies new risk loci for Alzheimer’s disease. Nat Genet 53:1276–1282. https://doi.org/10.1038/s41588-021-00921-z
Zhang Z, Wang M, Liu X (2022) C-reactive protein and risk of Alzheimer’s disease. Neurobiol Aging 109:259–263. https://doi.org/10.1016/j.neurobiolaging.2021.08.010
Acknowledgements
We are thankful to families who have participated in the ABCD Study as well as study staff and investigators. We thank Carlos Cruchaga for guidance on APOE genotype coding.
Funding
This study was funded by R01DA054750 (RB, AA). AJG was supported by NSF DGE-213989. SEP was supported by F31AA029934. NRK was supported by K23MH12179201. ECJ was supported by K01DA051759. ASH was supported by K01AA030083. Data for this study were provided by the Adolescent Brain Cognitive Development (ABCD) study, which was funded by the National Institutes of Health (grants U01DA041022, U01DA041025, U01DA041028, U01DA041048, U01DA041089, U01DA041093, U01DA041106, U01DA041117, U01DA041120, U01DA041134, U01DA041148, U01DA041156, U01DA041174, U24DA041123, and U24DA041147) and additional federal partners (https://abcdstudy.org/federal-partners.html).
Author information
Authors and Affiliations
Contributions
AJG, SEP, NRK, IN, LB, ISH cleaned the phenotypic data. AJG, SEP, ECJ, SC, ASH cleaned the genomic data and calculated/coded genetic risk (i.e., PRS, APOE4). AJG, SEP, ASH conducted analyses. AJG and RB conceptualized the study and AJG, RB, ASH, ECJ, NRK, and SEP drafted the initial manuscript. All coauthors provided input on study conceptualization and edited the manuscript with important intellectual content.
Corresponding author
Ethics declarations
Conflict of interest
Aaron J. Gorelik, Sarah E. Paul, Nicole R. Karcher, Emma C. Johnson, Isha Nagella, Lauren Blaydon, Hailey Modi, Isabella S. Hansen, Sarah M.C. Colbert, David A.A. Baranger, Sara A. Norton, Isaiah Spears, Brian Gordon, Wei Zhang, Patrick L. Hill, Thomas F. Oltmanns, Janine D. Bjisterbosch, Arpana Agrawal, Alexander S. Hatoum, and Ryan Bogdan declare that they have no conflict of interest.
Ethical approval
Working with ABCD NDA data was approved by the Washington University in St. Louis Institutional Review Board: IRB ID#201708123.
Human and Animal Rights and Informed consent
All data was obtained through the Adolescent Brain and Cognitive Development study. Informed consent was handled by each site. The ABCD data is publicly available (secondary data analysis). Informed consent was obtained from each site before data collection from the Childs parent or guardian. Data was deanonymized before download via the public NDA App. ABCD has extensive protocols for participant consent, safety, and anonymity, see https://doi.org/10.1016/j.dcn.2017.06.005.
Additional information
Handling Editor: Chun Chieh Fan.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gorelik, A.J., Paul, S.E., Karcher, N.R. et al. A Phenome-Wide Association Study (PheWAS) of Late Onset Alzheimer Disease Genetic Risk in Children of European Ancestry at Middle Childhood: Results from the ABCD Study. Behav Genet 53, 249–264 (2023). https://doi.org/10.1007/s10519-023-10140-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10519-023-10140-3