Abstract
Background
Glioma is the prevailing malignant tumor affecting the brain and central nervous system, constituting over 80% of all malignant brain tumors. HOXD9 has been implicated in the development of glioma, but the specific mechanism of its influence on glioma pathogenesis remains incompletely understood. The purpose of this study was to investigate the role of HOXD9 in glioma and examine the changes in HOXD9 expression during the progression of glioma, thus contributing new insights into the pathogenesis of glioma.
Methods
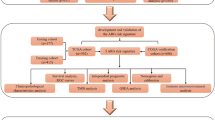
Glioma samples from the Cancer Gene Atlas (TCGA) and Chinese Glioma Genome Atlas (CGGA) datasets were included in this study. Variations in HOXD9 expression in gliomas between different subgroups of multiple clinical characteristics were explored, and the expression was validated in glioma samples using qRT-PCR and western blotting. Next, the impact of HOXD9 on the prognosis of gliomas was explored by survival analysis, receiver operating characteristic curve, and nomogram plots. Subsequently, the association between HOXD9 and the tumor immune microenvironment was explored using the ssGSEA algorithm and the ESTIMATE algorithm. Then, immune-related pathways associated with HOXD9 were determined by differential express analysis and GSEA. Finally, HOXD9-related genomic alterations were identified.
Results
HOXD9 expression is upregulated and correlated with malignant properties in glioma. Similarly, our validation results showed significantly upregulated protein and mRNA levels of HOXD9 in glioma brain tissues. In addition, high HOXD9 expression was indicative of a poor prognosis for glioma patients. Additionally, elevated HOXD9 levels were associated with reduced tumor purity and higher levels of immune invasion. Finally, HOXD9 was significantly associated with genomic alterations.
Conclusion
Overall, this study has unveiled a significant association between HOXD9 and the prognosis and survival of glioma patients. Our findings highlight the potential of HOXD9 as a prognostic biomarker, implicating its role in influencing the glioma immune microenvironment.










Similar content being viewed by others
Availability of data and materials
The publicly available datasets used in this study can be found in the TCGA database (https://portal.gdc.cancer.gov/).
References
Balkwill F, Mantovani A (2001) Inflammation and cancer: back to Virchow? Lancet 357(9255):539–545. https://doi.org/10.1016/s0140-6736(00)04046-0
Batash R, Asna N, Schaffer P, Francis N, Schaffer M (2017) Glioblastoma multiforme, diagnosis and treatment; recent literature review. Curr Med Chem 24(27):3002–3009. https://doi.org/10.2174/0929867324666170516123206
Bhatlekar S, Fields JZ, Boman BM (2018) Role of Hox genes in stem cell differentiation and cancer. Stem Cells Int 2018:3569493. https://doi.org/10.1155/2018/3569493
Bhattacharya S, Andorf S, Gomes L, Dunn P, Schaefer H, Pontius J et al (2014) Immport: disseminating data to the public for the future of immunology. Immunol Res 58(2–3):234–239. https://doi.org/10.1007/s12026-014-8516-1
Boix E, Torrent M, Sánchez D, Nogués MV (2008) The antipathogen activities of eosinophil cationic protein. Curr Pharm Biotechnol 9(3):141–152. https://doi.org/10.2174/138920108784567353
Bosma I, Douw L, Bartolomei F, Heimans JJ, van Dijk BW, Postma TJ et al (2008) Synchronized brain activity and neurocognitive function in patients with low-grade glioma: a magnetoencephalography study. Neuro Oncol 10(5):734–744. https://doi.org/10.1215/15228517-2008-034
Charoentong P, Finotello F, Angelova M, Mayer C, Efremova M, Rieder D et al (2017) Pan-cancer immunogenomic analyses reveal genotype-immunophenotype relationships and predictors of response to checkpoint blockade. Cell Rep 18(1):248–262. https://doi.org/10.1016/j.celrep.2016.12.019
Chen Z, Hambardzumyan D (2018) Immune microenvironment in glioblastoma subtypes. Front Immunol 9:1004. https://doi.org/10.3389/fimmu.2018.01004
Chen B, Gao C, Wang H, Sun J, Han Z (2022) Molecular analysis of prognosis and immune infiltration of ovarian cancer based on Homeobox D genes. Comput Math Methods Med 2022:3268386. https://doi.org/10.1155/2022/3268386
Ge X, Xu M, Cheng T, Hu N, Sun P, Lu B et al (2022) Tp53i13 promotes metastasis in glioma via macrophages, neutrophils, and fibroblasts and is a potential prognostic biomarker. Front Immunol 13:974346. https://doi.org/10.3389/fimmu.2022.974346
Guo X, Wang Y, Zhang H, Qin C, Cheng A, Liu J et al (2020) Identification of the prognostic value of immune-related genes in esophageal cancer. Front Genet 11:989. https://doi.org/10.3389/fgene.2020.00989
Hänzelmann S, Castelo R, Guinney J (2013) Gsva: Gene set variation analysis for microarray and rna-seq data. BMC Bioinform 14:7. https://doi.org/10.1186/1471-2105-14-7
He J, Jiang M, Liu J, Zhu R, Lv W, Lian R et al (2023) Homeobox D9 drives the malignant phenotypes and enhances the programmed death ligand-1 expression in non-small cell lung cancer cells via binding to angiopoietin-2 promoter. World J Surg Oncol 21(1):93. https://doi.org/10.1186/s12957-023-02969-z
Huang S, Song Z, Zhang T, He X, Huang K, Zhang Q et al (2020) Identification of immune cell infiltration and immune-related genes in the tumor microenvironment of glioblastomas. Front Immunol 11:585034. https://doi.org/10.3389/fimmu.2020.585034
Jin Z, Sun D, Song M, Zhu W, Liu H, Wang J et al (2022) Comprehensive analysis of Hox family members as novel diagnostic and prognostic markers for hepatocellular carcinoma. J Oncol 2022:5758601. https://doi.org/10.1155/2022/5758601
Komohara Y, Horlad H, Ohnishi K, Fujiwara Y, Bai B, Nakagawa T et al (2012) Importance of direct macrophage-tumor cell interaction on progression of human glioma. Cancer Sci 103(12):2165–2172. https://doi.org/10.1111/cas.12015
Liberzon A, Birger C, Thorvaldsdóttir H, Ghandi M, Mesirov JP, Tamayo P (2015) The molecular signatures database (Msigdb) Hallmark gene set collection. Cell Syst 1(6):417–425. https://doi.org/10.1016/j.cels.2015.12.004
Liu M, Xiao Y, Tang W, Li J, Hong L, Dai W et al (2020) Hoxd9 promote epithelial-mesenchymal transition and metastasis in colorectal carcinoma. Cancer Med 9(11):3932–3943. https://doi.org/10.1002/cam4.2967
Lu X, Maturi NP, Jarvius M, Yildirim I, Dang Y, Zhao L et al (2022) Cell-lineage controlled epigenetic regulation in glioblastoma stem cells determines functionally distinct subgroups and predicts patient survival. Nat Commun 13(1):2236. https://doi.org/10.1038/s41467-022-29912-2
Miao X, Leng X, Zhang Q (2017) The current state of nanoparticle-induced macrophage polarization and reprogramming research. Int J Mol Sci. https://doi.org/10.3390/ijms18020336
Ohgaki H, Kleihues P (2013) The definition of primary and secondary glioblastoma. Clin Cancer Res 19(4):764–772. https://doi.org/10.1158/1078-0432.Ccr-12-3002
O’Shea JJ, Schwartz DM, Villarino AV, Gadina M, McInnes IB, Laurence A (2015) The Jak-Stat pathway: impact on human disease and therapeutic intervention. Annu Rev Med 66:311–328. https://doi.org/10.1146/annurev-med-051113-024537
Ou A, Ott M, Fang D, Heimberger AB (2021) The Role and therapeutic targeting of Jak/Stat signaling in glioblastoma. Cancers (basel). https://doi.org/10.3390/cancers13030437
Propper DJ, Balkwill FR (2022) Harnessing Cytokines and chemokines for cancer therapy. Nat Rev Clin Oncol 19(4):237–253. https://doi.org/10.1038/s41571-021-00588-9
Saito N, Hirai N, Aoki K, Sato S, Suzuki R, Hiramoto Y et al (2019) Genetic and lineage classification of glioma-initiating cells identifies a clinically relevant glioblastoma model. Cancers (basel). https://doi.org/10.3390/cancers11101564
Shen W, Song Z, Zhong X, Huang M, Shen D, Gao P et al (2022) Sangerbox: a comprehensive, interaction-friendly clinical bioinformatics analysis platform. Imeta 1(3):e36
Sherman BT, Hao M, Qiu J, Jiao X, Baseler MW, Lane HC et al (2022) David: a web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res 50(W1):W216–W221. https://doi.org/10.1093/nar/gkac194
Tabuse M, Ohta S, Ohashi Y, Fukaya R, Misawa A, Yoshida K et al (2011) Functional analysis of Hoxd9 in human gliomas and glioma cancer stem cells. Mol Cancer 10:60. https://doi.org/10.1186/1476-4598-10-60
Wang JJ, Wang H, Zhu BL, Wang X, Qian YH, Xie L et al (2021) Development of a prognostic model of glioma based on immune-related genes. Oncol Lett 21(2):116. https://doi.org/10.3892/ol.2020.12377
Wang L, Qiao C, Cao L, Cai S, Ma X, Song X et al (2022) Significance of Hoxd transcription factors family in progression, migration and angiogenesis of cancer. Crit Rev Oncol Hematol 179:103809. https://doi.org/10.1016/j.critrevonc.2022.103809
Wei J, Liu J, Liang S, Sun M, Duan J (2020) Low-dose exposure of silica nanoparticles induces neurotoxicity via neuroactive ligand-receptor interaction signaling pathway in zebrafish embryos. Int J Nanomed 15:4407–4415. https://doi.org/10.2147/ijn.S254480
Wu X, Yang T, Qian L, Zhang D, Yang H (2021) Construction of a new tumor immunity-related signature to assess and classify the prognostic risk of colorectal cancer. Int J Gen Med 14:6661–6676. https://doi.org/10.2147/ijgm.S325511
Xu S, Tang L, Li X, Fan F, Liu Z (2020) Immunotherapy for glioma: current management and future application. Cancer Lett 476:1–12. https://doi.org/10.1016/j.canlet.2020.02.002
Yang HH, Duan JX, Liu SK, Xiong JB, Guan XX, Zhong WJ et al (2020) A Cox-2/Seh dual inhibitor Ptupb alleviates lipopolysaccharide-induced acute lung injury in mice by inhibiting Nlrp3 inflammasome activation. Theranostics 10(11):4749–4761. https://doi.org/10.7150/thno.43108
Yang HH, Jiang HL, Tao JH, Zhang CY, Xiong JB, Yang JT et al (2022) Mitochondrial citrate accumulation drives alveolar epithelial cell necroptosis in lipopolysaccharide-induced acute lung injury. Exp Mol Med 54(11):2077–2091. https://doi.org/10.1038/s12276-022-00889-8
Yoshihara K, Shahmoradgoli M, Martínez E, Vegesna R, Kim H, Torres-Garcia W et al (2013) Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun 4:2612. https://doi.org/10.1038/ncomms3612
Zhang CY, Zhong WJ, Liu YB, Duan JX, Jiang N, Yang HH et al (2023) Eets alleviate alveolar epithelial cell senescence by inhibiting endoplasmic reticulum stress through the Trim25/Keap1/Nrf2 Axis. Redox Biol 63:102765. https://doi.org/10.1016/j.redox.2023.102765
Zhong WJ, Yang HH, Guan XX, Xiong JB, Sun CC, Zhang CY et al (2019) Inhibition of glycolysis alleviates lipopolysaccharide-induced acute lung injury in a mouse model. J Cell Physiol 234(4):4641–4654. https://doi.org/10.1002/jcp.27261
Zhu H, Dai W, Li J, Xiang L, Wu X, Tang W et al (2019) Hoxd9 promotes the growth, invasion and metastasis of gastric cancer cells by transcriptional activation of Rufy3. J Exp Clin Cancer Res 38(1):412. https://doi.org/10.1186/s13046-019-1399-1
Acknowledgements
We are grateful to the Bioinformatics Center, Xiangya Hospital, Central South University, for partial support of this work.
Funding
This research was funded by the Natural Science Foundation of Hunan Province (2022JJ80120).
Author information
Authors and Affiliations
Contributions
SX, TS, YZ and CW designed the study, analyzed the data and wrote the manuscript. HX and RG performed validation in human brain tissues. TC and ZH assisted in analyzing the data and revising the manuscript. DL and MJ critically read and edited the manuscript. All authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Authorship
The authors have no relevant financial or non-financial interests to disclose.
Ethics approval
This study was approved by the Medical Ethics Committee of Xiangya Hospital, Central South University (201907786).
Consent to participate
Not applicable.
Consent to publish
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xu, S., Xiao, H., Song, T. et al. HOXD9 is a potential prognostic biomarker involved in immune microenvironment of glioma. J Cancer Res Clin Oncol 149, 14911–14926 (2023). https://doi.org/10.1007/s00432-023-05275-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-023-05275-z