Abstract
Background
Gastric cancer (GC) is characterized by an immunosuppressive and treatment-resistant tumor immune microenvironment (TIME). Here, we investigated the roles of different immunosuppressive cell types in the development of the GC TIME.
Methods
Single-cell RNA sequencing (scRNA-seq) and multiplex immunostaining of samples from untreated or immune checkpoint inhibitor (ICI)-resistant GC patients were used to examine the correlation between certain immunosuppressive cells and the prognosis of GC patients.
Results
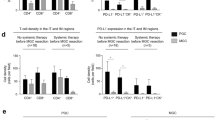
The results of the scRNA-seq analysis revealed that tumor-infiltrating monocytic myeloid-derived suppressor cells (TI-M-MDSCs) expressed higher levels of genes with immunosuppressive functions than other immunosuppressive cell types. Additionally, M-MDSCs in GC tissues expressed significantly higher levels of these markers than adjacent normal tissues. The M-MDSCs were most enriched in GC tissues relative to adjacent normal tissues. Among the immunosuppressive cell types assessed, the M-MDSCs were most enriched in GC tissues relative to adjacent normal tissues; moreover, their presence was most strongly associated with a poor prognosis. Immediate early response 3 (IER3), which we identified as a differentially expressed gene between M-MDSCs of GC and adjacent normal tissues, was an independent poor prognostic factor in GC patients (P = 0.0003). IER3+ M-MDSCs expressed higher levels of genes with immunosuppressive functions than IER3− M-MDSCs and were abundant in treatment-resistant GC patients.
Conclusions
The present study suggests that TI-M-MDSCs, especially IER3+ ones, may play a predominant role in the development of the immunosuppressive and ICI-resistant GC TIME.





Similar content being viewed by others
Data availability and materials
The scRNA-seq data generated in this study are available from the GEO database under accession code GSE231540. The remaining data are available within the manuscript, Supplementary Information, or from the corresponding author upon reasonable request.
References
Shitara K, Özgüroğlu M, Bang YJ, Di Bartolomeo M, Mandalà M, Ryu MH, et al. Pembrolizumab versus paclitaxel for previously treated, advanced gastric or gastro-oesophageal junction cancer (KEYNOTE-061): a randomised, open-label, controlled, phase 3 trial. Lancet. 2018;392:123–33.
Kang YK, Boku N, Satoh T, Ryu MH, Chao Y, Kato K, et al. Nivolumab in patients with advanced gastric or gastro-oesophageal junction cancer refractory to, or intolerant of, at least two previous chemotherapy regimens (ONO-4538-12, ATTRACTION-2): a randomised, double-blind, placebo-controlled, phase 3 trial. Lancet. 2017;390:2461–71. https://doi.org/10.1016/S0140-6736(17)31827-5.
Eum HH, Kwon M, Ryu D, Jo A, Chung W, Kim N, et al. Tumor-promoting macrophages prevail in malignant ascites of advanced gastric cancer. Exp Mol Med. 2020;52:1976–88. https://doi.org/10.1038/s12276-020-00538-y.
Sathe A, Grimes SM, Lau BT, Chen J, Suarez C, Huang RJ, Poultsides GJH. Single-cell genomic characterization reveals the cellular reprogramming of the gastric tumor microenvironment. Clin Cancer Res. 2020;26:2640–53.
Fu K, Hui B, Wang Q, Lu C, Shi W, Zhang Z, et al. Single-cell RNA sequencing of immune cells in gastric cancer patients. Aging. 2020;12:2747–63.
Joyce JA, Pollard JW. Microenvironmental regulation of metastasis. Nat Rev Cancer. 2009;9:239–52.
Movahedi K, Guilliams M, Van Den Bossche J, Van Den Bergh R, Gysemans C, Beschin A, et al. Identification of discrete tumor-induced myeloid-derived suppressor cell subpopulations with distinct T cell suppressive activity. Blood. 2008;111:4233–44.
Dolcetti L, Peranzoni E, Ugel S, Marigo I, Gomez AF, Mesa C, et al. Hierarchy of immunosuppressive strength among myeloid-derived suppressor cell subsets is determined by GM-CSF. Eur J Immunol. 2010;40:22–35.
Bronte V, Brandau S, Chen SH, Colombo MP, Frey AB, Greten TF, et al. Recommendations for myeloid-derived suppressor cell nomenclature and characterization standards. Nat Commun. 2016;7:1–10. https://doi.org/10.1038/ncomms12150.
Veglia F, Perego M, Gabrilovich D. Myeloid-derived suppressor cells coming of age review-article. Nat Immunol. 2018;19:108–19.
Corzo CA, Condamine T, Lu L, Cotter MJ, Youn JI, Cheng P, et al. HIF-1α regulates function and differentiation of myeloid-derived suppressor cells in the tumor microenvironment. J Exp Med. 2010;207:2439–53.
Koh J, Hur JY, Lee KY, Kim MS, Heo JY, Ku BM, et al. Regulatory (FoxP3+) T cells and TGF-β predict the response to anti-PD-1 immunotherapy in patients with non-small cell lung cancer. Sci Rep. 2020. https://doi.org/10.1038/s41598-020-76130-1.
Filipazzi P, Valenti R, Huber V, Pilla L, Canese P, Iero M, et al. Identification of a new subset of myeloid suppressor cells in peripheral blood of melanoma patients with modulation by a granulocyte-macrophage colony-stimulation factor-based antitumor vaccine. J Clin Oncol. 2007;25:2546–53.
Cole KE, Ly QP, Hollingsworth MA, Cox JL, Padussis JC, Foster JM, et al. Human splenic myeloid derived suppressor cells: phenotypic and clustering analysis. Cell Immunol. 2021;363:104317.
Nefedova Y, Fishman M, Sherman S, Wang X, Beg AA, Gabrilovich DI. Mechanism of all-trans retinoic acid effect on tumor-associated myeloid-derived suppressor cells. Cancer Res. 2007;67:11021–8.
Kodumudi KN, Woan K, Gilvary DL, Sahakian E, Wei S, Djeu JY. A novel chemoimmunomodulating property of docetaxel: Suppression of myeloid-derived suppressor cells in tumor bearers. Clin Cancer Res. 2010;16:4583–94.
Diaz-Montero CM, Salem ML, Nishimura MI, Garrett-Mayer E, Cole DJ, Montero AJ. Increased circulating myeloid-derived suppressor cells correlate with clinical cancer stage, metastatic tumor burden, and doxorubicin-cyclophosphamide chemotherapy. Cancer Immunol Immunother. 2009;58:49–59.
Nestorowa S, Hamey FK, Sala BP, Diamanti E, Shepherd M, Laurenti E, et al. Plenary Paper HEMATOPOIESIS AND STEM CELLS e-Blood A single-cell resolution map of mouse hematopoietic stem and progenitor cell differentiation. 2016; Available from: https://www.r-project.org. Accessed 1 Dec 2023.
Hafemeister C, Satija R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 2019;20:296. https://doi.org/10.1101/576827.
McGinnis CS, Murrow LM, Gartner ZJ. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 2019;8:329-337.e4.
Browaeys R, Saelens W, Saeys Y. NicheNet: modeling intercellular communication by linking ligands to target genes. Nat Methods. 2020;17:159–62. https://doi.org/10.1038/s41592-019-0667-5.
Zhou Y, Zhou B, Pache L, Chang M, Khodabakhshi AH, Tanaseichuk O, et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat Commun. 2019. https://doi.org/10.1038/s41467-019-09234-6.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102:15545–50.
Tang Z, Kang B, Li C, Chen T, Zhang Z. GEPIA2: an enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res. 2019;47:W556–60.
Alshetaiwi H, Pervolarakis N, Mcintyre LL, Ma D, Rath JA, Nee K, et al. Defining the emergence of myeloid-derived suppressor cells in breast cancer using single-cell transcriptomics. Sci Immunol. 2020;5:eaay6017.
Veglia F, Sanseviero E, Gabrilovich DI. Myeloid-derived suppressor cells in the era of increasing myeloid cell diversity. Nat Rev Immunol. 2021;21:485–98. https://doi.org/10.1038/s41577-020-00490-y.
Obradovic A, Graves D, Korrer M, Wang Y, Roy S, Naveed A, et al. Immunostimulatory cancer-associated fibroblast subpopulations can predict immunotherapy response in head and neck cancer. Clin Cancer Res. 2022;28:2094–109.
Togashi Y, Shitara K, Nishikawa H. Regulatory T cells in cancer immunosuppression—implications for anticancer therapy. Nat Rev Clin Oncol. 2019;16:356–71. https://doi.org/10.1038/s41571-019-0175-7.
Kumar V, Ramnarayanan K, Sundar R, Padmanabhan N, Srivastava S, Koiwa M, et al. Single-cell atlas of lineage states, tumor microenvironment, and subtype-specific expression programs in gastric cancer. Cancer Discov. 2022;12:670–91.
Lee HO, Hong Y, Etlioglu HE, Cho YB, Pomella V, Van den Bosch B, et al. Lineage-dependent gene expression programs influence the immune landscape of colorectal cancer. Nat Genet. 2020;52:594–603. https://doi.org/10.1038/s41588-020-0636-z.
Farshidpour M, Ahmed M, Junna S, Merchant JL. Myeloid-derived suppressor cells in gastrointestinal cancers: A systemic review. World J Gastrointest Oncol. 2021;13:1–11.
Xiao F, Dai Y, Hu Y, Lu M, Dai Q. Expression profile analysis identifies IER3 to predict overall survival and promote lymph node metastasis in tongue cancer. Cancer Cell Int. 2019;19:1–12. https://doi.org/10.1186/s12935-019-1028-2.
Witcher M, Pettersson F, Dupéré-richer D, Padovani A, Summers-deluca L, Baldwin AS, et al. Retinoic acid modulates chromatin to potentiate tumor necrosis factor alpha signaling on the DIF2 promoter. Nucleic Acids Res. 2008;36:435–43.
Ostrand-Rosenberg S, Sinha P, Beury DW, Clements VK. Cross-talk between myeloid-derived suppressor cells (MDSC), macrophages, and dendritic cells enhances tumor-induced immune suppression. Semin Cancer Biol. 2012;25:275–81.
Weber R, Fleming V, Hu X, Nagibin V, Groth C, Altevogt P, et al. Myeloid-derived suppressor cells hinder the anti-cancer activity of immune checkpoint inhibitors. Front Immunol. 2018;9:1310. https://doi.org/10.3389/fimmu.2018.01310.
Kim W, Chu TH, Nienhüser H, Jiang Z, Del Portillo A, Remotti HE, et al. PD-1 signaling promotes tumor-infiltrating myeloid-derived suppressor cells and gastric tumorigenesis in mice. Gastroenterology. 2021;160:781–96.
Oliveira G, Stromhaug K, Klaeger S, Kula T, Frederick DT, Le PM, et al. Phenotype, specificity and avidity of antitumour CD8+ T cells in melanoma. Nature. 2021;596:119–25. https://doi.org/10.1038/s41586-021-03704-y.
Miller BC, Sen DR, Al Abosy R, Bi K, Virkud YV, LaFleur MW, et al. Subsets of exhausted CD8+ T cells differentially mediate tumor control and respond to checkpoint blockade. Nat Immunol. 2019;20:326–36.
Kumar V, Patel S, Tcyganov E, Gabrilovich DI. Virulence-associated characteristics of actinobacillus actinomycetemcomitans. Trends Immunol. 2016;37:208–20.
Arlt A, Schäfer H. Role of the immediate early response 3 (IER3) gene in cellular stress response, inflammation and tumorigenesis. Eur J Cell Biol. 2011;90:545–52. https://doi.org/10.1016/j.ejcb.2010.10.002.
Law AMK, Valdes-Mora F, Gallego-Ortega D. Myeloid-Derived Suppressor Cells as a Therapeutic Target for Cancer. Cells. NLM (Medline). 2020;9:561.
Zhang Y, Schlossman SF, Edwards RA, Ou CN, Gu J, Wu MX. Impaired apoptosis, extended duration of immune responses, and a lupus-like autoimmune disease in IEX-1-transgenic mice. Proc Natl Acad Sci USA. 2002;99:878–83.
Caronni N, La Terza F, Vittoria FM, Barbiera G, Mezzanzanica L, Cuzzola V, et al. IL-1β+ macrophages fuel pathogenic inflammation in pancreatic cancer. Nature. 2023;623:415–22.
Hou J, Wang X, Zhang M, Wang M, Gao P, Jiang Y. Circulating CD14 + CD163 + CD209 + M2-like monocytes are associated with the severity of infection in Helicobacter pylori-positive patients. Mol Immunol. 2019;108:13–22.
Liu J, Zhang F, Zhang Z, Zheng C. Everolimus ameliorates Helicobacter pylori infection-induced inflammation in gastric epithelial cells. Bioengineered. 2022;13:11361–72.
Acknowledgements
We are grateful to E. Manabe and S. Sadatomi (Kyushu University) for their expert technical assistance. We also thank Edanz (https://jp.edanz.com/ac) for editing a draft of this manuscript.
Funding
K. Ohuchida has received grants from JSPS KAKENHI (grant numbers JP20K17621, JP21K08800, JP21K07244, JP22H00480 and JP23H02770). C. Tsutsumi has received grants from JSPS Research Fellow DC2 (23KJ1698) and Support for Pioneering Research Initiated by the Next Generation (SPRING) “Future-Creation (MIRAI)” Course (JPMJSP2136). The other authors have nothing to disclose.
Author information
Authors and Affiliations
Contributions
CT designed and performed the experiments, analyzed the data, coordinated the research, and wrote the manuscript. KO designed the experiments, analyzed the data, wrote the manuscript, coordinated the research, and contributed to data interpretation and discussion. NK, YO, SN, and KO performed the experiments. CI, YM, NI, KN, EN, and TM contributed to data interpretation and discussion. KO, NT, KH, and KS obtained and prepared human samples. Immunohistochemically stained samples were analyzed by CT, YY, and YO. MN coordinated the research, wrote the manuscript, and contributed to data interpretation and discussion.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest.
Ethical approval and consent to participate
Clinical data, including histopathological findings, were obtained from the patients’ electronic medical records. Written informed consent was obtained from all participating patients and the study was approved by the Ethics Committee of Kyushu University (approval number: 22002-00 and 2020-7882020-503) and conducted in accordance with the Ethical Guidelines for Human Genome/Gene Research enacted by the Japanese Government and the Declaration of Helsinki.
Consent for publication
All authors have read and agreed with the manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tsutsumi, C., Ohuchida, K., Katayama, N. et al. Tumor-infiltrating monocytic myeloid-derived suppressor cells contribute to the development of an immunosuppressive tumor microenvironment in gastric cancer. Gastric Cancer 27, 248–262 (2024). https://doi.org/10.1007/s10120-023-01456-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10120-023-01456-4