Abstract
Purpose
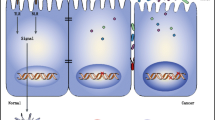
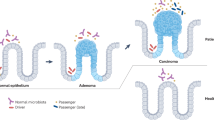
Colorectal cancer (CRC) is the cancer of the colon and rectum. Recent research has found a link between CRC and human gut microbiota. This review explores the effect of gut microbiota on colorectal carcinogenesis and the development of chemoresistance.
Methods
A literature overview was performed to identify the gut microbiota species that showed altered abundance in CRC patients and the mechanisms by which some of them aid in the development of chemoresistance.
Results
Types of gut microbiota present and methods of analyzing them were discussed. We observed that numerous microbiota showed altered abundance in CRC patients and could act as a biomarker for CRC diagnosis and treatment. Further, it was demonstrated that microbes also have a role in the development of chemoresistance by mechanisms like immune system activation, drug modification, and autophagy modulation. Finally, the key issue of the growing global problem of antimicrobial resistance and its relationship with CRC was highlighted.
Conclusion
This review discussed the role of gut microbiota dysbiosis on colorectal cancer progression and the development of chemoresistance.




Similar content being viewed by others
References
Mármol I, Sánchez-de-Diego C, Dieste AP et al (2017) Colorectal carcinoma: a general overview and future perspectives in colorectal cancer. Int J Mol Sci 18. https://doi.org/10.3390/ijms18010197
Araghi M, Soerjomataram I, Jenkins M et al (2019) Global trends in colorectal cancer mortality: projections to the year 2035. Int J Cancer 144:2992–3000. https://doi.org/10.1002/ijc.32055
Mulyawan IM (2019) Role of Ki67 protein in colorectal cancer. Int J Res Med Sci 7:644. https://doi.org/10.18203/2320-6012.ijrms20190374
Levin B, Lieberman DA, McFarland B et al (2008) Screening and surveillance for the early detection of colorectal cancer and adenomatous polyps, 2008: a joint guideline from the American Cancer Society, the US Multi-Society Task Force on Colorectal Cancer, and the American College of Radiology. CA Cancer J Clin 58:130–160. https://doi.org/10.3322/ca.2007.0018
Bibbò S, Ianiro G, Giorgio V et al (2016) The role of diet on gut microbiota composition. Eur Rev Med Pharmacol Sci 20:4742–4749
Sekirov I, Russell SL, Antunes LC, Finlay BB (2010) Gut microbiota in health and disease. Physiol Rev 90:859–904. https://doi.org/10.1152/physrev.00045.2009
Turnbaugh PJ, Ley RE, Hamady M et al (2007) The human microbiome project. Nature 449:804–810. https://doi.org/10.1038/nature06244
Dzutsev A, Goldszmid RS, Viaud S et al (2015) The role of the microbiota in inflammation, carcinogenesis, and cancer therapy. Eur J Immunol 45:17–31. https://doi.org/10.1002/eji.201444972
Saus E, Iraola-Guzmán S, Willis JR et al (2019) Microbiome and colorectal cancer: roles in carcinogenesis and clinical potential. Mol Aspects Med 69:93–106. https://doi.org/10.1016/j.mam.2019.05.001
Schwabe RF, Jobin C (2013) The microbiome and cancer. Nat Rev Cancer 13:800–812. https://doi.org/10.1038/nrc3610
Elinav E, Nowarski R, Thaiss CA et al (2013) Inflammation-induced cancer: crosstalk between tumours, immune cells and microorganisms. Nat Rev Cancer 13:759–771. https://doi.org/10.1038/nrc3611
Kostic AD, Gevers D, Pedamallu CS et al (2012) Genomic analysis identifies association of Fusobacterium with colorectal carcinoma. Genome Res 22:292–298. https://doi.org/10.1101/gr.126573.111
Wu S, Rhee KJ, Albesiano E et al (2009) A human colonic commensal promotes colon tumorigenesis via activation of T helper type 17 T cell responses. Nat Med 15:1016–1022. https://doi.org/10.1038/nm.2015
Gill CIR, Rowland IR (2002) Diet and cancer: assessing the risk. Br J Nutr 88:s73–s87. https://doi.org/10.1079/bjn2002632
Ferlay J, Soerjomataram I, Dikshit R et al (2015) Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer 136:E359–E386. https://doi.org/10.1002/ijc.29210
Fearon EF, Vogelstein B (1990) A genetic model for colorectal tumorigenesis. G Ital Cardiol 19:170–172. https://doi.org/10.1016/0092-8674(90)90186-I
Kolligs FT, Nieman MT, Winer I et al (2002) ITF-2, a downstream target of the Wnt/TCF pathway, is activated in human cancers with β-catenin defects and promotes neoplastic transformation. Cancer Cell 1:145–155. https://doi.org/10.1016/s1535-6108(02)00035-1
Stoffel EM, Kastrinos F (2014) Familial colorectal cancer, beyond lynch syndrome. Clin Gastroenterol Hepatol 12:1059–1068. https://doi.org/10.1016/j.cgh.2013.08.015
Levin B, Lieberman DA, McFarland B et al (2008) Screening and surveillance for the early detection of colorectal cancer and adenomatous polyps, 2008: a joint guideline from the American Cancer Society, the US Multi-Society Task Force on Colorectal Cancer, and the American College of Radiology. Gastroenterology 134:1570–1595. https://doi.org/10.1053/j.gastro.2008.02.002
Moskal A, Freisling H, Byrnes G et al (2016) Main nutrient patterns and colorectal cancer risk in the European Prospective Investigation into Cancer and Nutrition study. Br J Cancer 115:1430–1440. https://doi.org/10.1038/bjc.2016.334
Favoriti P, Carbone G, Greco M et al (2016) Worldwide burden of colorectal cancer: a review. Updates Surg 68:7–11. https://doi.org/10.1007/s13304-016-0359-y
Simon K (2016) Colorectal cancer development and advances in screening. Clin Interv Aging 11:967–976. https://doi.org/10.2147/CIA.S109285
Mackie RI, Sghir A, Gaskins HR (1999) Developmental microbial ecology of the neonatal gastrointestinal tract. Am J Clin Nutr 69. https://doi.org/10.1093/ajcn/69.5.1035s
Mandar R, Mikelsaar M (1996) Transmission of mothers microflora to newborn at birth. 68–70. https://doi.org/10.1159/000244275
Turnbaugh PJ, Bäckhed F, Fulton L, Gordon JI (2008) Diet-induced obesity is linked to marked but reversible alterations in the mouse distal gut microbiome. Cell Host Microbe 3:213–223. https://doi.org/10.1016/j.chom.2008.02.015
Eckburg PB, Bik EM, Bernstein CN et al (2005) Microbiology: diversity of the human intestinal microbial flora. Science (80- ) 308:1635–1638. https://doi.org/10.1126/science.1110591
Gagnière J, Raisch J, Veziant J et al (2016) Gut microbiota imbalance and colorectal cancer. World J Gastroenterol 22:501–518. https://doi.org/10.3748/wjg.v22.i2.501
Tlaskalová-Hogenová H, Štěpánková R, Hudcovic T et al (2004) Commensal bacteria (normal microflora), mucosal immunity and chronic inflammatory and autoimmune diseases. Immunol Lett 93:97–108. https://doi.org/10.1016/j.imlet.2004.02.005
Frank DN, St. Amand AL, Feldman RA, et al (2007) Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proc Natl Acad Sci U S A 104:13780–13785. https://doi.org/10.1073/pnas.0706625104
Swidsinski A, Loening-Baucke V, Lochs H, Hale LP (2005) Spatial organization of bacterial flora in normal and inflamed intestine: a fluorescence in situ hybridization study in mice. World J Gastroenterol 11:1131–1140. https://doi.org/10.3748/wjg.v11.i8.1131
Zoetendal EG, Von Wright A, Vilpponen-Salmela T et al (2002) Mucosa-associated bacteria in the human gastrointestinal tract are uniformly distributed along the colon and differ from the community recovered from feces. Appl Environ Microbiol 68:3401–3407. https://doi.org/10.1128/AEM.68.7.3401-3407.2002
Carroll IM, Chang YH, Park J et al (2010) Luminal and mucosal-associated intestinal microbiota in patients with diarrhea-predominant irritable bowel syndrome. Gut Pathog 2:1–9. https://doi.org/10.1186/1757-4749-2-19
Tap J, Derrien M, Törnblom H et al (2017) Identification of an intestinal microbiota signature associated with severity of irritable bowel syndrome. Gastroenterology 152:111-123.e8. https://doi.org/10.1053/j.gastro.2016.09.049
Rangel I, Sundin J, Fuentes S et al (2015) The relationship between faecal-associated and mucosal-associated microbiota in irritable bowel syndrome patients and healthy subjects. Aliment Pharmacol Ther 42:1211–1221. https://doi.org/10.1111/apt.13399
Zmora N, Zilberman-Schapira G, Suez J et al (2018) Personalized gut mucosal colonization resistance to empiric probiotics is associated with unique host and microbiome features. Cell 174:1388-1405.e21. https://doi.org/10.1016/j.cell.2018.08.041
Cardona S, Eck A, Cassellas M et al (2012) Storage conditions of intestinal microbiota matter in metagenomic analysis for standardization/RNA and DNA degradation/Metagenomics/16S ribosomal RNA Background. BMC Microbiol 12:1. https://doi.org/10.1186/1471-2180-12-158
Flemer B, Lynch DB, Brown JMR et al (2017) Tumour-associated and non-tumour-associated microbiota in colorectal cancer. Gut 66:633–643. https://doi.org/10.1136/gutjnl-2015-309595
Yu J, Feng Q, Wong SH et al (2017) Metagenomic analysis of faecal microbiome as a tool towards targeted non-invasive biomarkers for colorectal cancer. Gut 66:70–78. https://doi.org/10.1136/gutjnl-2015-309800
Harrell L, Wang Y, Antonopoulos D et al (2012) Standard colonic lavage alters the natural state of mucosal-associated microbiota in the human colon. PLoS ONE 7. https://doi.org/10.1371/journal.pone.0032545
Budding AE, Grasman ME, Eck A et al (2014) Rectal swabs for analysis of the intestinal microbiota. PLoS ONE 9:5–12. https://doi.org/10.1371/journal.pone.0101344
Namsolleck P, Thiel R, Lawson P et al (2004) Molecular methods for the analysis of gut microbiota. Microb Ecol Health Dis 16:71–85. https://doi.org/10.1080/08910600410032367
Li J, Jia H, Cai X et al (2014) An integrated catalog of reference genes in the human gut microbiome. Nat Biotechnol 32:834–841. https://doi.org/10.1038/nbt.2942
Huttenhower C, Gevers D, Knight R et al (2012) Structure, function and diversity of the healthy human microbiome. Nature 486:207–214. https://doi.org/10.1038/nature11234
Schmidt TSB, Raes J, Bork P (2018) The human gut microbiome: from association to modulation. Cell 172:1198–1215. https://doi.org/10.1016/j.cell.2018.02.044
Anderson S (1984) Shotgun DNA sequencing using cloned DNase I-generated fragments. Methods 12:8235–8251. https://doi.org/10.1093/nar/9.13.3015
Weisburg WG, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703. https://doi.org/10.1128/jb.173.2.697-703.1991
Ranjan R, Rani A, McGee HS, Perkins DL (2016) Analysis of the microbiome: advantages of whole genome shotgun versus 16S amplicon sequencing. Physiol Behav 63:1–18. https://doi.org/10.1016/j.bbrc.2015.12.083
Aguiar-Pulido V, Huang W, Suarez-Ulloa V et al (2016) Metagenomics, metatranscriptomics, and metabolomics approaches for microbiome analysis. Evol Bioinforma 12:5–16. https://doi.org/10.4137/EBO.S36436
Ranjan R, Rani A, Finn PW, Perkins DL (2018) Multiomic strategies reveal diversity and important functional aspects of human gut microbiome. Biomed Res Int 2018. https://doi.org/10.1155/2018/6074918
Armitage EG, Barbas C (2014) Metabolomics in cancer biomarker discovery: current trends and future perspectives. J Pharm Biomed Anal 87:1–11. https://doi.org/10.1016/j.jpba.2013.08.041
Farshidfar F, Weljie AM, Kopciuk KA et al (2016) A validated metabolomic signature for colorectal cancer: exploration of the clinical value of metabolomics. Br J Cancer 115:848–857. https://doi.org/10.1038/bjc.2016.243
Liesenfeld DB, Grapov D, Fahrmann JF et al (2015) Metabolomics and transcriptomics identify pathway differences between visceral and subcutaneous adipose tissue in colorectal cancer patients: the ColoCare study. Am J Clin Nutr 102:433–443. https://doi.org/10.3945/ajcn.114.103804
Qiu Y, Cai G, Zhou B et al (2014) A distinct metabolic signature of human colorectal cancer with prognostic potential. Clin Cancer Res 20:2136–2146. https://doi.org/10.1158/1078-0432.CCR-13-1939
Tan B, Qiu Y, Zou X et al (2013) Metabonomics identifies serum metabolite markers of colorectal cancer. J Proteome Res 12:3000–3009. https://doi.org/10.1021/pr400337b
Uchiyama K, Yagi N, Mizushima K et al (2017) Serum metabolomics analysis for early detection of colorectal cancer. J Gastroenterol 52:677–694. https://doi.org/10.1007/s00535-016-1261-6
Fleming M, Ravula S, Tatishchev SF, Wang HL (2012) Colorectal carcinoma: pathologic aspects. J Gastrointest Oncol 3:153–173. https://doi.org/10.3978/j.issn.2078-6891.2012.030
Carethers JM, Jung BH (2015) Genetics and genetic biomarkers in sporadic colorectal cancer. Gastroenterology 149:1177-1190.e3. https://doi.org/10.1053/j.gastro.2015.06.047
Cheng Y, Ling Z, Li L (2020) The intestinal microbiota and colorectal cancer. Front Immunol 11:1–13. https://doi.org/10.3389/fimmu.2020.615056
Marchesi JR, Dutilh BE, Hall N et al (2011) Towards the human colorectal cancer microbiome. PLoS ONE 6. https://doi.org/10.1371/journal.pone.0020447
Gao Z, Guo B, Gao R et al (2015) Microbiota disbiosis is associated with colorectal cancer. Front Microbiol 6:1–9. https://doi.org/10.3389/fmicb.2015.00020
Gao R, Kong C, Huang L et al (2017) Mucosa-associated microbiota signature in colorectal cancer. Eur J Clin Microbiol Infect Dis 36:2073–2083. https://doi.org/10.1007/s10096-017-3026-4
Feng Q, Liang S, Jia et al H (2015) Gut microbiome development along the colorectal adenoma-carcinoma sequence. Nat Commun 6. https://doi.org/10.1038/ncomms7528
Shang FM, Liu HL (2018) Fusobacterium nucleatum and colorectal cancer: a review. World J Gastrointest Oncol 10:71–81. https://doi.org/10.4251/wjgo.v10.i3.71
Wang T, Cai G, Qiu Y et al (2012) Structural segregation of gut microbiota between colorectal cancer patients and healthy volunteers. ISME J 6:320–329. https://doi.org/10.1038/ismej.2011.109
Yu TC, Guo F, Yu Y et al (2017) Fusobacterium nucleatum promotes chemoresistance to colorectal cancer by modulating autophagy. Cell 170:548-563.e16. https://doi.org/10.1016/j.cell.2017.07.008
Zhang S, Yang Y, Weng W et al (2019) Fusobacterium nucleatum promotes chemoresistance to 5-fluorouracil by upregulation of BIRC3 expression in colorectal caner. J Exp Clin Cancer Res 38:1–13. https://doi.org/10.1186/s13046-018-0985-y
Arrieta MC, Bistritz L, Meddings JB (2006) Alterations in intestinal permeability Gut 55:1512–1520. https://doi.org/10.1136/gut.2005.085373
Bischoff SC, Barbara G, Buurman W et al (2014) Intestinal permeability - a new target for disease prevention and therapy. BMC Gastroenterol 14:1–25. https://doi.org/10.1186/s12876-014-0189-7
Owyang C, Wu GD (2014) The gut microbiome in health and disease. Gastroenterology 146:1433–1436. https://doi.org/10.1053/j.gastro.2014.03.032
Maloy KJ, Powrie F (2001) Regulatory T cells in the control of immune pathology. Nat Immunol 2:816–822. https://doi.org/10.1038/ni0901-816
Lamont RJ, Hajishengallis G (2015) Polymicrobial synergy and dysbiosis in inflammatory disease. Trends Mol Med 21:172–183. https://doi.org/10.1016/j.molmed.2014.11.004
Tibbs TN, Lopez LR, Arthur JC (2019) The influence of the microbiota on immune development, chronic inflammation, and cancer in the context of aging. Microb Cell 6:324–334. https://doi.org/10.15698/mic2019.08.685
Goodwin AC, Destefano Shields CE, Wu S et al (2011) Polyamine catabolism contributes to enterotoxigenic Bacteroides fragilis-induced colon tumorigenesis. Proc Natl Acad Sci U S A 108:15354–15359. https://doi.org/10.1073/pnas.1010203108
Wu S, Morin PJ, Maouyo D, Sears CL (2003) Bacteroides fragilis enterotoxin induces c-Myc expression and cellular proliferation. Gastroenterology 124:392–400. https://doi.org/10.1053/gast.2003.50047
Rubinstein MR, Wang X, Liu W et al (2013) Fusobacterium nucleatum promotes colorectal carcinogenesis by modulating E-cadherin/β-catenin signaling via its FadA adhesin. Cell Host Microbe 14:195–206. https://doi.org/10.1016/j.chom.2013.07.012
Kostic AD, Chun E, Robertson L et al (2013) Fusobacterium nucleatum potentiates intestinal tumorigenesis and modulates the tumor-immune microenvironment. Cell Host Microbe 14:207–215. https://doi.org/10.1016/j.chom.2013.07.007
Cheng WY, Wu CY, Yu J (2020) The role of gut microbiota in cancer treatment: friend or foe? Gut 69:1867–1876. https://doi.org/10.1136/gutjnl-2020-321153
Navarro M, Nicolas A, Ferrandez A, Lanas A (2017) Colorectal cancer population screening programs worldwide in 2016: an update. World J Gastroenterol 23:3632–3642. https://doi.org/10.3748/wjg.v23.i20.3632
Zhang J, Cheng Z, Ma Y et al (2017) Effectiveness of screening modalities in colorectal cancer: a network meta-analysis. Clin Colorectal Cancer 16:252–263. https://doi.org/10.1016/j.clcc.2017.03.018
Konstantinov SR, Kuipers EJ, Peppelenbosch MP (2013) Functional genomic analyses of the gut microbiota for crc screening. Nat Rev Gastroenterol Hepatol 10:741–745. https://doi.org/10.1038/nrgastro.2013.178
Zeller G, Tap J, Voigt AY et al (2014) Potential of fecal microbiota for early‐stage detection of colorectal cancer. Mol Syst Biol 10:766. https://doi.org/10.15252/msb.20145645
Rezasoltani S, Sharafkhah M, Asadzadeh Aghdaei H et al (2018) Applying simple linear combination, multiple logistic and factor analysis methods for candidate fecal bacteria as novel biomarkers for early detection of adenomatous polyps and colon cancer. J Microbiol Methods 155:82–88. https://doi.org/10.1016/j.mimet.2018.11.007
Ai L, Tian H, Chen Z et al (2017) Systematic evaluation of supervised classifiers for fecal microbiota-based prediction of colorectal cancer. Oncotarget 8:9546–9556. https://doi.org/10.18632/oncotarget.14488
Castellarin M, Warren R, Douglas F, Dreolini L (2018) Fusobacterium nucleatum infection is prevalent in human colorectal carcinoma. Elife 7:299–306. https://doi.org/10.1101/gr.126516.111
Flanagan L, Schmid J, Ebert M et al (2014) Fusobacterium nucleatum associates with stages of colorectal neoplasia development, colorectal cancer and disease outcome. Eur J Clin Microbiol Infect Dis 33:1381–1390. https://doi.org/10.1007/s10096-014-2081-3
Dai Z, Coker OO, Nakatsu G et al (2018) Multi-cohort analysis of colorectal cancer metagenome identified altered bacteria across populations and universal bacterial markers. Microbiome 6:70. https://doi.org/10.1186/s40168-018-0451-2
Shah MS, DeSantis TZ, Weinmaier T et al (2018) Leveraging sequence-based faecal microbial community survey data to identify a composite biomarker for colorectal cancer. Gut 67:882–891. https://doi.org/10.1136/gutjnl-2016-313189
Balish E, Warner T (2002) Enterococcus faecalis induces inflammatory bowel disease in interleukin-10 knockout mice. Am J Pathol 160:2253–2257. https://doi.org/10.1016/S0002-9440(10)61172-8
D’asheesh TA, Hussen BM, Al-Marzoqi AH, Ghasemian A (2020) Assessment of oncogenic role of intestinal microbiota in colorectal cancer patients. J Gastrointest Cancer. https://doi.org/10.1007/s12029-020-00531-8
Thomas DW, Greer FR, Bhatia JJS et al (2010) Clinical report - probiotics and prebiotics in pediatrics. Pediatrics 126:1217–1231. https://doi.org/10.1542/peds.2010-2548
Isolauri E, Salminen S, Ouwehand AC (2019) Probiotics. Compr Biotechnol 18:649–661. https://doi.org/10.1053/ybega.2004.443
Hutkins RW, Krumbeck JA, Bindels LB et al (2016) Prebiotics – why definitions matter. Curr Oin Biotechnol 37:1–7. https://doi.org/10.1016/j.copbio.2015.09.001
Lynch SV, Pedersen O (2016) The human intestinal microbiome in health and disease. N Engl J Med 375:2369–2379. https://doi.org/10.1056/nejmra1600266
Juul FE, Garborg K, Bretthauer M et al (2018) Fecal microbiota transplantation for primary clostridium difficile infection. N Engl J Med 378:2535–2536. https://doi.org/10.1056/nejmc1803103
Filip M, Tzaneva V, Dumitrascu DANL (2018) Fecal transplantation : digestive and extradigestive clinical applications Intestinal microbiota. 91:259–265. https://doi.org/10.15386/cjmed-946
Rosshart SP, Vassallo BG, Angeletti D et al (2017) Wild mouse gut microbiota promotes host fitness and improves disease resistance article wild mouse gut microbiota promotes host fitness and improves disease resistance. Cell 1–14. https://doi.org/10.1016/j.cell.2017.09.016
Wong SH, Zhao L, Zhang X et al (2017) Gavage of fecal samples from patients with colorectal cancer promotes intestinal carcinogenesis in germ-free and conventional mice. Gastroenterology 153:1621-1633.e6. https://doi.org/10.1053/j.gastro.2017.08.022
Sbahi H, Di Palma JA (2016) Faecal microbiota transplantation: applications and limitations in treating gastrointestinal disorders. BMJ Open Gastroenterol 3:1–7. https://doi.org/10.1136/bmjgast-2016-000087
Bullman S, Pedamallu CS, Sicinska E et al (2017) Analysis of Fusobacterium persistence and antibiotic response in colorectal cancer. Science (80- ) 358:1443–1448. https://doi.org/10.1126/science.aal5240
Cao H, Xu M, Dong W et al (2017) Secondary bile acid-induced dysbiosis promotes intestinal carcinogenesis. Int J Cancer 140:2545–2556. https://doi.org/10.1002/ijc.30643
Donohoe DR, Holley D, Collins LB et al (2014) A gnotobiotic mouse model demonstrates that dietary fiber protects against colorectal tumorigenesis in a microbiota- and butyrate-dependent manner. Cancer Discov 4:1387–1397. https://doi.org/10.1158/2159-8290.CD-14-0501
Ho CL, Tan HQ, Chua KJ et al (2018) Engineered commensal microbes for diet-mediated colorectal-cancer chemoprevention. Nat Biomed Eng 2:27–37. https://doi.org/10.1038/s41551-017-0181-y
Camp ER, Patterson LD, Kester M, Voelkel-Johnson C (2017) Therapeutic implications of bioactive sphingolipids: a focus on colorectal cancer. Cancer Biol Ther 18:640–650. https://doi.org/10.1080/15384047.2017.1345396
West NR, Powrie F (2015) Immunotherapy not working? Check your microbiota. Cancer Cell 28:687–689. https://doi.org/10.1016/j.ccell.2015.11.010
Raman M, Ambalam P, Kondepudi KK et al (2013) Potential of probiotics, prebiotics and synbiotics for management of colorectal cancer. Gut Microbes 4:181–192. https://doi.org/10.4161/gmic.23919
Escamilla J, Lane MA, Maitin V (2012) Cell-free supernatants from probiotic lactobacillus casei and lactobacillus rhamnosus GG decrease colon cancer cell invasion in vitro. Nutr Cancer 64:871–878. https://doi.org/10.1080/01635581.2012.700758
Lin C, Cai X, Zhang J et al (2019) Role of gut microbiota in the development and treatment of colorectal cancer. Digestion 100:72–78. https://doi.org/10.1159/000494052
Chen ZF, Ai LY, Wang JL et al (2015) Probiotics Clostridium butyricum and Bacillus subtilis ameliorate intestinal tumorigenesis. Future Microbiol 10:1433–1445. https://doi.org/10.2217/fmb.15.66
Lenoir M, del Carmen S, Cortes-Perez NG et al (2016) Lactobacillus casei BL23 regulates Treg and Th17 T-cell populations and reduces DMH-associated colorectal cancer. J Gastroenterol 51:862–873. https://doi.org/10.1007/s00535-015-1158-9
Konishi H, Fujiya M, Tanaka H et al (2016) Probiotic-derived ferrichrome inhibits colon cancer progression via JNK-mediated apoptosis. Nat Commun 7. https://doi.org/10.1038/ncomms12365
Chang CW, Liu CY, Lee HC et al (2018) Lactobacillus casei variety rhamnosus probiotic preventively attenuates 5-fluorouracil/oxaliplatin-induced intestinal injury in a syngeneic colorectal cancer model. Front Microbiol 9:1–15. https://doi.org/10.3389/fmicb.2018.00983
Liang S, Xu L, Zhang D, Wu Z (2016) Effect of probiotics on small intestinal bacterial overgrowth in patients with gastric and colorectal cancer. Turkish J Gastroenterol 27:227–232. https://doi.org/10.5152/tjg.2016.15375
Cousin FJ, Jouan-Lanhouet S, Théret N et al (2016) The probiotic Propionibacterium freudenreichii as a new adjuvant for TRAIL-based therapy in colorectal cancer. Oncotarget 7:7161–7178. https://doi.org/10.18632/oncotarget.6881
Pandey KR, Naik SR, Vakil BV (2015) Probiotics, prebiotics and synbiotics- a review. J Food Sci Technol 52:7577–7587. https://doi.org/10.1007/s13197-015-1921-1
Bozkurt HS, Quigley EM, Kara B (2019) Bifidobacterium animalis subspecies lactis engineered to produce mycosporin-like amino acids in colorectal cancer prevention. SAGE Open Med 7:205031211982578. https://doi.org/10.1177/2050312119825784
Ding C, Tang W, Fan X, Wu G (2018) Intestinal microbiota: a novel perspective in colorectal cancer biotherapeutics. Onco Targets Ther 11:4797–4810. https://doi.org/10.2147/OTT.S170626
Zitvogel L, Ma Y, Raoult D et al (2018) The microbiome in cancer immunotherapy: diagnostic tools and therapeutic strategies. Science (80- ) 359:1366–1370. https://doi.org/10.1126/science.aar6918
Routy B, Le Chatelier E, Derosa L et al (2018) Gut microbiome influences efficacy of PD-1-based immunotherapy against epithelial tumors. Science (80- ) 359:91–97. https://doi.org/10.1126/science.aan3706
Matson V, Fessler J, Bao R et al (2018) The commensal microbiome is associated with anti-PD-1 efficacy in metastatic melanoma patients. Science (80- ) 359:104–108. https://doi.org/10.1126/science.aao3290
Sivan A, Corrales L, Hubert N et al (2015) Commensal Bifidobacterium promotes antitumor immunity and facilitates anti-PD-L1 efficacy. Science (80- ) 350:1084–1089. https://doi.org/10.1126/science.aac4255
Vétizou M, Pitt JM, Daillère R et al (2015) Anticancer immunotherapy by CTLA-4 blockade relies on the gut microbiota. Science (80- ) 350:1079–1084. https://doi.org/10.1126/science.aad1329
Daillère R, Vétizou M, Waldschmitt N et al (2016) Enterococcus hirae and Barnesiella intestinihominis facilitate cyclophosphamide-induced therapeutic immunomodulatory effects. Immunity 45:931–943. https://doi.org/10.1016/j.immuni.2016.09.009
Lee V, Le DT (2016) Efficacy of PD-1 blockade in tumors with MMR deficiency. Immunotherapy 8:1–3. https://doi.org/10.2217/imt.15.97
Goyle S, Maraveyas A (2006) Chemotherapy for colorectal cancer. Dig Surg 22:401–414. https://doi.org/10.1159/000091441
Miwa M, Ura M, Nishida M et al (1998) Design of a novel oral fluoropyrimidine carbamate, capecitabine, which generates 5 fluorouracil selectively in tumours by enzymes concentrated in human liver and cancer tissue. Eur J Cancer 34:1274–1281. https://doi.org/10.1016/S0959-8049(98)00058-6
Walko CM, Lindley C (2005) Capecitabine: a review. Clin Ther 27:23–44. https://doi.org/10.1016/j.clinthera.2005.01.005
Kelland L (2007) The resurgence of platinum-based cancer chemotherapy. Nat Rev Cancer 7:573–584. https://doi.org/10.1038/nrc2167
Fuchs C, Mitchell EP, Hoff PM (2006) Irinotecan in the treatment of colorectal cancer. Cancer Treat Rev 32:491–503. https://doi.org/10.1016/j.ctrv.2006.07.001
Kciuk M, Marciniak B, Kontek R (2020) Irinotecan—still an important player in cancer chemotherapy: A comprehensive overview. Int J Mol Sci 21:1–21. https://doi.org/10.3390/ijms21144919
Cremolini C, Schirripa M, Antoniotti C et al (2015) First-line chemotherapy for mCRC-a review and evidence-based algorithm. Nat Rev Clin Oncol 12:607–619. https://doi.org/10.1038/nrclinonc.2015.129
Marin JJG, Sanchez De Medina F, Castão B et al (2012) Chemoprevention, chemotherapy, and chemoresistance in colorectal cancer. Drug Metab Rev 44:148–172. https://doi.org/10.3109/03602532.2011.638303
Lin M, Zhang J, Chen X (2018) Bioactive flavonoids in Moringa oleifera and their health-promoting properties. J Funct Foods 47:469–479. https://doi.org/10.1016/j.jff.2018.06.011
Raghavan R, Koyande N, Beher R et al (2022) Plakophilin3 loss leads to increased adenoma formation and rectal prolapse in APCmin mice. Biochem Biophys Res Commun 586:14–19. https://doi.org/10.1016/j.bbrc.2021.11.071
Chaudhary N, Choudhary BS, Shah SG et al (2021) Lipocalin 2 expression promotes tumor progression and therapy resistance by inhibiting ferroptosis in colorectal cancer. Int J Cancer 149:1495–1511. https://doi.org/10.1002/ijc.33711
Cammareri P, Scopelliti A, Todaro M et al (2010) Aurora-A is essential for the tumorigenic capacity and chemoresistance of colorectal cancer stem cells. Cancer Res 70:4655–4665. https://doi.org/10.1158/0008-5472.CAN-09-3953
Tsoumas D, Nikou S, Giannopoulou E et al (2018) ILK expression in colorectal cancer is associated with EMT, cancer stem cell markers and chemoresistance. Cancer Genom Proteom 15:127–141. https://doi.org/10.21873/cgp.20071
Wasserman I, Lee LH, Ogino S et al (2018) (SMAD4 loss in colorectal cancer patients correlates with recurrence, loss of immune infiltrate, and chemoresistance. 2015 IEEE Summer Top Meet Ser SUM 2015 10:1–13. https://doi.org/10.1158/1078-0432.CCR-18-1726
Izumi D, Ishimoto T, Miyake K et al (2017) Colorectal cancer stem cells acquire chemoresistance through the upregulation of F-Box/WD repeat-containing protein 7 and the consequent degradation of c-Myc. Stem Cells 35:2027–2036. https://doi.org/10.1002/stem.2668
Iida N, Dzutsev A, Stewart CA et al (2013) Commensal bacteria control cancer response to therapy by modulating the tumor microenvironment. Science (80- ) 342:967–970. https://doi.org/10.1126/science.1240527
Yuan L, Zhang S, Li H et al (2018) The influence of gut microbiota dysbiosis to the efficacy of 5-fluorouracil treatment on colorectal cancer. Biomed Pharmacother 108:184–193. https://doi.org/10.1016/j.biopha.2018.08.165
Ramos A, Hemann MT (2017) Drugs, bugs, and cancer: Fusobacterium nucleatum promotes chemoresistance in colorectal cancer. Cell 170:411–413. https://doi.org/10.1016/j.cell.2017.07.018
Kelly MG, Alvero AB, Chen R et al (2006) TLR-4 signaling promotes tumor growth and paclitaxel chemoresistance in ovarian cancer. Cancer Res 66:3859–3868. https://doi.org/10.1158/0008-5472.CAN-05-3948
Choy ATF, Carnevale I, Coppola S et al (2018) The microbiome of pancreatic cancer: from molecular diagnostics to new therapeutic approaches to overcome chemoresistance caused by metabolic inactivation of gemcitabine. Expert Rev Mol Diagn 18:1005–1009. https://doi.org/10.1080/14737159.2018.1544495
Vande VJ, Sabuncuoǧlu S, Noppen S et al (2014) Nucleoside-catabolizing enzymes in mycoplasma-infected tumor cell cultures compromise the cytostatic activity of the anticancer drug gemcitabine. J Biol Chem 289:13054–13065. https://doi.org/10.1074/jbc.M114.558924
Lehouritis P, Cummins J, Stanton M et al (2015) Local bacteria affect the efficacy of chemotherapeutic drugs. Sci Rep 5:1–12. https://doi.org/10.1038/srep14554
Vande Voorde J, Vervaeke P, Liekens S, Balzarini J (2015) Mycoplasma hyorhinis-encoded cytidine deaminase efficiently inactivates cytosine-based anticancer drugs. FEBS Open Bio 5:634–639. https://doi.org/10.1016/j.fob.2015.07.007
Liu Y, Baba Y, Ishimoto T et al (2020) Fusobacterium nucleatum confers chemoresistance by modulating autophagy in oesophageal squamous cell carcinoma. Br J Cancer. https://doi.org/10.1038/s41416-020-01198-5
Geller LT, Barzily-rokni M, Danino T et al (2017) Potential role of intratumor bacteria in mediating tumor resistance to the chemotherapeutic drug gemcitabine Leore 1160:1156–1160. https://doi.org/10.1126/science.aah5043
Solomon SL, Oliver B, Disease C (2014) Antibiotic resistance threats in the United States: stepping back from the brink. American family physician, 89(12), 938–941.
Kumar A, Schweizer HP (2005) Bacterial resistance to antibiotics: active efflux and reduced uptake. Adv Drug Deliv Rev 57:1486–1513. https://doi.org/10.1016/j.addr.2005.04.004
Ruiz N, Montero T, Hernandez-Borrell J, Viñas M (2003) The role of Serratia marcescens porins in antibiotic resistance. Microb Drug Resist 9:257–264. https://doi.org/10.1089/107662903322286463
Abraham EP, Chain E (1940) An enzyme from bacteria able to. Nature 146:837. https://doi.org/10.1038/146837a0
Poole K (2004) Resistance to β-lactam antibiotics. Cell Mol Life Sci 61:2200–2223. https://doi.org/10.1007/s00018-004-4060-9
Levy SB (2002) Active efflux, a common mechanism for biocide and antibiotic resistance. Symp Ser Soc Appl Microbiol 65–71. https://doi.org/10.1046/j.1365-2672.92.5s1.4.x
Poole K (2007) Efflux pumps as antimicrobial resistance mechanisms. Ann Med 39:162–176. https://doi.org/10.1080/07853890701195262
Roberts MC (2005) Update on acquired tetracycline resistance genes. FEMS Microbiol Lett 245:195–203. https://doi.org/10.1016/j.femsle.2005.02.034
Schelenz S, Nwaka D, Hunter PR (2013) Longitudinal surveillance of bacteraemia in haematology and oncology patients at a uk cancer centre and the impact of ciprofloxacin use on antimicrobial resistance. J Antimicrob Chemother 68:1431–1438. https://doi.org/10.1093/jac/dkt002
Munita JM, Arias CA (2015) Mechanisms of antibiotic resistance. Annu Rep Med Chem 17:119–127. https://doi.org/10.1128/microbiolspec.VMBF-0016-2015
Iizumi T, Battaglia T, Ruiz V, Perez Perez GI (2017) Gut microbiome and antibiotics. Arch Med Res 48:727–734. https://doi.org/10.1016/j.arcmed.2017.11.004
Holman DB, Chénie MR (2012) Antimicrobial use in swine production and its effect on the swine gut microbiota and antimicrobial resistance. Journal 348–349. https://doi.org/10.1139/cjm-2015-0239
Korpela K, Salonen A, Virta LJ et al (2016) Intestinal microbiome is related to lifetime antibiotic use in Finnish pre-school children. Nat Commun 7. https://doi.org/10.1038/ncomms10410
Rutter MD, Nickerson C, Rees CJ, et al (2014) Risk factors for adverse events related to polypectomy in the english bowel cancer screening programme. Endoscopy 46:90–97. https://doi.org/10.1055/s-0033-1344987
Lee J, LILES E, Bent S (2011) Accuracy of Fecal Immunochemical Tests for Colorectal Cancer: Systematic Review and Meta-analysis. Bone 23:1–7. https://doi.org/10.7326/M13-1484
Robertson DJ, Lee JK, Boland CR, et al (2017) Recommendations on fecal immunochemical testing to screen for colorectal neoplasia: A consensus statement by the US multi-society task force on colorectal cancer. Gastroenterology 152:1217-1237.e3. https://doi.org/10.1053/j.gastro.2016.08.053
Coghlin C, Murray G (2015) Biomarkers of colorectal cancer: recent advances and future challenges. 1–24. https://doi.org/10.1002/prca.201400082
Pouncey AL, Scott AJ, Alexander JL et al (2018) Gut microbiota, chemotherapy and the host: the influence of the gut microbiota on cancer treatment. ecancermedicalscience 12:1–9. https://doi.org/10.3332/ecancer.2018.868
Acknowledgements
The authors are thankful to Dr. Sangeetha CJ for her constant support and valuable comments regarding the content of the article. The authors also wish to acknowledge Mr. Vikas Sonkar, Mr. Arun Kashyap, and Miss Rupali Srivastava for language correction and proofreading. All the figures were created using Biorender.com.
Funding
The study was supported by the Indo-UK project entitled “AMRflows UK-India project: antimicrobials and resistance from manufacturing flows to people: joined up experiments, mathematical modelling and risk analysis” (AMRflows; grant number DBT/CE/F064/2020–21/G307) funded by the Department of Biotechnology (Government of India).
Author information
Authors and Affiliations
Contributions
Navami Koyande: conceptualization; formal analysis; methodology; investigation: writing-original draft; and writing-review and editing. Madhusree Gangopadhyay: writing-original draft and formal analysis. Shashidhar Thatikonda: funding acquisition; project administration and supervision. Aravind Kumar Rengan: resources; project administration; and supervision.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Koyande, N., Gangopadhyay, M., Thatikonda, S. et al. The role of gut microbiota in the development of colorectal cancer: a review. Int J Colorectal Dis 37, 1509–1523 (2022). https://doi.org/10.1007/s00384-022-04192-w
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00384-022-04192-w