Abstract
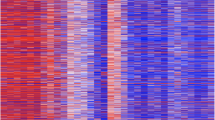
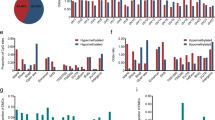
Research on the DNA methylation status of gastric cancer (GC) has primarily focused on identifying invasive GC to develop biomarkers for diagnostic. However, DNA methylation in noninvasive GC remains unclear. We conducted a comprehensive DNA methylation profiling study of differentiated-type intramucosal GCs (IMCs). Illumina 850K microarrays were utilized to assess the DNA methylation profiles of formalin-fixed paraffin-embedded tissues from eight patients who were Epstein-Barr virus-negative and DNA mismatch repair proficient, including IMCs and paired adjacent nontumor mucosa. Gene expression profiling microarray data from the GEO database were analyzed via bioinformatics to identify candidate methylation genes. The final validation was conducted using quantitative real-time PCR, the TCGA methylation database, and single-sample gene set enrichment analysis (GSEA). Genome-wide DNA methylation profiling revealed a global decrease in methylation in IMCs compared with nontumor tissues. Differential methylation analysis between IMCs and nontumor tissues identified 449 differentially methylated probes, with a majority of sites showing hypomethylation in IMCs compared with nontumor tissues (66.1% vs 33.9%). Integrating two RNA-seq microarray datasets, we found one hypomethylation-upregulated gene: eEF1A2, overlapped with our DNA methylation data. The mRNA expression of eEF1A2 was higher in twenty-four IMC tissues than in their paired adjacent nontumor tissues. GSEA indicated that the functions of eEF1A2 were associated with the development of IMCs. Furthermore, TCGA data indicated that eEF1A2 is hypomethylated in advanced GC. Our study illustrates the implications of DNA methylation alterations in IMCs and suggests that aberrant hypomethylation and high mRNA expression of eEF1A2 might play a role in IMCs development.





Similar content being viewed by others
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author upon reasonable request.
References
Yang K, Lu L, Liu H, Wang X, Gao Y, Yang L, Li Y, Su M, Jin M, Khan S (2021) A comprehensive update on early gastric cancer: defining terms, etiology, and alarming risk factors. Expert Rev Gastroenterol Hepatol 15(3):255–73. https://doi.org/10.1080/17474124.2021.1845140
Correa P (1988) A human model of gastric carcinogenesis. Cancer Res 48(13):3554–3560
Lauren P (1965) The two histological main types of gastric carcinoma: diffuse and so-called intestinal-type carcinoma. an attempt at a histo-clinical classification. Acta Pathol Microbiol Scand 64:31–49. https://doi.org/10.1111/apm.1965.64.1.31
Nagtegaal ID, Odze RD, Klimstra D, Paradis V, Rugge M, Schirmacher P, Washington KM, Carneiro F, Cree IA (2020) The 2019 WHO classification of tumours of the digestive system. Histopathology 76(2):182–8. https://doi.org/10.1111/his.13975
Bonelli P, Borrelli A, Tuccillo FM, Silvestro L, Palaia R, Buonaguro FM (2019) Precision medicine in gastric cancer. World J Gastrointest Oncol 11(10):804–29. https://doi.org/10.4251/wjgo.v11.i10.804
Usui G, Matsusaka K, Mano Y, Urabe M, Funata S, Fukayama M, Ushiku T, Kaneda A (2021) DNA methylation and genetic aberrations in gastric cancer. Digestion 102(1):25–32. https://doi.org/10.1159/000511243
Niwa T, Tsukamoto T, Toyoda T, Mori A, Tanaka H, Maekita T, Ichinose M, Tatematsu M, Ushijima T (2010) Inflammatory processes triggered by Helicobacter pylori infection cause aberrant DNA methylation in gastric epithelial cells. Cancer Res 70(4):1430–40. https://doi.org/10.1158/0008-5472.Can-09-2755
Meng Q, Lu YX, Ruan DY, Yu K, Chen YX, Xiao M, Wang Y, Liu ZX, Xu RH, Ju HQ, Qiu MZ (2021) DNA methylation regulator-mediated modification patterns and tumor microenvironment characterization in gastric cancer. Mol Ther Nucleic Acids 24:695–710. https://doi.org/10.1016/j.omtn.2021.03.023
Liu D, Ma X, Yang F, Xiao D, Jia Y, Wang Y (2020) Discovery and validation of methylated-differentially expressed genes in Helicobacter pylori-induced gastric cancer. Cancer Gene Ther 27(6):473–85. https://doi.org/10.1038/s41417-019-0125-7
Necula L, Matei L, Dragu D, Neagu AI, Mambet C, Nedeianu S, Bleotu C, Diaconu CC, Chivu-Economescu M (2019) Recent advances in gastric cancer early diagnosis. World J Gastroenterol 25(17):2029–44. https://doi.org/10.3748/wjg.v25.i17.2029
Kaneda A, Feinberg AP (2005) Loss of imprinting of IGF2: a common epigenetic modifier of intestinal tumor risk. Cancer Res 65(24):11236–40. https://doi.org/10.1158/0008-5472.Can-05-2959
Du P, Zhang X, Huang CC, Jafari N, Kibbe WA, Hou L, Lin SM (2010) Comparison of Beta-value and M-value methods for quantifying methylation levels by microarray analysis. BMC Bioinformatics 11:587. https://doi.org/10.1186/1471-2105-11-587
Anderson KJ, Cormier RT, Scott PM (2019) Role of ion channels in gastrointestinal cancer. World J Gastroenterol 25(38):5732–72. https://doi.org/10.3748/wjg.v25.i38.5732
Xu X, Feng L, Liu Y, Zhou WX, Ma YC, Fei GJ, An N, Li Y, Wu X, Yao F, Cheng SJ, Lu XH (2014) Differential gene expression profiling of gastric intraepithelial neoplasia and early-stage adenocarcinoma. World journal of gastroenterology 20(47):17883–93. https://doi.org/10.3748/wjg.v20.i47.17883
Zhang Y, Wu X, Zhang C, Wang J, Fei G, Di X, Lu X, Feng L, Cheng S, Yang A (2020) Dissecting expression profiles of gastric precancerous lesions and early gastric cancer to explore crucial molecules in intestinal-type gastric cancer tumorigenesis. J Pathol 251(2):135–46. https://doi.org/10.1002/path.5434
Watanabe Y, Kim HS, Castoro RJ, Chung W, Estecio MR, Kondo K, Guo Y, Ahmed SS, Toyota M, Itoh F, Suk KT, Cho MY, Shen L, Jelinek J, Issa JP (2009) Sensitive and specific detection of early gastric cancer with DNA methylation analysis of gastric washes. Gastroenterology 136(7):2149–58. https://doi.org/10.1053/j.gastro.2009.02.085
Cristiano L (2022) The pseudogenes of eukaryotic translation elongation factors (EEFs): Role in cancer and other human diseases. Genes Dis 9(4):941–58. https://doi.org/10.1016/j.gendis.2021.03.009
Tan Y, Sun R, Liu L, Yang D, Xiang Q, Li L, Tang J, Qiu Z, Peng W, Wang Y, Ye L, Ren G, Xiang T (2021) Tumor suppressor DRD2 facilitates M1 macrophages and restricts NF-κB signaling to trigger pyroptosis in breast cancer. Theranostics 11(11):5214–31. https://doi.org/10.7150/thno.58322
Sun Y, Du C, Wang B, Zhang Y, Liu X, Ren G (2014) Up-regulation of eEF1A2 promotes proliferation and inhibits apoptosis in prostate cancer. Biochem Biophys Res Commun 450(1):1–6. https://doi.org/10.1016/j.bbrc.2014.05.045
Zang W, Wang Y, Wang T, Du Y, Chen X, Li M, Zhao G (2015) miR-663 attenuates tumor growth and invasiveness by targeting eEF1A2 in pancreatic cancer. Mol Cancer 14:37. https://doi.org/10.1186/s12943-015-0315-3
Rhodes DR, Yu J, Shanker K, Deshpande N, Varambally R, Ghosh D, Barrette T, Pandey A, Chinnaiyan AM (2004) ONCOMINE: a cancer microarray database and integrated data-mining platform. Neoplasia 6(1):1–6. https://doi.org/10.1016/s1476-5586(04)80047-2
Long K, Wang H, Song Z, Yin X, Wang Y (2020) EEF1A2 mutations in epileptic encephalopathy/intellectual disability: understanding the potential mechanism of phenotypic variation. Epilepsy Behav 105:106955. https://doi.org/10.1016/j.yebeh.2020.106955
Hassan MK, Kumar D, Naik M, Dixit M (2018) The expression profile and prognostic significance of eukaryotic translation elongation factors in different cancers. PLoS One 13(1):e0191377. https://doi.org/10.1371/journal.pone.0191377
Wang Q, Xiong F, Wu G, Liu W, Chen J, Wang B, Chen Y (2022) Gene body methylation in cancer: molecular mechanisms and clinical applications. Clin Epigenet 14(1):154. https://doi.org/10.1186/s13148-022-01382-9
Di Mario F, Goni E (2014) Gastric acid secretion: changes during a century. Best Pract Res Clin Gastroenterol 28(6):953–65. https://doi.org/10.1016/j.bpg.2014.10.006
Duarte HO, Freitas D, Gomes C, Gomes J, Magalhães A, Reis CA (2016) Mucin-type O-glycosylation in gastric carcinogenesis. Biomolecules 6(3). https://doi.org/10.3390/biom6030033
Akyala AI, Peppelenbosch MP (2018) Gastric cancer and Hedgehog signaling pathway: emerging new paradigms. Genes Cancer 9(1–2):1–10. https://doi.org/10.18632/genesandcancer.168
Oliveira D, Hentze J, O’Rourke CJ, Andersen JB, Høgdall C, Høgdall EV (2021) DNA methylation in ovarian tumors-a comparison between fresh tissue and FFPE samples. Reprod Sci 28(11):3212–8. https://doi.org/10.1007/s43032-021-00589-0
Kling T, Wenger A, Beck S, Carén H (2017) Validation of the MethylationEPIC BeadChip for fresh-frozen and formalin-fixed paraffin-embedded tumours. Clin Epigenet 9:33. https://doi.org/10.1186/s13148-017-0333-7
Acknowledgements
This work was supported by Innovation Technology Project, Beijing Chao-Yang Hospital, Capital Medical University, Beijing, China (No. 21kcjj-9).
Author information
Authors and Affiliations
Contributions
ZS, MJ, and XJ designed the study. ZS, XG, and RL completed the experiment. ZS, XH, JL, and XL collected the data. ZS, XH, JL, and XL analyzed and interpreted the data. ZS, XJ, and MJ prepared the manuscript. All authors contributed to the article and approved the submitted version.
Corresponding authors
Ethics declarations
Ethics approval
Approval of the research protocol by the local ethics committee (Beijing Chao-Yang Hospital, Capital Medical University, NO. 2020-KE-323).
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Shi, Z., Guo, X., Hu, X. et al. DNA methylation profiling identifies epigenetic signatures of early gastric cancer. Virchows Arch 484, 687–695 (2024). https://doi.org/10.1007/s00428-024-03765-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00428-024-03765-0