Abstract
Introduction
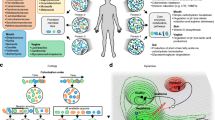
Infective endocarditis (IE) is an inflammatory disease usually caused by bacteria that enter the bloodstream and establish infections in the inner linings or valves of the heart, including blood vessels. Despite the availability of modern antimicrobial and surgical treatments, IE continues to cause substantial morbidity and mortality. Oral microbiota is considered one of the most significant risk factors for IE. The objective of this study was to evaluate the microbiota present in root canal (RC) and periodontal pocket (PP) clinical samples in cases with combined endo-periodontal lesions (EPL) to detect species related to IE using NGS.
Methods
Microbial samples were collected from 15 RCs and their associated PPs, also from 05 RCs with vital pulp tissues (negative control, NC). Genomic studies associated with bioinformatics, combined with structuring of a database (genetic sequences of bacteria reported for infective endocarditis), allowed for the assessment of the microbial community at both sites. Functional prediction was conducted using PICRUSt2.
Results
Parvimonas, Streptococcus, and Enterococcus were the major genera detected in the RCs and PPs. A total of 79, 96, and 11 species were identified in the RCs, PPs, and NCs, respectively. From them, a total of 34 species from RCs, 53 from PPs, and 2 from NCs were related to IE. Functional inference demonstrated that CR and PP microbiological profiles may not be the only risk factors for IE but may also be associated with systemic diseases, including myocarditis, human cytomegalovirus infection, bacterial invasion of epithelial cells, Huntington’s disease, amyotrophic lateral sclerosis, and hypertrophic cardiomyopathy. Additionally, it was possible to predict antimicrobial resistance variants for broad-spectrum drugs, including ampicillin, tetracycline, and macrolides.
Conclusion
Microorganisms present in the combined EPL may not be the only risk factor for IE but also for systemic diseases. Antimicrobial resistance variants for broad-spectrum drugs were inferred based on PICRUSt-2. State-of-the-art sequencing combined with bioinformatics has proven to be a powerful tool for conducting studies on microbial communities and could considerably assist in the diagnosis of serious infections.
Clinical relevance
Few studies have investigated the microbiota in teeth compromised by combined endo-periodontal lesions (EPL), but none have correlated the microbiological findings to any systemic condition, particularly IE, using NGS techniques. In such cases, the presence of apical periodontitis and periodontal disease can increase IE risk in susceptible patients.






Similar content being viewed by others
References
Carinci F, Martinelli M, Contaldo M et al (2018) Focus on periodontal disease and development of endocarditis. J Biol Regul Homeost Agents 32:143–147
Dajani AS, Taubert KA, Wilson W et al (1997) Prevention of bacterial endocarditis. Recommendations by the American Heart Association. JAMA 277:1794–1801
Gomez CA, Gerber DA, Zambrano E et al (2015) First case of infectious endocarditis caused by Parvimonas micra. Anaerobe 36:53–55. https://doi.org/10.1016/j.anaerobe.2015.10.007
Morris NA, Matiello M, Lyons JL, Samuels MA (2014) Neurologic complications in infective endocarditis: identification, management, and impact on cardiac surgery. Neurohospitalist 4:213–222. https://doi.org/10.1177/1941874414537077
Seymour RA, Lowry R, Whitworth JM, Martin MV (2000) Infective endocarditis, dentistry and antibiotic prophylaxis; time for a rethink? Br Dent J 189:610–616. https://doi.org/10.1038/sj.bdj.4800845
Debelian GJ, Olsen I, Tronstad L (1994) Systemic diseases caused by oral microorganisms. Dent Traumatol 10:57–65. https://doi.org/10.1111/j.1600-9657.1994.tb00061.x
Habib G, Lancellotti P, Antunes MJ et al (2015) 2015 ESC Guidelines for the management of infective endocarditis: the task force for the management of infective endocarditis of the European Society of Cardiology (ESC). Endorsed by: European Association for Cardio-Thoracic Surgery (EACTS), the European Association of Nuclear Medicine (EANM). Eur Heart J 36:3075–3128. https://doi.org/10.1093/eurheartj/ehv319
O’Connor EA, Cornwallis CK, Hasselquist D et al (2018) The evolution of immunity in relation to colonization and migration. Nat Ecol Evol 2:841–849. https://doi.org/10.1038/s41559-018-0509-3
Gomes BPFA, Berber VB, Kokaras AS et al (2015) Microbiomes of endodontic-periodontal lesions before and after chemomechanical preparation. J Endod 41:1975–1984. https://doi.org/10.1016/j.joen.2015.08.022
Kamada N, Chen GY, Inohara N, Núñez G (2013) Control of pathogens and pathobionts by the gut microbiota. Nat Immunol 14:685–690. https://doi.org/10.1038/ni.2608
Berg G, Rybakova D, Fischer D et al (2020) Microbiome definition re-visited: old concepts and new challenges. Microbiome 8:103. https://doi.org/10.1186/s40168-020-00875-0
Martín R, Miquel S, Langella P, Bermúdez-Humarán LG (2014) The role of metagenomics in understanding the human microbiome in health and disease. Virulence 5:413–423. https://doi.org/10.4161/viru.27864
Dahlén G, Pipattanagovit P, Rosling B, Möller ÅJR (1993) A comparison of two transport media for saliva and subgingival samples. Oral Microbiol Immunol 8:375–382. https://doi.org/10.1111/j.1399-302X.1993.tb00614.x
Mougeot J-LC, Stevens CB, Cotton SL et al (2016) Concordance of HOMIM and HOMI NGS technologies in the microbiome analysis of clinical samples. J Oral Microbiol 8:30379. https://doi.org/10.3402/jom.v8.30379
Lopes EM, Passini MRZ, Kishi LT et al (2021) Interrelationship between the microbial communities of the root canals and periodontal pockets in combined endodontic-periodontal diseases. Microorganisms 9:1925. https://doi.org/10.3390/microorganisms9091925
Andermann T, Antonelli A, Barrett RL, Silvestro D (2022) Estimating alpha, beta, and gamma diversity through deep learning. Front Plant Sci 13:839407. https://doi.org/10.3389/fpls.2022.839407
Chao A, Lee S-M (1992) Estimating the number of classes via sample coverage. J Am Stat Assoc 87:210–217. https://doi.org/10.1080/01621459.1992.10475194
Konopiński MK (2020) Shannon diversity index: a call to replace the original Shannon’s formula with unbiased estimator in the population genetics studies. PeerJ 8:e9391. https://doi.org/10.7717/peerj.9391
Maziarz M, Pfeiffer RM, Wan Y, Gail MH (2018) Using standard microbiome reference groups to simplify beta-diversity analyses and facilitate independent validation. Bioinformatics 34:3249–3257. https://doi.org/10.1093/bioinformatics/bty297
Segata N, Izard J, Waldron L et al (2011) Metagenomic biomarker discovery and explanation. Genome Biol 12:R60. https://doi.org/10.1186/gb-2011-12-6-r60
Chong J, Liu P, Zhou G, Xia J (2020) Using MicrobiomeAnalyst for comprehensive statistical, functional, and meta-analysis of microbiome data. Nat Protoc 15:799–821. https://doi.org/10.1038/s41596-019-0264-1
Douglas GM, Maffei VJ, Zaneveld JR et al (2020) PICRUSt2 for prediction of metagenome functions. Nat Biotechnol 38:685–688. https://doi.org/10.1038/s41587-020-0548-6
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680. https://doi.org/10.1093/nar/22.22.4673
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797. https://doi.org/10.1093/nar/gkh340
Soding J (2005) Protein homology detection by HMM-HMM comparison. Bioinformatics 21:951–960. https://doi.org/10.1093/bioinformatics/bti125
Kent WJ (2002) BLAT—The BLAST-Like Alignment Tool. Genome Res 12:656–664. https://doi.org/10.1101/gr.229202
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461. https://doi.org/10.1093/bioinformatics/btq461
Zaura E, Keijser BJ, Huse SM, Crielaard W (2009) Defining the healthy “core microbiome” of oral microbial communities. BMC Microbiol 9:259. https://doi.org/10.1186/1471-2180-9-259
Dewhirst FE, Chen T, Izard J et al (2010) The human oral microbiome. J Bacteriol 192:5002–5017. https://doi.org/10.1128/JB.00542-10
Del Giudice C, Vaia E, Liccardo D et al (2021) Infective endocarditis: a focus on oral microbiota. Microorganisms 9:1218. https://doi.org/10.3390/microorganisms9061218
Thomas C, Minty M, Vinel A et al (2021) Oral microbiota: a major player in the diagnosis of systemic diseases. Diagnostics 11:1376. https://doi.org/10.3390/diagnostics11081376
Kilian M, Chapple ILC, Hannig M et al (2016) The oral microbiome – an update for oral healthcare professionals. Br Dent J 221:657–666. https://doi.org/10.1038/sj.bdj.2016.865
Dhadse P, Gattani D, Mishra R (2010) The link between periodontal disease and cardiovascular disease: how far we have come in last two decades? J Indian Soc Periodontol 14:148. https://doi.org/10.4103/0972-124X.75908
Humphrey LL, Fu R, Buckley DI et al (2008) Periodontal disease and coronary heart disease incidence: a systematic review and meta-analysis. J Gen Intern Med 23:2079–2086. https://doi.org/10.1007/s11606-008-0787-6
Sanz M, Marco del Castillo A, Jepsen S et al (2020) Periodontitis and cardiovascular diseases: consensus report. J Clin Periodontol 47:268–288. https://doi.org/10.1111/jcpe.13189
Kumar PS (2017) From focal sepsis to periodontal medicine: a century of exploring the role of the oral microbiome in systemic disease: oral microbiome and systemic disease. J Physiol 595:465–476. https://doi.org/10.1113/JP272427
Debelian GJ, Olsen I, Tronstad L (1995) Bacteremia in conjunction with endodontic therapy. Dent Traumatol 11:142–149. https://doi.org/10.1111/j.1600-9657.1995.tb00476.x
Debelian GJ, Olsen I, Tronstad L (1998) Anaerobic bacteremia and fungemia in patients undergoing endodontic therapy: an overview. Ann Periodontol 3:281–287. https://doi.org/10.1902/annals.1998.3.1.281
Fernández-Hidalgo N, Escolà-Vergé L, Pericàs JM (2020) Enterococcus faecalis endocarditis: what’s next? Future Microbiol 15:349–364. https://doi.org/10.2217/fmb-2019-0247
Jordal S, Kittang BR, Salminen P-R et al (2018) Infective endocarditis in Western Norway: a 20-year retrospective survey. Infect Dis 50:757–763. https://doi.org/10.1080/23744235.2018.1482419
Gomes BPFA, Jacinto RC, Montagner F et al (2011) Analysis of the antimicrobial susceptibility of anaerobic bacteria isolated from endodontic infections in brazil during a period of nine years. J Endod 37:1058–1062. https://doi.org/10.1016/j.joen.2011.05.015
Pietropaoli D, Del Pinto R, Ferri C et al (2019) Definition of hypertension-associated oral pathogens in NHANES. J Periodontol 90:866–876. https://doi.org/10.1002/JPER.19-0046
Yakob M, Söder B, Meurman JH et al (2011) Prevotella nigrescens and Porphyromonas gingivalis are associated with signs of carotid atherosclerosis in subjects with and without periodontitis: periodontal microorganisms and atherosclerosis. J Periodontal Res 46:749–755. https://doi.org/10.1111/j.1600-0765.2011.01398.x
Kaplan JB, Ragunath C, Velliyagounder K et al (2004) Enzymatic detachment of Staphylococcus epidermidis biofilms. Antimicrob Agents Chemother 48:2633–2636. https://doi.org/10.1128/AAC.48.7.2633-2636.2004
Freires IA, Avilés-Reyes A, Kitten T et al (2017) Heterologous expression of Streptococcus mutans Cnm in Lactococcus lactis promotes intracellular invasion, adhesion to human cardiac tissues and virulence. Virulence 8:18–29. https://doi.org/10.1080/21505594.2016.1195538
Inenaga C, Hokamura K, Nakano K et al (2018) A potential new risk factor for stroke: streptococcus mutans with collagen-binding protein. World Neurosurg 113:e77–e81. https://doi.org/10.1016/j.wneu.2018.01.158
Tonomura S, Naka S, Tabata K et al (2019) Relationship between Streptococcus mutans expressing Cnm in the oral cavity and non-alcoholic steatohepatitis: a pilot study. BMJ Open Gastroenterol 6:e000329. https://doi.org/10.1136/bmjgast-2019-000329
Misaki T, Naka S, Hatakeyama R et al (2016) Presence of Streptococcus mutans strains harbouring the cnm gene correlates with dental caries status and IgA nephropathy conditions. Sci Rep 6:36455. https://doi.org/10.1038/srep36455
Tagliabue A, Rappuoli R (2018) Changing priorities in vaccinology: antibiotic resistance moving to the top. Front Immunol 9:1068. https://doi.org/10.3389/fimmu.2018.01068
Nishimura RA, Otto CM, Bonow RO et al (2017) 2017 AHA/ACC Focused Update of the 2014 AHA/ACC Guideline for the management of patients with valvular heart disease: a report of the American College of Cardiology/American Heart Association Task Force on Clinical Practice Guidelines. Circulation 135:e1159–e1195. https://doi.org/10.1161/CIR.0000000000000503
Thanavaro KL, Nixon JV (Ian) (2014) Endocarditis 2014: an update. Heart Lung 43:334–337. https://doi.org/10.1016/j.hrtlng.2014.03.009
Acknowledgements
This study was supported by São Paulo Research Foundation (FAPESP 2015/23479-5, 2017/14912-2, 19/14441-8, 2021/13871-6), National Scientific and Technological Development Council (CNPq 303852/2019-4, 421801/2021-2), Coordination for the Improvement of Higher Education Personnel (CAPES Finance Code 001), and Fund for the Support of Education, Research and Extension- State University of Campinas (FAEPEX 2036/17).
Author information
Authors and Affiliations
Contributions
Conceptualization: Brenda P. F. A. Gomes. Methodology: Brenda P. F. A. Gomes, Vanessa B. Berber, Rafaela C. Chapola, Vito M. Chiarelli-Neto, Emelly Aveiro, Maicon R. Z. Passini, Erica M Lopes, Tsute Chen, and Bruce J. Paster. Validation: Brenda P. F. A. Gomes, Vanessa B. Berber, Rafaela C. Chapola, Vito M. Chiarelli-Neto, Emelly Aveiro, Maicon R. Z. Passini, Erica M. Lopes, Tsute Chen, and Bruce J. Paster. Formal analysis: Brenda P. F. A. Gomes, Vanessa B. Berber, Rafaela C. Chapola, Vito M. Chiarelli-Neto, Emelly Aveiro, Maicon R. Z. Passini, Erica M. Lopes, Tsute Chen, and Bruce J. Paster. Investigation: Brenda P. F. A. Gomes, Vanessa B. Berber, Rafaela C. Chapola, Vito M. Chiarelli-Neto, Emelly Aveiro, Maicon R. Z. Passini, Erica M. Lopes, Tsute Chen, and Bruce J. Paster. Resources: Brenda P. F. A. Gomes. Data curation: Brenda P. F. A. Gomes, Vanessa B. Berber, Rafaela C. Chapola, Vito M. Chiarelli-Neto, Emelly Aveiro, Maicon R. Z. Passini, Erica M. Lopes, Tsute Chen, and Bruce J. Paster. Writing—original draft: Brenda P. F. A. Gomes, Vanessa B. Berber, Rafaela C. Chapola, Vito M. Chiarelli-Neto, Emelly Aveiro, Maicon R. Z. Passini, Erica M. Lopes, Tsute Chen, and Bruce J. Paster. Writing—review and editing preparation: Brenda P. F. A. Gomes, Vanessa B. Berber, Rafaela C. Chapola, Vito M. Chiarelli-Neto, Emelly Aveiro, Maicon R. Z. Passini, Erica M. Lopes, Tsute Chen, and Bruce J. Paster. Visualization: Brenda P. F. A. Gomes, Vanessa B. Berber, Rafaela C. Chapola, Vito M. Chiarelli-Neto, Emelly Aveiro, Maicon R. Z. Passini, Erica M. Lopes, Tsute Chen, and Bruce J. Paster. Project administration: Brenda P. F. A. Gomes, Vanessa B. Berber, Rafaela C. Chapola, Vito M. Chiarelli-Neto, Emelly Aveiro, Maicon R. Z. Passini, Erica M. Lopes, Tsute Chen, and Bruce J. Paster. Funding acquisition: Brenda P. F. A. Gomes.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent to participate
For this type of study, formal consent is not required. Informed consent was obtained from all individual participants included in the study.
Conflict of interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gomes, B.P.F.A., Berber, V.B., Chiarelli-Neto, V.M. et al. Microbiota present in combined endodontic-periodontal diseases and its risks for endocarditis. Clin Oral Invest 27, 4757–4771 (2023). https://doi.org/10.1007/s00784-023-05104-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00784-023-05104-0