Abstract
Context
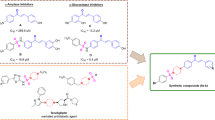
The regioselectivity and diastereoselectivity of the 1,3-dipolar cycloaddition reaction between azomethine ylides and acrolein were investigated. The DFT studies revealed that the favored pathway leads to the formation of cis-cycloadduct pyrrolidine and these computational findings align with experimental observations. The cis-cycloadduct pyrrolidine product serves as an advanced intermediate in the synthesis of a hepatitis C virus inhibitor. For this, the antiviral activity of cis-cycloadduct pyrrolidine against cyclophilin A, the co-factor responsible for hepatitis C virus, was also evaluated through molecular docking simulations which revealed intriguing interactions and a high C-score, which were further confirmed by molecular dynamics simulations, demonstrating stability over a 100-ns simulation period. Furthermore, the cis-cycloadduct pyrrolidine exhibits favorable drug-like properties and a better ADMET profile compared to hepatitis C virus inhibitor.
Methods
Chemical reactivity studies were performed using DFT method by the functional B3LYP at 6-31G (d, p) computational level by GAUSSIAN 16 program. Frontal molecular orbitals theory used to investigate HOMO/LUMO interactions between azomethine ylides and acrolein. Findings of this approach were confirmed by global reactivity indices and electron displacement was investigated based on Fukui functions. Furthermore, the activation energies were determined after frequency calculations using TS Berny algorithm and transition states were confirmed by the presence of a single imaginary frequency. Moreover, antiviral activity of cis-cycloadduct was explored through molecular docking using Surflex-Dock suite SYBYL X 2.0, and molecular dynamics simulation using GROMACS program. Finally, drug-like properties were investigated with SwissADME and ADMETlab 2.0.











Similar content being viewed by others
Data availability
All the data are available.
References
Sardari M, Fotooh FK, Nateghi MR (2018) A DFT study of the structural and electronic properties of periodic forms of aniline and pyrrole polymers and aniline–pyrrole copolymer. J Mol Model 24:148. https://doi.org/10.1007/s00894-018-3667-y
Abdessadak O, Hajji H, El Khatabi K, Mehanned S, Lakhlifi T, Ajana MA, Bouachrine M (2020) 2D-QSAR analysis on compounds (N-phenylpyrolidin-2-onesand N-phenyl-1H-pyrrol-2-ones) whose inhibitory activity of PPO (protoporphyrinogen oxidase). RHAZES: Green Appl Chem 9:92–101
Vitaku E, Smith DT, Njardarson JT (2014) Analysis of the structural diversity, substitution patterns, and frequency of nitrogen heterocycles among U.S. FDA approved. pharmaceuticals. J Med Chem 57:10257–10274. https://doi.org/10.1021/jm501100b
Giovanna Li P, Virginia S, Roberto S, Ralph H, Raimondi MV, Paola B, Montalbano A (2020) Bioactive pyrrole-based compounds with target selectivity. Eur J Med Chem 208:112–783. https://doi.org/10.1016/j.ejmech.2020.112783
Jeelan Basha N, Basavarajaiah SM, Shyamsunder K (2022) Therapeutic potential of pyrrole and pyrrolidine analogs: an update. Mol Divers. https://doi.org/10.1007/s11030-022-10387-8
Munnaluri RK, Peddi SR, Sivan SK, Manga V (2019) Computational studies on N-phenyl pyrrole derivatives as MmpL3 inhibitors in Mycobacterium tuberculosis. Comput Biol Chem 78:1476–9271. https://doi.org/10.1016/j.compbiolchem.2018.11.007
Uddin R, Jalal K, Khan K, Ul-Haq Z (2022) Re-purposing of hepatitis C virus FDA approved direct acting antivirals as potential SARS-CoV-2 protease inhibitors. J Mol Struct 15:131920. https://doi.org/10.1016/j.molstruc.2021.131920
Fischer CS, Fischer G, Braun M (2022) Non-immunosuppressive cyclophilin inhibitors. GDCh 61:e202201597. https://doi.org/10.1002/anie.202201597
André PG (2015) Roles of prolyl isomerases in RNA-mediated gene expression. Biomolecules 5:974–999
Ma Y, Peng J, Liu W, Zhang P, Huang L, Gao B, Shen T, Zhou Y, Chen H, Chu Z, Zhang M, Qin H (2009) Proteomics identification of desmin as a potential oncofetal diagnostic and prognostic biomarker in colorectal cancer. Mol Cell Proteomics 8:1878–1890. https://doi.org/10.1074/mcp.M800541-MCP200
Gallay PA (2012) Cyclophilin inhibitors: a novel class of promising host-targeting anti-HCV agents. Immunol Res 52:200–210. https://doi.org/10.1007/s12026-011-8263-5
Ugarriza I, Reyes E, Prieto L, Uria U, Carrillo L, Vicario JL (2022) An approach to the synthesis of a hepatitis C virus inhibitor through a proline-catalyzed 1,3-dipolar cycloaddition using acrolein. Synth 54:1101–1107. https://doi.org/10.1055/s-0040-1719858
Abdessadak O, Hajji H, Koubi Y, Lakhlifi T, Ajana MA, Bouachrine M (2020) Study of dipolar 1.3 cycloaddition reaction by DFT method, as well as a study of antibacterial activity of two isomers 1.4 and 1.5 on two therapeutic targets E. coli and Helicobacter pylori, by molecular docking. Turkish Comp Theo Chem 4:46–55. https://doi.org/10.33435/tcandtc.1020278
Fischer G, Gallay P, Hopkins S (2010) Cyclophilin inhibitors for the treatment of HCV infection. Curr Opin Investig Drugs 11:911–918
Jain AN (2003) Surflex: fully automatic flexible molecular docking using a molecular similarity-based search engine. J Med Chem 46:499–511
Malathi K, Ramaiah S (2016) Molecular docking and molecular dynamics studies to identify potential OXA-10 extended spectrum β-Lactamase non-hydrolysing inhibitors for Pseudomonas aeruginosa. Cell Biochem Biophys 74:141–155
Tabti K, Abdessadak O, Sbai A, Maghat H, Bouachrine M, Lakhlifi T (2023) Design and development of novel spiro-oxindoles as potent antiproliferative agents using quantitative structure activity-based Monte Carlo method, docking molecular, molecular dynamics, free energy calculations, and pharmacokinetics /toxicity studies. J Mol Struct 1284:0022–2860. https://doi.org/10.1016/j.molstruc.2023.135404
Abdessadak O, Tabti K, Alaqarbeh M, Elmchichi L, Koubi Y, Sbai A, Ajana MA, Lakhlifi T, Bouachrine M (2023) Computational studies for development of triazole-pyrimidines as inhibitor of α-tubulin receptor. ChemistrySelect 8:e202301992. https://doi.org/10.1002/slct.202301992
Abdessadak O, Hajji H, Mehanned S, Ajana MA, Lakhlifi T, Bouachrine M (2022) Reverse docking on five original PPO structures: plant, bacterial, and human. MCJ 10:442–451. https://doi.org/10.48317/IMIST.PRSM/morjchem-v10i3.33065
Abdessadak O, Alaqarbeh M, Zaki H et al (2022) Computational approaches to discover a Kaempferol derivative extracted from Senna alexandrina as Escherichia coli enzyme (MurF) inhibitor by molecular docking, molecular dynamics simulation, and ADME-Tox. Struct Chem 34:1173–1187. https://doi.org/10.1007/s11224-022-02068-x
Haroon M, Akhtar T, Yousuf M et al (2022) Synthesis, crystal structure, Hirshfeld surface investigation and comparative DFT studies of ethyl 2-[2-(2-nitrobenzylidene)hydrazinyl]thiazole-4-carboxylate. BMC Chem 16. https://doi.org/10.1186/s13065-022-00805-1
Tabti K, Sbai A, Maghat H, Lakhlifi T, Bouachrine M (2024) Computational assessment of the reactivity and pharmaceutical potential of novel triazole derivatives: an approach combining DFT calculations, molecular dynamics simulations, and molecular docking. Arab J Chem 17:105376. https://doi.org/10.1016/j.arabjc.2023.105376
Shen H, Zhang C, Zhou H, Li C, Yuan H, Jiang J, Wang C (2023) Preparation of chlorophyll compounds and simulation of DFT optical properties assisted screening of photosensitizers from Ginkgo biloba leaves. Arab J Chem 105549. https://doi.org/10.1016/j.arabjc.2023.105549
Chafaa F, Hellel D, Khorief Nacereddine A, Djerourou A (2016) A theoretical study of the mechanism and selectivity of the intramolecular 1,3-dipolar cycloaddition reaction of the nitrone–alkene derived from 2-allylthiobenzaldehyde for the synthesis of tricyclic isoxazolidines. Tetrahedron Lett 57:67–70. https://doi.org/10.1016/j.tetlet.2015.11.069
Morgante P, Autschbach J (2023) Strategies to calculate fukui functions and applications to radicals with SOMO–HOMO inversion. J Chem Theory Comput 19:3929–3942. https://doi.org/10.1021/acs.jctc.3c00256
Lokhande KB, Tiwari A, Gaikwad S, Kore S, Nawani N, Wani M, Swamy KV, Pawar SV (2023) Computational docking investigation of phytocompounds from bergamot essential oil against Serratia marcescens protease and FabI: alternative pharmacological strategy. Comput Biol Chem 104:1476–9271. https://doi.org/10.1016/j.compbiolchem.2023.107829
Suresh PS, Kesarwani V, Kumari S, Shankar R, Sharma U (2023) Bisbenzylisoquinolines from Cissampelos pareira L. as antimalarial agents: molecular docking, pharmacokinetics analysis, and molecular dynamic simulation studies. Comput Biol Chem 104:1476–9271. https://doi.org/10.1016/j.compbiolchem.2023.107826
Tian H, Ketkar R, Tao P (2022) ADMETboost: a web server for accurate ADMET prediction. J Mol Model 28:408. https://doi.org/10.1007/s00894-022-05373-8
Karabacak Atay Ç, Dilek Ö, Tilki T et al (2023) A novel imidazole-based azo molecule: synthesis, characterization, quantum chemical calculations, molecular docking, molecular dynamics simulations and ADMET properties. J Mol Model 29:226. https://doi.org/10.1007/s00894-023-05625-1
Jin HS, Jong GB, Ri KH et al (2022) MEAM potential–based MD simulations of melting transition on Ni surfaces. J Mol Model 28:368. https://doi.org/10.1007/s00894-022-05357-8
Mary YS, Mary YS, Bielenica A et al (2021) Investigation of the reactivity properties of a thiourea derivative with anticancer activity by DFT and MD simulations. J Mol Model 27:217. https://doi.org/10.1007/s00894-021-04835-9
Joseph E, Swaminathan N, Kannan K (2020) Material identification for improving the strength of silica/SBR interface using MD simulations. J Mol Model 26:234. https://doi.org/10.1007/s00894-020-04489-z
Al-Otaibi JS, Mary YS, Mary YS et al (2023) TD-DFT, DFT, docking, MD simulations, and concentration-dependent SERS investigations of a bioactive trifluoromethyl derivative having human acetylcholinesterase and butyrylcholinesterase in silver colloids. J Mol Model 29:271. https://doi.org/10.1007/s00894-023-05679-1
Singh A (2018) Kumar D (2018) Effect of temperature on elastic properties of CNT-polyethylene nanocomposite and its interface using MD simulations. J Mol Model 24:178. https://doi.org/10.1007/s00894-018-3716-6
Němec T (2022) Nucleation parameters of SPC/E and TIP4P/2005 water vapor measured in NPT molecular dynamics simulations. J Mol Model 28:174. https://doi.org/10.1007/s00894-022-05130-x
Addo JK, Skaff DA, Miziorko HM (2016) Active site binding modes of inhibitors of Staphylococcus aureus mevalonate diphosphate decarboxylase from docking and molecular dynamics simulations. J Mol Model 22:13. https://doi.org/10.1007/s00894-015-2873-0
Pan QW, Henry SD, Scholte BJ, Tilanus HW, Janssen HL, Van der Laan LJ (2007) New therapeutic opportunities for hepatitis C based on small RNA. World J Gastroenterol 13:4431–4436. https://doi.org/10.3748/wjg.v13.i33.4431
Acknowledgements
The authors express their gratitude to the “Association Marcaine des chimistes théoriciens” (AMCT) for technical support.
Author information
Authors and Affiliations
Contributions
Data collection, material preparation, analysis, and writing by “Abdessadak O”; material preparation and analysis of DFT simulation part by “Kandwal P”; material and analysis of MD simulation part by “Alaqarbeh M”; material and analysis by “Tabti K”; data analysis, study justification and supervision by “Sbai A” data analysis, study justification and supervision by “Ajana M A”; supervision and project administration by “Lakhlifi T”; review, study justification and supervision by “Bouachrine M”.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Abdessadak, O., Kandwal, P., Alaqarbeh, M. et al. Exploring azomethine ylides reactivity with acrolein through cycloaddition reaction and computational antiviral activity assessment against hepatitis C virus. J Mol Model 30, 23 (2024). https://doi.org/10.1007/s00894-023-05818-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-023-05818-8