Abstract
Proteins are the material basis of life and the primary carriers of life activities, containing various impurities that must be removed before use. To keep pace with the increasing complexity of protein samples, it is essential to constantly work on developing new purification technologies for downstream processes. While traditional downstream purification methods rely heavily on protein A affinity chromatography, there is still a lot of interest in finding safer and more cost-effective alternatives to protein A. Many non-affinity ligands and technologies have also been developed in biological purification recently. Here, the current status of biotechnology and the progress of protein separation technology from 2018 to 2023 are reviewed from the aspects of new preparation methods and new composite materials of commonly used separation media. The research status of new ligands with different mechanisms of action was reviewed, including the expanded application of affinity ligands, the development prospect of biotechnology such as polymer grafting, continuous column technology, and its new applications.
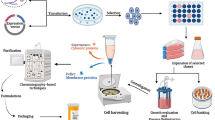
Graphical Abstract

















Similar content being viewed by others
References
Hondius DC, Eigenhuis KN, Morrema THJ, van der Schors RC, van Nierop P, Bugiani M et al (2018) Proteomics analysis identifies new markers associated with capillary cerebral amyloid angiopathy in Alzheimer’s disease. Acta Neuropathol Commun 6(1):46. https://doi.org/10.1186/s40478-018-0540-2
Rolland DCM, Lim MS, Elenitoba-Johnson KSJ (2019) Mass spectrometry and proteomics in hematology. Semin Hematol 56(1):52–57. https://doi.org/10.1053/j.seminhematol.2018.05.009
Kim S-Y, Seong H. (2021) Modulation of physicochemical properties of magnetic agarose microspheres by hydrolysis-suppressive sequential crosslinking. Colloids Surf A Physicochem Eng Asp 630:127607. https://doi.org/10.1016/j.colsurfa.2021.127607.
Li Q, Xue Z, Zhao J, Ao C, Jia X, Xia T, et al. (2020) Mass production of high thermal conductive boron nitride/nanofibrillated cellulose composite membranes. Chem Eng J 383:123101. https://doi.org/10.1016/j.cej.2019.123101
Li X-Q, Li Q, Gong F-L, Lei J-D, Zhao X, Ma G-H et al (2015) Preparation of large-sized highly uniform agarose beads by novel rotating membrane emulsification. J Membr Sci 476:30–39. https://doi.org/10.1016/j.memsci.2014.11.017
Shukla AA, Thommes J (2010) Recent advances in large-scale production of monoclonal antibodies and related proteins. Trends Biotechnol 28(5):253–261. https://doi.org/10.1016/j.tibtech.2010.02.001
Liu HF, Ma J, Winter C, Bayer R (2010) Recovery and purification process development for monoclonal antibody production. MAbs 2(5):480–499. https://doi.org/10.4161/mabs.2.5.12645
Kruljec N, Bratkovic T (2017) Alternative affinity ligands for immunoglobulins. Bioconjug Chem 28(8):2009–2030. https://doi.org/10.1021/acs.bioconjchem.7b00335
Kruljec N, Molek P, Bratkovic T (2016) Peptide ligands of the immunoglobulin G Fc region identified by screening phage libraries and site-directed mutagenesis. Febs Journal 283:166
Arakawa T, Ponce S, Young G (2015) Isoform separation of proteins by mixed-mode chromatography. Protein Expr Purif 116:144–151. https://doi.org/10.1016/j.pep.2015.08.013
Hong Y, Liu N, Wei W, Yu LL, Ma G, Sun Y (2014) Protein adsorption to poly(ethylenimine)-modified Sepharose FF: III. Comparison between different proteins. J Chromatogr A 1342:30–6. https://doi.org/10.1016/j.chroma.2014.03.036
Yu L, Zhang L, Sun Y (2015) Protein behavior at surfaces: orientation, conformational transitions and transport. J Chromatogr A 1382:118–134. https://doi.org/10.1016/j.chroma.2014.12.087
Pagkaliwangan M, Hummel J, Gjoka X, Bisschops M, Schofield M (2019) Optimized continuous multicolumn chromatography enables increased productivities and cost savings by employing more columns. Biotechnol J 14(2):e1800179. https://doi.org/10.1002/biot.201800179
Otes O, Flato H, Winderl J, Hubbuch J, Capito F (2017) Feasibility of using continuous chromatography in downstream processing: comparison of costs and product quality for a hybrid process vs. a conventional batch process. J Biotechnol 259:213–20. https://doi.org/10.1016/j.jbiotec.2017.07.001
Jiang W, Wang J, Yuan D, Gao Z, Hu B, Li Y et al (2023) Fabrication, characterization and emulsifying properties of agarose microgel. Int J Biol Macromol 241:124565. https://doi.org/10.1016/j.ijbiomac.2023.124565
Martínez-Sanz M, Gomez-Barrio LP, Zhao M, Tiwari B, Knutsen SH, Ballance S, et al. (2021) Alternative protocols for the production of more sustainable agar-based extracts from Gelidium sesquipedale. Algal Res 55:102254. https://doi.org/10.1016/j.algal.2021.102254.
Xiao Q, Ma M, Chen J, Zhang Y, Chen F, Weng H et al (2022) Preparation of macroporous rigid agarose microspheres by pre-crosslinking with cyclic anhydride. Int J Biol Macromol 222(Pt A):41–54. https://doi.org/10.1016/j.ijbiomac.2022.09.146
Zhao L, Huang Y, Zhu K, Miao Z, Zhao J, Che XJ et al (2020) Manipulation of pore structure during manufacture of agarose microspheres for bioseparation. Eng Life Sci 20(11):504–513. https://doi.org/10.1002/elsc.202000023
Xiao Q, Weng H, Chen G, Xiao A (2019) Preparation and characterization of octenyl succinic anhydride modified agarose derivative. Food Chem 279:30–39. https://doi.org/10.1016/j.foodchem.2018.11.133
Zhao L, Huang L, Huang Y, Zhu K, Che X, Du Y et al (2022) Preparation and structural regulation of macroporous agarose microspheres for highly efficient adsorption of giant biomolecules. Colloid Polym Sci 300(6):691–705. https://doi.org/10.1007/s00396-022-04968-0
Zhao X, Huang L, Wu J, Huang YD, Zhao L, Wu N et al (2019) Fabrication of rigid and macroporous agarose microspheres by pre-cross-linking and surfactant micelles swelling method. Colloids Surf B Biointerfaces 182:110377. https://doi.org/10.1016/j.colsurfb.2019.110377
Siar E-H, Arana-Peña S, Barbosa O, Zidoune MN, Fernandez-Lafuente R (2018) Solid phase chemical modification of agarose glyoxyl-ficin: improving activity and stability properties by amination and modification with glutaraldehyde. Process Biochem 73:109–116. https://doi.org/10.1016/j.procbio.2018.07.013
Siar E-H, Zaak H, Kornecki JF, Zidoune MN, Barbosa O, Fernandez-Lafuente R (2017) Stabilization of ficin extract by immobilization on glyoxyl agarose. Preliminary characterization of the biocatalyst performance in hydrolysis of proteins. Process Biochemistry 58:98–104. https://doi.org/10.1016/j.procbio.2017.04.009
Zhao L, Li S, Liang C, Qiao L, Du K. (2021) High-strength and low-crystallinity cellulose/agarose composite microspheres: fabrication, characterization and protein adsorption. Biochem Eng J 166:107826. https://doi.org/10.1016/j.bej.2020.107826.
Hajizadeh S, Ye L (2019) Hierarchical macroporous material with dual responsive copolymer brushes and phenylboronic acid ligands for bioseparation of proteins and living cells. Sep Purif Technol 224:95–105. https://doi.org/10.1016/j.seppur.2019.05.002
Wulan DR, Rahmaniyah WR, Zulfikar MA, Nurachman Z (2022) Enhancement of microsphere specificity to purify human serum albumin from blood plasma. J Chromatogr A 1683:463535. https://doi.org/10.1016/j.chroma.2022.463535
Mofidian R, Barati A, Jahanshahi M, Shahavi MH (2020) Fabrication of novel agarose–nickel bilayer composite for purification of protein nanoparticles in expanded bed adsorption column. Chem Eng Res Des 159:291–299. https://doi.org/10.1016/j.cherd.2020.03.024
Arana-Pena S, Rios NS, Mendez-Sanchez C, Lokha Y, Goncalves LRB, Fernandez-Lafuente R (2020) Use of polyethylenimine to produce immobilized lipase multilayers biocatalysts with very high volumetric activity using octyl-agarose beads: avoiding enzyme release during multilayer production. Enzyme Microb Technol 137:109535. https://doi.org/10.1016/j.enzmictec.2020.109535
Urner LH, Mohammadifar E, Ludwig K, Shutin D, Fiorentino F, Liko I et al (2021) Anionic dendritic polyglycerol for protein purification and delipidation. ACS Applied Polymer Materials 3(11):5903–5911. https://doi.org/10.1021/acsapm.1c01127
Xue Z-X, Yang G-P, Wang G-C, Niu J-F, Cao X-Y (2007) Preparation of porous chitosan/agarose microsphere and its R-phycoerythrin release properties. J Appl Polym Sci 103(4):2759–2766. https://doi.org/10.1002/app.25335
Lei Y, Liu X, Lu L, Liu C, Xu R, Huang S et al (2020) Rapid preparation of 1-vinylimidazole based non-affinity polymers for the highly-selective purification of antibodies from multiple biological sources. J Chromatogr A 1632:461607. https://doi.org/10.1016/j.chroma.2020.461607
Xiong Z, Zhou W, Sun L, Li X, Zhao D, Chen Y et al (2014) Konjac glucomannan microspheres for low-cost desalting of protein solution. Carbohydr Polym 111:56–62. https://doi.org/10.1016/j.carbpol.2014.04.059
Cuatrecasas P, and Wilchek, M. (1968) Single-step purification of avidine from egg white by affinity chromatography on biocytinSepharose columns. Biochem Biophys Res Commun 33:235−239
Ayyar BV, Arora S, Murphy C, O’Kennedy R (2012) Affinity chromatography as a tool for antibody purification. Methods 56(2):116–129. https://doi.org/10.1016/j.ymeth.2011.10.007
Tao P, Poddar S, Sun Z, Hage DS, Chen J (2018) Analysis of solute-protein interactions and solute-solute competition by zonal elution affinity chromatography. Methods 146:3–11. https://doi.org/10.1016/j.ymeth.2018.01.020
Grom M, Kozorog M, Caserman S, Pohar A, Likozar B (2018) Protein A affinity chromatography of Chinese hamster ovary (CHO) cell culture broths containing biopharmaceutical monoclonal antibody (mAb): experiments and mechanistic transport, binding and equilibrium modeling. J Chromatogr B Analyt Technol Biomed Life Sci 1083:44–56. https://doi.org/10.1016/j.jchromb.2018.02.032
Zarrineh M, Mashhadi IS, Farhadpour M, Ghassempour A (2020) Mechanism of antibodies purification by protein A. Anal Biochem 609:113909. https://doi.org/10.1016/j.ab.2020.113909
Hagemann F, Adametz P, Wessling M, Thom V (2020) Modeling hindered diffusion of antibodies in agarose beads considering pore size reduction due to adsorption. J Chromatogr A 1626:461319. https://doi.org/10.1016/j.chroma.2020.461319
Wang Y, Zhang X, Han N, Wu Y, Wei D (2018) Oriented covalent immobilization of recombinant protein A on the glutaraldehyde activated agarose support. Int J Biol Macromol 120(Pt A):100–108. https://doi.org/10.1016/j.ijbiomac.2018.08.074
Zheng H, Wei F, Tian J, Wang C, Xue C (2022) Preparation of nickel-chelated iminodiacetate-functionalized macroporous agarose monolith using modular and clickable building blocks for affinity separation of histidine-tagged recombinant proteins. J Chromatogr A 1682:463509. https://doi.org/10.1016/j.chroma.2022.463509
Arakawa T, Tomioka Y, Nakagawa M, Sakuma C, Kurosawa Y, Ejima D, et al. (2023) Non-affinity purification of antibodies. Antibodies (Basel) 12(1):12010015. https://doi.org/10.3390/antib12010015.
Lin DQ, Tong HF, Wang HY, Shao S, Yao SJ (2012) Molecular mechanism of hydrophobic charge-induction chromatography: interactions between the immobilized 4-mercaptoethyl-pyridine ligand and IgG. J Chromatogr A 1260:143–153. https://doi.org/10.1016/j.chroma.2012.08.080
Lin DQ, Tong HF, Wang HY, Yao SJ (2012) Molecular insight into the ligand-IgG interactions for 4-mercaptoethyl-pyridine based hydrophobic charge-induction chromatography. J Phys Chem B 116(4):1393–1400. https://doi.org/10.1021/jp206817b
Chi H, Tian S, Li X, Chen Y, Xu Q, Wang Q et al (2023) Construction of lipid raft-coupled agarose gels as bioaffinity chromatography materials and validation with tropomyosin-related kinase A-targeted drugs. J Chromatogr A 1691:463803. https://doi.org/10.1016/j.chroma.2023.463803
Hou Z, Han X, Wang Z, Ghazanfar S, Yang J, Liu H (2021) A terminal alkyne and disulfide functionalized agarose resin specifically enriches azidohomoalanine labeled nascent proteins. J Chromatogr B Analyt Technol Biomed Life Sci 1165:122527. https://doi.org/10.1016/j.jchromb.2021.122527
Chung JA, Wollack JW, Hovlid ML, Okesli A, Chen Y, Mueller JD et al (2009) Purification of prenylated proteins by affinity chromatography on cyclodextrin-modified agarose. Anal Biochem 386(1):1–8. https://doi.org/10.1016/j.ab.2008.09.007
Reese HR, Shanahan CC, Lembo J, Tsonev L, Hirsh A, Menegatti S (2020) Chromatographic assay to probe the binding energy and mechanisms of homologous proteins to surface-bound ligands. J Chromatogr B Analyt Technol Biomed Life Sci 1136:121927. https://doi.org/10.1016/j.jchromb.2019.121927
Kittler S, Ebner J, Besleaga M, Larsbrink J, Darnhofer B, Birner-Gruenberger R et al (2022) Recombinant protein L: production, purification and characterization of a universal binding ligand. J Biotechnol 359:108–115. https://doi.org/10.1016/j.jbiotec.2022.10.002
King C, Patel R, Ponniah G, Nowak C, Neill A, Gu Z et al (2018) Characterization of recombinant monoclonal antibody variants detected by hydrophobic interaction chromatography and imaged capillary isoelectric focusing electrophoresis. J Chromatogr B Analyt Technol Biomed Life Sci 1085:96–103. https://doi.org/10.1016/j.jchromb.2018.03.049
Rodler A, Beyer B, Ueberbacher R, Hahn R, Jungbauer A (2018) Hydrophobic interaction chromatography of proteins: studies of unfolding upon adsorption by isothermal titration calorimetry. J Sep Sci 41(15):3069–3080. https://doi.org/10.1002/jssc.201800016
Tassi M, De Vos J, Chatterjee S, Sobott F, Bones J, Eeltink S (2018) Advances in native high-performance liquid chromatography and intact mass spectrometry for the characterization of biopharmaceutical products. J Sep Sci 41(1):125–144. https://doi.org/10.1002/jssc.201700988
Evans C, Morimitsu Y, Nishi R, Yoshida M, Takei T (2022) Novel hydrophobically modified agarose cryogels fabricated using dimethyl sulfoxide. J Biosci Bioeng 133(4):390–395. https://doi.org/10.1016/j.jbiosc.2021.12.009
Rajesh S, Crandall C, Schneiderman S, Menkhaus TJ (2018) Cellulose-graft-polyethyleneamidoamine anion-exchange nanofiber membranes for simultaneous protein adsorption and virus filtration. ACS Applied Nano Materials 1(7):3321–3330. https://doi.org/10.1021/acsanm.8b00519
Lou Y, Ji G, Liu Q, Wang P, Zhang R, Zhang Y et al (2018) Secretory expression and scale-up production of recombinant human thyroid peroxidase via baculovirus/insect cell system in a wave-type bioreactor. Protein Expr Purif 149:7–12. https://doi.org/10.1016/j.pep.2018.04.005
Huang Y, Bi J, Zhao L, Ma G, Su Z (2010) Regulation of protein multipoint adsorption on ion-exchange adsorbent and its application to the purification of macromolecules. Protein Expr Purif 74(2):257–263. https://doi.org/10.1016/j.pep.2010.07.002
Moreno-González M, Chuekitkumchorn P, Silva M, Groenewoud R, Ottens M (2021) High throughput process development for the purification of rapeseed proteins napin and cruciferin by ion exchange chromatography. Food Bioprod Process 125:228–241. https://doi.org/10.1016/j.fbp.2020.11.011
Hendricks R, Reese D, Fedesco M, Chinn M, Zhang J, Hutchinson M (2022) Simplified strategy for developing purification processes for antibody-drug conjugates using cation-exchange chromatography in flow-through mode. J Chromatogr A 1666:462865. https://doi.org/10.1016/j.chroma.2022.462865
Chern MK, Shiah WJ, Chen JJ, Tsai TY, Lin HY, Liu CW (2009) Single-step protein purification by back flush in ion exchange chromatography. Anal Biochem 392(2):174–176. https://doi.org/10.1016/j.ab.2009.05.045
Zhu M, Carta G (2015) Protein adsorption equilibrium and kinetics in multimodal cation exchange resins. Adsorption 22(2):165–179. https://doi.org/10.1007/s10450-015-9735-z
Jing S-Y, Gou J-X, Gao D, Wang H-B, Yao S-J, Lin D-Q. (2020) Separation of monoclonal antibody charge variants using cation exchange chromatography: resins and separation conditions optimization. Sep Purif Technol 235:116136. https://doi.org/10.1016/j.seppur.2019.116136.
Huang C, Wang Y, Xu X, Mills J, Jin W, Ghose S et al (2020) Hydrophobic property of cation-exchange resins affects monoclonal antibody aggregation. J Chromatogr A 1631:461573. https://doi.org/10.1016/j.chroma.2020.461573
Roberts JA, Kimerer L, Carta G (2020) Effects of molecule size and resin structure on protein adsorption on multimodal anion exchange chromatography media. J Chromatogr A 1628:461444. https://doi.org/10.1016/j.chroma.2020.461444
Zhao G, Dong XY, Sun Y (2009) Ligands for mixed-mode protein chromatography: principles, characteristics and design. J Biotechnol 144(1):3–11. https://doi.org/10.1016/j.jbiotec.2009.04.009
Gu J-L, Pi W-Y, Xie W-F, Gu J-W, Wang Y-J, Xiao L, et al. (2021) Improved antibody adsorption performance of phenyl-based mixed-mode adsorbents by adjusting the functional group of ligand. Biochem Eng J 176:108092. https://doi.org/10.1016/j.bej.2021.108092.
Li M, Zou X, Zhang Q, Lin D, Yao S. (2020) Binding mechanism of functional moieties of a mixed-mode ligand in antibody purification. Chem Eng J. 400. https://doi.org/10.1016/j.cej.2020.125887.
Li M, Lin D, Yao S, Zhang Q (2022) Study on antibody adsorption and elution performance of carboxyl and hydrophobic groups on mixed-mode ligands. J Sep Sci 45(15):2946–2955. https://doi.org/10.1002/jssc.202200342
Ge CT, Cai QY, Zhang QL, Chu WN, Yao SJ, Lin DQ (2020) Rational design of specific ligands for human serum albumin separation and applications. J Sep Sci 43(21):4028–4035. https://doi.org/10.1002/jssc.202000409
Luo YD, Zhang QL, Yao SJ, Lin DQ (2018) Evaluation of adsorption selectivity of immunoglobulins M, A and G and purification of immunoglobulin M with mixed-mode resins. J Chromatogr A 1533:77–86. https://doi.org/10.1016/j.chroma.2017.12.018
Silva-Santos AR, Paulo PMR, F. Prazeres DM. (2022) Scalable purification of single stranded DNA scaffolds for biomanufacturing DNA-origami nanostructures: exploring anion-exchange and multimodal chromatography. Sep Purif Technol 298:121623. https://doi.org/10.1016/j.seppur.2022.121623.
Koehnlein W, Holzgreve A, Schwendner K, Skudas R, Schelter F (2023) Purification of hydrophobic complex antibody formats using a moderately hydrophobic mixed mode cation exchange resin. J Chromatogr A 1687:463696. https://doi.org/10.1016/j.chroma.2022.463696
Ren J, Xiang X, Ji F, Gao X, Han L, Jia L (2021) Benzotriazole-5-carboxylic as a mixed-mode ligand for chromatographic separation of antibody with enhanced adsorption capacity. J Chromatogr B Analyt Technol Biomed Life Sci 1179:122652. https://doi.org/10.1016/j.jchromb.2021.122652
Bresolin IT, de Souza MC, Bueno SM (2010) A new process of IgG purification by negative chromatography: adsorption aspects of human serum proteins onto omega-aminodecyl-agarose. J Chromatogr B Analyt Technol Biomed Life Sci 878(23):2087–2093. https://doi.org/10.1016/j.jchromb.2010.06.009
Aoyama S, Matsumoto Y, Mori C, Sota K (2022) Application of novel mixed mode chromatography (MMC) resins having a hydrophobic modified polyallylamine ligand for monoclonal antibody purification. J Chromatogr B Analyt Technol Biomed Life Sci 1191:123072. https://doi.org/10.1016/j.jchromb.2021.123072
Zhang Q, Schimpf F, Lu H-L, Lin D-Q, Yao S-J (2016) Binary adsorption processes of albumin and immunoglobulin on hydrophobic charge-induction resins. J Chem Eng Data 61(3):1353–1360. https://doi.org/10.1021/acs.jced.5b01108
Ye J, Zhang Y, Meng J. (2022) Protein–ligand interactions for hydrophobic charge-induction chromatography: a QCM-D study. Appl Surf Sci 572:151420. https://doi.org/10.1016/j.apsusc.2021.151420.
Shi W, Zhang X, Kong X, Fu J, Zhang S, Li K et al (2022) Evaluation of hydrophobic charge-induction ligand efficiency for protein adsorption in one single cycle. J Chromatogr A 1668:462923. https://doi.org/10.1016/j.chroma.2022.462923
Fang YM, Lin DQ, Yao SJ (2018) Review on biomimetic affinity chromatography with short peptide ligands and its application to protein purification. J Chromatogr A 1571:1–15. https://doi.org/10.1016/j.chroma.2018.07.082
Zou X, Zhang Q, Lu H, Lin D, Yao S (2019) Development of a hybrid biomimetic ligand with high selectivity and mild elution for antibody purification. Chem Eng J 368:678–686. https://doi.org/10.1016/j.cej.2019.03.014
Fang YM, Chen SG, Lin DQ, Yao SJ (2019) A new tetrapeptide biomimetic chromatographic resin for antibody separation with high adsorption capacity and selectivity. J Chromatogr A 1604:460474. https://doi.org/10.1016/j.chroma.2019.460474
Liu T, Lin D-Q, Wu Q-C, Zhang Q-L, Wang C-X, Yao S-J (2016) A novel polymer-grafted hydrophobic charge-induction chromatographic resin for enhancing protein adsorption capacity. Chem Eng J 304:251–258. https://doi.org/10.1016/j.cej.2016.06.074
Fang YM, Zhu HY, Lin DQ, Yao SJ (2020) A novel dextran-grafted tetrapeptide resin for antibody purification. J Sep Sci 43(19):3816–3823. https://doi.org/10.1002/jssc.202000325
Gu J, Zhang Y, Tong H, Liu Y, Sun L, Wang Y et al (2019) Preparation and evaluation of dextran-grafted mixed-mode chromatography adsorbents. J Chromatogr A 1599:1–8. https://doi.org/10.1016/j.chroma.2019.04.021
Zhao Y, Dong X, Yu L, Liu Y, Sun Y (2018) Characterization of new polymer-grafted protein cation exchangers developed by partial neutralization of carboxyl groups derivatized by modification of poly(ethylenimine)-Sepharose with succinic anhydride. J Chromatogr A 1550:28–34. https://doi.org/10.1016/j.chroma.2018.03.046
Liu T, Angelo JM, Lin DQ, Lenhoff AM, Yao SJ (2017) Characterization of dextran-grafted hydrophobic charge-induction resins: structural properties, protein adsorption and transport. J Chromatogr A 1517:44–53. https://doi.org/10.1016/j.chroma.2017.07.090
Qiao L, Du Y, Du K. (2022) Grafting diethylaminoethyl dextran to macroporous cellulose microspheres: a protein anion exchanger of high capacity and fast uptake rate. Sep Purif Technol 297:121434. https://doi.org/10.1016/j.seppur.2022.121434.
Hao D, Zhang R, Ge J, Ye P, Song C, Zhu K et al (2021) Rapid and high-capacity loading of IgG monoclonal antibodies by polymer brush and peptides functionalized microspheres. J Chromatogr A 1640:461948. https://doi.org/10.1016/j.chroma.2021.461948
Xenopoulos A (2015) A new, integrated, continuous purification process template for monoclonal antibodies: process modeling and cost of goods studies. J Biotechnol 213:42–53. https://doi.org/10.1016/j.jbiotec.2015.04.020
Gjoka X, Gantier R, Schofield M (2017) Transfer of a three step mAb chromatography process from batch to continuous: optimizing productivity to minimize consumable requirements. J Biotechnol 242:11–18. https://doi.org/10.1016/j.jbiotec.2016.12.005
Gjoka X, Rogler K, Martino RA, Gantier R, Schofield M (2015) A straightforward methodology for designing continuous monoclonal antibody capture multi-column chromatography processes. J Chromatogr A 1416:38–46. https://doi.org/10.1016/j.chroma.2015.09.005
Nestola P, Silva RJ, Peixoto C, Alves PM, Carrondo MJ, Mota JP (2014) Adenovirus purification by two-column, size-exclusion, simulated countercurrent chromatography. J Chromatogr A 1347:111–121. https://doi.org/10.1016/j.chroma.2014.04.079
Nestola P, Silva RJ, Peixoto C, Alves PM, Carrondo MJ, Mota JP (2015) Robust design of adenovirus purification by two-column, simulated moving-bed, size-exclusion chromatography. J Biotechnol 213:109–119. https://doi.org/10.1016/j.jbiotec.2015.01.030
Ren X, Zhang K, Gao D, Fu Q, Zeng J, Zhou D et al (2018) Mixed-mode liquid chromatography with a stationary phase co-functionalized with ionic liquid embedded C18 and an aryl sulfonate group. J Chromatogr A 1564:137–144. https://doi.org/10.1016/j.chroma.2018.06.017
Zaveckas M, Snipaitis S, Pesliakas H, Nainys J, Gedvilaite A (2015) Purification of recombinant virus-like particles of porcine circovirus type 2 capsid protein using ion-exchange monolith chromatography. J Chromatogr B Analyt Technol Biomed Life Sci 991:21–28. https://doi.org/10.1016/j.jchromb.2015.04.004
Xi H, Yu J, Sun Q, Lu J, Gu T, Guo X et al (2018) Expression and purification of pneumococcal surface protein A of clade 4 in Escherichia coli using hydroxylapatite and ion-exchange column chromatography. Protein Expr Purif 151:56–61. https://doi.org/10.1016/j.pep.2018.06.008
Heidebrecht HJ, Kainz B, Schopf R, Godl K, Karcier Z, Kulozik U et al (2018) Isolation of biofunctional bovine immunoglobulin G from milk- and colostral whey with mixed-mode chromatography at lab and pilot scale. J Chromatogr A 1562:59–68. https://doi.org/10.1016/j.chroma.2018.05.046
Sun YN, Shi C, Zhong XZ, Chen XJ, Chen R, Zhang QL et al (2022) Model-based evaluation and model-free strategy for process development of three-column periodic counter-current chromatography. J Chromatogr A 1677:463311. https://doi.org/10.1016/j.chroma.2022.463311
Shi C, Gao ZY, Zhang QL, Yao SJ, Slater NKH, Lin DQ (2020) Model-based process development of continuous chromatography for antibody capture: a case study with twin-column system. J Chromatogr A 1619:460936. https://doi.org/10.1016/j.chroma.2020.460936
Sun YN, Shi C, Zhang QL, Slater NKH, Jungbauer A, Yao SJ et al (2021) Comparison of protein A affinity resins for twin-column continuous capture processes: process performance and resin characteristics. J Chromatogr A 1654:462454. https://doi.org/10.1016/j.chroma.2021.462454
Funding
This work was supported by the Tianjin Research Innovation Project for Postgraduate Students (2022SKY056 and 2021YJSB194).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Qi, C., Chen, L. Progress of ligand-modified agarose microspheres for protein isolation and purification. Microchim Acta 191, 149 (2024). https://doi.org/10.1007/s00604-024-06224-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-024-06224-4