Abstract
Objectives
Sjögren’s syndrome (SS), a systemic autoimmune disorder, is characterized by dry mouth and eyes. However, SS pathogenesis is poorly understood. We performed bioinformatics analysis to investigate the potential targets and molecular pathogenesis of SS.
Methods
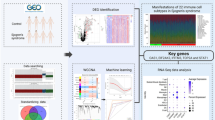
Gene expression profiles (GSE157159) and methylation data (GSE110007) associated with SS patients were obtained from the Gene Expression Omnibus (GEO) database. Differentially methylated positions (DMPs) and differentially expressed genes (DEGs) were identified by the R package limma. The potential biological functions of DEGs were determined using Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analyses. Key DMPs were selected by overlap and the shrunken centroid algorithm, and corresponding genes were identified as hub genes, with their diagnostic value assessed by receiver operating characteristic (ROC) curves. The potential molecular mechanisms of hub genes were analyzed by protein–protein interaction (PPI) networks and single-gene gene set enrichment analysis (GSEA). Peripheral blood mononuclear cells (PBMCs) were collected from control and SS patients at The Affiliated Hospital of Southwest Medical University and Dazhou Central Hospital. The mRNA levels of hub genes were verified by quantitative real-time polymerase chain reaction (qRT–PCR).
Results
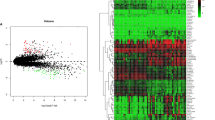
We identified 788 DMPs and 2457 DEGs between the two groups. Functional enrichment analysis suggested that the DEGs were significantly enriched in T cell activation, leukocyte cell–cell adhesion, and cytokine–cytokine receptor interaction. TSS200, TSS1500, and 1stExon were identified as highly enriched areas of differentially methylated promoter CpG islands (DMCIs). In total, 61 differentially methylated genes (DMGs) were identified by the overlap of 2457 DEGs and 507 genes related to DMPs (DMPGs), of which 21 genes located near TSS200, TSS1500, and 1stExon were selected. Then, three key DMPs and the corresponding hub genes (RUNX3, HLA-DPA1, and CD6) were screened by the shrunken centroid algorithm and calculated to have areas under the ROC curve of 1.000, 0.931, and 0.986, respectively, indicating good diagnostic value. The GSEA results suggested that all three hub genes were highly associated with the immune response. Finally, positive mRNA expression of the three hub genes in clinical SS samples was verified by qRT–PCR, consistent with the GSE157159 data.
Conclusions
The identification of three hub genes provides novel insight into molecular mechanisms and therapeutic targets for SS.
Key Points | |
• Hub genes were screened by DNA methylation and transcriptome analyses. • The relative expression of hub genes in peripheral blood samples was verified by qRT–PCR. • HLA-DPA1 was correlated with the pathogenic mechanism of SS. |






Similar content being viewed by others
Abbreviations
- ACR:
-
American College of Rheumatology
- BP:
-
biological process
- CC:
-
cellular component
- CpG:
-
cytosine-phosphate-guanine
- DEG:
-
differentially expressed gene
- DMG:
-
differentially methylation gene
- DMPG:
-
differentially methylated position gene
- EULAR:
-
European League Against Rheumatism
- GO:
-
Gene Ontology
- GEO:
-
Gene Expression Omnibus
- GSEA:
-
gene set enrichment analysis
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- MF:
-
molecular function
- NCBI:
-
National Center for Biotechnology Information
- PBMC:
-
peripheral blood mononuclear cells
- PPI:
-
protein–protein interaction
- qRT-PCR:
-
quantitative real-time polymerase chain reaction
- SS:
-
Sjögren’s syndrome
- TSS:
-
transcriptional start site
References
Psianou K, Panagoulias I, Papanastasiou AD, de Lastic AL, Rodi M, Spantidea PI et al (2018) Clinical and immunological parameters of Sjogren’s syndrome. Autoimmun Rev 17(10):1053–1064
Mariette X, Criswell LA (2018) Primary Sjögren’s syndrome. N Engl J Med 379(1):97
Qin B, Wang J, Yang Z, Yang M, Ma N, Huang F et al (2015) Epidemiology of primary Sjogren’s syndrome: a systematic review and meta-analysis. Ann Rheum Dis 74(11):1983–1989
Guellec D, Cornec D, Jousse-Joulin S, Marhadour T, Marcorelles P, Pers JO et al (2013) Diagnostic value of labial minor salivary gland biopsy for Sjogren’s syndrome: a systematic review. Autoimmun Rev 12(3):416–420
Ramos-Casals M, Brito-Zeron P, Bombardieri S, Bootsma H, De Vita S, Dorner T et al (2020) EULAR recommendations for the management of Sjogren’s syndrome with topical and systemic therapies. Ann Rheum Dis 79(1):3–18
Smith ZD, Meissner A (2013) DNA methylation: roles in mammalian development. Nat Rev Genet 14(3):204–220
Jones PA (2012) Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat Rev Genet 13(7):484–492
Bansal A, Pinney SE (2017) DNA methylation and its role in the pathogenesis of diabetes. Pediatr Diabetes 18(3):167–177
Yang IV, Lozupone CA, Schwartz DA (2017) The environment, epigenome, and asthma. J Allergy Clin Immunol 140(1):14–23
Long H, Yin H, Wang L, Gershwin ME, Lu Q (2016) The critical role of epigenetics in systemic lupus erythematosus and autoimmunity. J Autoimmun 74:118–138
Miao L, Yin R-X, Zhang Q-H, Hu X-J, Huang F, Chen W-X et al (2019) Integrated DNA methylation and gene expression analysis in the pathogenesis of coronary artery disease. Aging (Albany NY) 11(5):1486–1500
Coit P, Kaushik P, Caplan L, Kerr GS, Walsh JA, Dubreuil M et al (2019) Genome-wide DNA methylation analysis in ankylosing spondylitis identifies HLA-B*27 dependent and independent DNA methylation changes in whole blood. J Autoimmun 102:126–132
Lanata CM, Chung SA, Criswell LA (2018) DNA methylation 101: what is important to know about DNA methylation and its role in SLE risk and disease heterogeneity. Lupus Sci Med 5(1):e000285
Nakano K, Whitaker JW, Boyle DL, Wang W, Firestein GS (2013) DNA methylome signature in rheumatoid arthritis. Ann Rheum Dis 72(1):110–117
Oyelakin A, Horeth E, Song E-AC, Min S, Che M, Marzullo B et al (2020) Transcriptomic and network analysis of minor salivary glands of patients with primary Sjögren’s syndrome. Front Immunol 11:606268
Cole MB, Quach H, Quach D, Baker A, Taylor KE, Barcellos LF et al (2016) Epigenetic signatures of salivary gland inflammation in Sjogren’s syndrome. Arthritis Rheumatol 68(12):2936–2944
Cook NR (2007) Use and misuse of the receiver operating characteristic curve in risk prediction. Circulation 115(7):928–935
Shiboski CH, Shiboski SC, Seror R, Criswell LA, Labetoulle M, Lietman TM et al (2016) 2016 American College of Rheumatology/European league against rheumatism classification criteria for primary Sjögren’s syndrome: a consensus and data-driven methodology involving three international patient cohorts. Arthritis Rheumatol 69(1):35–45
Shiboski SC, Shiboski CH, Criswell LA, Baer AN, Challacombe S, Lanfranchi H et al (2012) American College of Rheumatology classification criteria for Sjögren’s syndrome: a data-driven, expert consensus approach in the Sjögren’s international collaborative clinical alliance cohort. Arthritis Care Res 64(4):475–487
Patel R, Shahane A (2014) The epidemiology of Sjogren’s syndrome. Clin Epidemiol 6:247–255
Segal B, Bowman SJ, Fox PC, Vivino FB, Murukutla N, Brodscholl J et al (2009) Primary Sjogren’s syndrome: health experiences and predictors of health quality among patients in the United States. Health Qual Life Outcomes 7:46
Mougeot JL, Noll BD, Bahrani Mougeot FK (2019) Sjogren’s syndrome X-chromosome dose effect: an epigenetic perspective. Oral Dis 25(2):372–384
Imgenberg-Kreuz J, Almlof JC, Leonard D, Sjowall C, Syvanen AC, Ronnblom L et al (2019) Shared and unique patterns of DNA methylation in systemic lupus erythematosus and primary Sjogren’s syndrome. Front Immunol 10:1686
Miceli-Richard C, Wang-Renault SF, Boudaoud S, Busato F, Lallemand C, Bethune K et al (2016) Overlap between differentially methylated DNA regions in blood B lymphocytes and genetic at-risk loci in primary Sjogren’s syndrome. Ann Rheum Dis 75(5):933–940
Altorok N, Coit P, Hughes T, Koelsch KA, Stone DU, Rasmussen A et al (2014) Genome-wide DNA methylation patterns in naive CD4+ T cells from patients with primary Sjogren’s syndrome. Arthritis Rheumatol 66(3):731–739
Konsta OD, Le Dantec C, Charras A, Cornec D, Kapsogeorgou EK, Tzioufas AG et al (2016) Defective DNA methylation in salivary gland epithelial acini from patients with Sjogren’s syndrome is associated with SSB gene expression, anti-SSB/LA detection, and lymphocyte infiltration. J Autoimmun 68:30–38
Thabet Y, Le Dantec C, Ghedira I, Devauchelle V, Cornec D, Pers JO et al (2013) Epigenetic dysregulation in salivary glands from patients with primary Sjogren’s syndrome may be ascribed to infiltrating B cells. J Autoimmun 41:175–181
Christodoulou MI, Kapsogeorgou EK, Moutsopoulos HM (2010) Characteristics of the minor salivary gland infiltrates in Sjögren’s syndrome. J Autoimmun 34(4):400–407
Du W, Han M, Zhu X, Xiao F, Huang E, Che N et al (2021) The multiple roles of B cells in the pathogenesis of Sjögren’s syndrome. Front Immunol 12:684999
Corneth OBJ, de Bruijn MJW, Rip J, Asmawidjaja PS, Kil LP, Hendriks RW (2016) Enhanced expression of Bruton’s tyrosine kinase in B cells drives systemic autoimmunity by disrupting T cell homeostasis. J Immunol 197(1):58–67
Verstappen GM, Meiners PM, Corneth OBJ, Visser A, Arends S, Abdulahad WH et al (2017) Attenuation of follicular helper T cell-dependent B cell hyperactivity by abatacept treatment in primary Sjögren’s syndrome. Arthritis Rheumatol (Hoboken, NJ) 69(9):1850–1861
Lavie F, Miceli-Richard C, Quillard J, Roux S, Leclerc P, Mariette X (2004) Expression of BAFF (BLyS) in T cells infiltrating labial salivary glands from patients with Sjögren’s syndrome. J Pathol 202(4):496–502
Roescher N, Tak PP, Illei GG (2009) Cytokines in Sjögren’s syndrome. Oral diseases 15(8):519–526
Abidi SM, Saifullah MK, Zafiropulos MD, Kaput C, Bowen MA, Cotton C et al (2006) CD166 expression, characterization, and localization in salivary epithelium: implications for function during sialoadenitis. J Clin Immunol 26(1):12–21
Ramos-Casals M, Font J, García-Carrasco M, Calvo J, Places L, Padilla O et al (2001) High circulating levels of soluble scavenger receptors (sCD5 and sCD6) in patients with primary Sjögren’s syndrome. Rheumatology (Oxford) 40(9):1056–1059
Chalmers SA, Ayilam Ramachandran R, Garcia SJ, Der E, Herlitz L, Ampudia J et al (2022) The CD6/ALCAM pathway promotes lupus nephritis via T cell-mediated responses. J Clin Invest 132(1)
Le Dantec C, Alonso R, Fali T, Montero E, Devauchelle V, Saraux A et al (2013) Rationale for treating primary Sjögren’s syndrome patients with an anti-CD6 monoclonal antibody (Itolizumab). Immunol Res 56(2-3):341–347
Soret P, Le Dantec C, Desvaux E, Foulquier N, Chassagnol B, Hubert S et al (2021) A new molecular classification to drive precision treatment strategies in primary Sjögren’s syndrome. Nat Commun 12(1):3523
Imgenberg-Kreuz J, Sandling JK, Norheim KB, Johnsen SJA, Omdal R, Syvänen A-C et al (2021) DNA methylation-based interferon scores associate with sub-phenotypes in primary Sjögren’s syndrome. Frontiers Immunol 12:702037
Djuretic IM, Levanon D, Negreanu V, Groner Y, Rao A, Ansel KM (2007) Transcription factors T-bet and Runx3 cooperate to activate Ifng and silence Il4 in T helper type 1 cells. Nat Immunol 8(2):145–153
Del Papa N, Minniti A, Lorini M, Carbonelli V, Maglione W, Pignataro F et al (2021) The role of interferons in the pathogenesis of Sjögren’s syndrome and future therapeutic perspectives. Biomolecules 11(2)
Nezos A, Gravani F, Tassidou A, Kapsogeorgou EK, Voulgarelis M, Koutsilieris M et al (2015) Type I and II interferon signatures in Sjogren’s syndrome pathogenesis: contributions in distinct clinical phenotypes and Sjogren’s related lymphomagenesis. J Autoimmun 63:47–58
Men S, Yu Y, Zhang Y, Wang Y, Qian Q, Li W et al (2019) Methylation landscape of RUNX3 promoter region as a predictive marker for Th1/Th2 imbalance in bronchiolitis. Med Sci Monit 25:7795–7807
Arman M, Aguilera-Montilla N, Mas V, Puig-Kroger A, Pignatelli M, Guigo R et al (2009) The human CD6 gene is transcriptionally regulated by RUNX and Ets transcription factors in T cells. Mol Immunol 46(11-12):2226–2235
Gunnell A, Webb HM, Wood CD, McClellan MJ, Wichaidit B, Kempkes B et al (2016) RUNX super-enhancer control through the Notch pathway by Epstein-Barr virus transcription factors regulates B cell growth. Nucleic Acids Res 44(10):4636–4650
Lauterbach N, Crivello P, Wieten L, Zito L, Groeneweg M, Voorter CE et al (2014) Allorecognition of HLA-DP by CD4+ T cells is affected by polymorphism in its alpha chain. Mol Immunol 59(1):19–29
Anczurowski M, Hirano N (2018) Mechanisms of HLA-DP antigen processing and presentation revisited. Trends Immunol 39(12):960–964
Thrane PS, Halstensen TS, Haanaes HR, Brandtzaeg P (1993) Increased epithelial expression of HLA-DQ and HLA-DP molecules in salivary glands from patients with Sjögren’s syndrome compared with obstructive sialadenitis. Clin Exp Immunol 92(2):256–262
Gupta S, Li D, Ostrov DA, Nguyen CQ (2022) Blocking IAg class II major histocompatibility complex by drug-like small molecules alleviated Sjögren’s syndrome in NOD mice. Life Sci 288:120182
Wang LY, Chen CF, Wu TW, Lai SK, Chu CC, Lin HH (2019) Response to hepatitis B vaccination is co-determined by HLA-DPA1 and -DPB1. Vaccine 37(43):6435–6440
Zhang J, Zhan W, Yang B, Tian A, Chen L, Liao Y et al (2017) Genetic polymorphisms of rs3077 and rs9277535 in HLA-DP associated with systemic lupus erythematosus in a Chinese population. Sci Rep 7:39757
Gregersen JW, Erikstrup C, Ivarsen P, Glerup R, Krarup E, Keller KK et al (2019) PR3-ANCA-associated vasculitis is associated with a specific motif in the peptide-binding cleft of HLA-DP molecules. Rheumatol (Oxford) 58(11):1942–1949
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Approved by the Ethics Committee of the affiliated Hospital of Southwest Medical University, this study obtained the consent of all the subjects and signed the informed consent form.
Disclosures
None.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information

ESM 1
Supplementary Fig. 1 The distribution of differentially methylated CpG positions (DMCPs) and the proportion of differentially methylated promoter CpG islands (DMCIs). A The pie charts illustrate the proportional distribution of DMCPs (upper panel) and DMCIs (lower panel). B Chromosomal distribution of CpG sites. (PNG 311 kb)

ESM 2
Supplementary Fig. 2 GO and KEGG enrichment analysis results for the differentially expressed genes (DEGs). A Ten terms each from the biological process, cellular component, and molecular function categories for GO enrichment based on all DEGs. B Top 10 enriched KEGG pathways for the DEGs. (PNG 184 kb)

ESM 3
Supplementary Fig. 3 Methylation levels of 7 differentially methylated CpG sites in the GSE110007 dataset. Heatmap showing the methylation of the 7 CpG sites in SS and control tissues. Red represents hypermethylation, and blue represents hypomethylation. (PNG 70 kb)

ESM 4
Supplementary Fig. 4 PPI analysis of hub genes via STRING. The interaction score was set to medium confidence (0.400). Hub genes are shown, and the thickness of the line indicates the extent of the relationship between two genes. (PNG 183 kb)
ESM 5
(XLSX 39 kb)
ESM 6
(XLSX 16 kb)
ESM 7
(XLSX 16 kb)
ESM 8
(XLSX 119 kb)
ESM 9
(XLSX 15 kb)
ESM 10
(XLSX 105 kb)
ESM 11
(XLSX 14 kb)
ESM 12
(XLSX 104 kb)
ESM 13
(XLSX 15 kb)
Rights and permissions
About this article
Cite this article
Du, Y., Li, J., Wu, J. et al. Exploration of the pathogenesis of Sjögren’s syndrome via DNA methylation and transcriptome analyses. Clin Rheumatol 41, 2765–2777 (2022). https://doi.org/10.1007/s10067-022-06200-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10067-022-06200-4