Abstract
Key message
The specifically expressed alleles of CsMCT, CsGGPPS2, CsHMGS2, CsHMGR3, CsTPS10, CsADH4, CsDAHPS, CsCM3, and CsAS2 may play critical roles in the aroma formation of Camellia sinensis 'Jinguanyin' cultivar. CsMVD, CsDXS2, CsTPS3, CsTPS9, CsTPS10, CsTPS13 and CsTPS14 play an essential role in the formation of the characteristic floral and fruit aromas of oolong tea.
Abstract
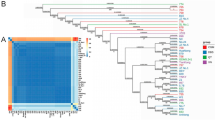
To explore the role of alleles in the aroma biosynthesis pathways of the tea plant, this study was conducted to identify and analyze the alleles of structural genes in the MVA, MEP, LOX and shikimate pathways based on Camellia sinensis ‘Jinguanyin’ haplotype-resolved genomes. The results showed that 69 pairs of alleles were identified in the four pathways, of which 24 genes showed allele-specific expression (ASE) in at least one tissue. TGY-allele and HD-allele correlations of buds, young leaves, and stems were higher in the six tissues of JGY. KEGG analysis revealed that 24 ASEGs were significantly enriched in GPI-anchor biosynthesis and monoterpenoid biosynthesis pathways. The results suggest that the specifically expressed alleles of CsMCT, CsGGPPS2, CsHMGS2, CsHMGR3, CsTPS10, CsADH4, CsDAHPS, CsCM3, and CsAS2 may play critical roles in the aroma formation of JGY cultivars. The qRT‒PCR results showed that the bias of CsTPS1, CsMVD, CsDHQS, and CsAS2 alleles was basically consistent with the expression of the parents (TGY and HD) of JGY in young leaves. In addition, qRT‒PCR was used to detect the expression of aroma-related genes during the manufacturing process of JGY and parents (TGY and HD) oolong tea. It showed that CsMVD, CsDXS2, CsTPS3, CsTPS9, CsTPS10, CsTPS13 and CsTPS14 play an essential role in the formation of the characteristic floral and fruit aromas of oolong tea.









Similar content being viewed by others
Data availability
The datasets analyzed in this research are available from the corresponding author upon request.
Abbreviations
- JGY:
-
Jin Guanyin
- TGY:
-
Tie Guanyin
- HD:
-
Huang Dan
- ASE:
-
Allele-specific expression
- ASEG:
-
Allele-specific expression gene
- FPKM:
-
Fragments per kilobase per million
- FDR:
-
False discovery rate
- GO:
-
Gene Ontology
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- LOX:
-
Lipoxygenase
References
Baldauf JA, Vedder L, Schoof H, Hochholdinger F (2020) Robust non-syntenic gene expression patterns in diverse maize hybrids during root development. J Exp Bot 71:865–876
Chen CJ, Chen H, Zhang Y, Thomas HR, Frank MH, He YH, Xia R (2020a) TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant 13:1194–1202
Chen XJ, Wang PJ, Lin XY, Gu MY, Zheng YC, Zheng ZL, Ye NX (2020b) Identification and expression analysis of terpene synthesis related genes during the withering of white tea. Tea Sci 40:363–374
Chen XJ, Wang PJ, Gu MY, Lin XY, Hou BH, Zheng YC, Sun Y, Jin S, Ye NX (2021) R2R3-MYB transcription factor family in tea plant (Camellia sinensis): Genome-wide characterization, phylogeny, chromosome location, structure and expression patterns. Genomics 113:1565–1578
Deng WW, Wu YL, Li YY, Tan Z, Wei CL (2016) Molecular cloning and characterization of hydroperoxide lyase gene in the leaves of tea plant (Camellia sinensis). J Agric Food Chem 64:1770–1776
Dudareva N, Klempien A, Muhlemann JK, Kaplan I (2013) Biosynthesis, function and metabolic engineering of plant volatile organic compounds. New Phytol 198:16–32
Gao T, Hou BH, Shao SX, Xu MT, Zheng YC, Jin S, Wang PJ, Ye NX (2023) Differential metabolites and transcriptional regulation of seven major tea cultivars (Camellia sinensis) in China. J Integrative Agric. https://doi.org/10.1016/j.jia.2023.02.009
Gaur U, Li K, Mei SQ, Liu GS (2013) Research progress in allele-specific expression and its regulatory mechanisms. J Appl Genet 54:271–283
Gui JD, Fu XM, Zhou Y, Katsuno T, Mei X, Deng RF, Xu XL, Zhang LY, Dong F, Watanabe N, Yang ZY (2015) Does enzymatic hydrolysis of glycosidically bound volatile compounds really contribute to the formation of volatile compounds during the oolong tea manufacturing process? J Agric Food Chem 63:6905–6914
Guo M, Rupe MA, Zinselmeier C, Habben J, Bowen BA, Smith OS (2004) Allelic variation of gene expression in maize hybrids. Plant Cell 16:1707–1716
Guo JC, Yang RX, Zhang WJ, Chen ZH, Chen J (2008) Breeding and application of two the hybrid Jinguanyin and Huangguanyin. Guizhou Sci, pp 20–24
Huang XH, Yang SH, Gong JY, Zhao Y, Feng Q, Gong H, Li WJ, Zhan QL, Cheng BY, Xia JH, Chen N, Hao ZN, Liu KY, Zhu CR, Huang T, Zhao Q, Zhang L, Fan DL, Zhou CC, Lu YQ, Weng QJ, Wang ZX, Li JY, Han B (2015) Genomic analysis of hybrid rice varieties reveals numerous superior alleles that contribute to heterosis. Nat Commun 6:6258
Klosinska M, Picard CL, Gehring M (2016) Conserved imprinting associated with unique epigenetic signatures in the Arabidopsis genus. Nat Plants 2:16145
Knight JC (2004) Allele-specific gene expression uncovered. Trends Genet 20:113–116
Liu GF, Liu JJ, He ZR, Wang FM, Yang H, Yan YF, Gao MJ, Gruber MY, Wan XC, Wei S (2018) Implementation of CsLIS/NES in linalool biosynthesis involves transcript splicing regulation in Camellia sinensis. Plant Cell Environ 41:176–186
Ma X, Xing F, Jia QX, Zhang QL, Hu T, Wu BG, Shao L, Zhao Y, Zhang QF, Zhou DX (2021) Parental variation in CHG methylation is associated with allelic-specific expression in elite hybrid rice. Plant Physiol 186:1025–1041
Paschold A, Jia Y, Marcon C, Lund S, Larson NB, Yeh CT, Ossowski S, Lanz C, Nettleton D, Schnable PS, Hochholdinger F (2012) Complementation contributes to transcriptome complexity in maize (Zea mays L.) hybrids relative to their inbred parents. Genome Res 22:2445–2454
Pham GM, Newton L, Wiegert-Rininger K, Vaillancourt B, Douches DS, Buell CR (2017) Extensive genome heterogeneity leads to preferential allele expression and copy number-dependent expression in cultivated potato. Plant J 92:624–637
Rodrigues AMG, Correr FH, Furtado A, Botha FC, Henry RJ (2022) Limited allele-specific gene expression in highly polyploid sugarcane. Genome Res 32:297–308
Shao L, Xing F, Xu CH, Zhang QH, Che J, Wang XM, Song JM, Li XH, Xiao JH, Chen LL, Ouyang YD, Zhang QF (2019) Patterns of genome-wide allele-specific expression in hybrid rice and the implications on the genetic basis of heterosis. PNAS 116:5653–5658
Springer NM, Stupar RM (2007) Allele-specific expression patterns reveal biases and embryo-specific parent-of-origin effects in hybrid maize. Plant Cell 19:2391–2402
von Korff M, Radovic S, Choumane W, Stamati K, Udupa SM, Grando S, Ceccarelli S, Mackay I, Powell W, Baum M, Morgante M (2009) Asymmetric allele-specific expression in relation to developmental variation and drought stress in barley hybrids. Plant J 59:14–26
Wang B, Tseng E, Baybayan P, Eng K, Regulski M, Jiao YP, Wang LY, Olson A, Chougule K, Buren PV, Ware D (2020) Variant phasing and haplotypic expression from long-read sequencing in maize. Commun Biol 3:78
Wang PJ, Yu JX, Jin S, Chen S, Yue C, Wang WL, Gao SL, Cao HL, Zheng YC, Gu MY, Chen XJ, Sun Y, Guo YQ, Yang JF, Zhang XT, Ye NX (2021) Genetic basis of high aroma and stress tolerance in the oolong tea cultivar genome. Hortic Res 8:107–121
Wang PJ, Fang JP, Lin HZ, Yang WW, Yu JX, Hong YP, Jiang MW, Gu MY, Chen QC, Zheng YC, Liao ZY, Chen GX, Yang JF, Jin S, Zhang XT, Ye NX (2022a) Genomes of single- and double-petal jasmines (Jasminum sambac) provide insights into their divergence time and structural variations. Plant Biotechnol J 20:1232–1234
Wang PJ, Gu MY, Shao SX, Chen XM, Hou BH, Ye NX, Zhang XT (2022b) Changes in non-volatile and volatile metabolites associated with heterosis in tea plants (Camellia sinensis). J Agr Food Chem 70:3067–3078
Wang PJ, Gu MY, Yu XK, Shao SX, Du JY, Wang YB, Wang FQ, Chen S, Liao ZY, Ye NX, Zhang XT (2022c) Allele-specific expression and chromatin accessibility contribute to heterosis in tea plants (Camellia sinensis). Plant J 112(5):1194–1211
Waters AJ, Makarevitch I, Noshay J, Burghardt LT, Hirsch CN, Hirsch CD, Springer NM (2017) Natural variation for gene expression responses to abiotic stress in maize. Plant J 89:706–717
Wiberley AE, Donohue AR, Westphal MM, Sharkey TD (2009) Regulation of isoprene emission from poplar leaves throughout a day. Plant Cell Environ 32:939–947
Yang ZY, Baldermann S, Watanabe N (2013) Recent studies of the volatile compounds in tea. Food Res Int 53:585–599
Zeng LT, Zhou Y, Gui JD, Fu XM, Mei X, Zhen YP, Ye TX, Du B, Dong F, Watanabe N, Yang ZY (2016) Formation of volatile tea constituent indole during the oolong tea manufacturing process. J Agric Food Chem 64:5011–5019
Zeng LT, Zhou Y, Fu XM, Liao YY, Yuan YF, Jia YX, Dong F, Yang ZY (2018) Biosynthesis of jasmine lactone in tea (Camellia sinensis) leaves and its formation in response to multiple stresses. J Agr Food Chem 66:3899–3909
Zeng LT, Watanabe N, Yang ZY (2019) Understanding the biosyntheses and stress response mechanisms of aroma compounds in tea (Camellia sinensis) to safely and effectively improve tea aroma. Crit Rev Food Sci Nutr 59:2321–2334
Zeng L, Jin S, Fu YQ, Chen LS, Yin JF, Xu YQ (2022) A targeted and untargeted metabolomics analysis of “Oriental Beauty” oolong tea during processing. Beverage Plant Res. https://doi.org/10.48130/BPR-2022-0020
Zhang X, Borevitz JO (2009) Global analysis of allele-specific expression in Arabidopsis thaliana. Genetics 182:943–954
Zhang XT, Chen S, Shi LQ, Gong DP, Zhang SC, Zhao Q, Zhan DL, Vasseur L, Wang YB, Yu JX, Liao ZY, Xu XD, Qi R, Wang WL, Ma YR, Wang PJ, Ye NX, Ma DN, Shi Y, Wang HF, Ma XK, Kong XR, Lin J, Wei LF, Ma YY, Li RY, Hu GP, He HF, Zhang L, Ming R, Wang G, Tang HB, You MS (2021) Haplotype-resolved genome assembly provides insights into evolutionary history of the tea plant Camellia sinensis. Nat Genet 53:1250–1259
Zheng YC, Wang PJ, Chen XJ, Sun Y, Yue C, Ye NX (2019) Transcriptome and metabolite profiling reveal novel insights into volatile heterosis in the tea plant (Camellia Sinensis). Molecules 24:3380
Zheng YC, Hu QC, Wu ZJ, Bi WJ, Chen B, Hao ZL, Wu LY, Ye NX, Sun Y (2022a) Volatile metabolomics and coexpression network analyses provide insight into the formation of the characteristic cultivar aroma of oolong tea (Camellia sinensis). LWT 164:113666
Zheng YC, Hu QC, Yang Y, Wu ZJ, Wu LY, Wang PJ, Deng HL, Ye NX, Sun Y (2022b) Architecture and dynamics of the wounding-induced gene regulatory network during the oolong tea manufacturing process (Camellia sinensis). Front Plant Sci 12:788469
Zhou Y, Zeng LT, Liu XY, Gui JD, Mei X, Fu XM, Dong F, Tang JC, Zhang LY, Yang ZY (2017) Formation of (E)-nerolidol in tea (Camellia sinensis) leaves exposed to multiple stresses during tea manufacturing. Food Chem 231:78–86
Zhou XC, Yang J, Jian GT, Zeng LT, Gu DC, Su XG, Yang ZY (2022) Role of histone deacetylase CsHDA8 in regulating the accumulation of indole during the oolong tea manufacturing process. Beverage Plant Res. https://doi.org/10.48130/BPR-2022-0017
Zhuang Y, Adams KL (2007) Extensive allelic variation in gene expression in populus F1 hybrids. Genetics 177:1987–1996
Funding
This research was funded by the National Natural Science Foundation of China (Grant No. 32202550), Major Special Project of Scientific and Technological Innovation on Anxi Tea (Grant No. AX2021001) and Special Fund for Science and Technology Innovation of Fujian Zhang Tianfu Tea Development Foundation (FJZTF01).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
We declare that there is no conflict of interest.
Additional information
Communicated by Gómez.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gu, M., Gao, T., Xu, M. et al. Identification and analysis of alleles in the aroma biosynthesis pathways based on Camellia sinensis ‘Jinguanyin’ haplotype-resolved genomes. Trees 37, 1627–1641 (2023). https://doi.org/10.1007/s00468-023-02447-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00468-023-02447-9