Abstract
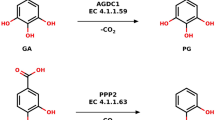
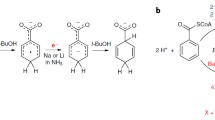
Benzoic acid decarboxylases offer an elegant alternative to CO2 fixation by reverse reaction-carboxylation, which is named the bio-Kolbe–Schmitt reaction, but they are unfavorable to carboxylation. Enhancing the carboxylation efficiency of reversible benzoic acid decarboxylases is restricted by the unexplained carboxylation mechanisms. The direction of reversible enzyme catalytic reactions depends on whether catalytic residues at the active center of the enzyme are protonated, which is subjected by the pH. Therefore, the forward and reverse reactions could be separated at different pH values. Reversible 2,3-dihydroxybenzoate acid decarboxylase undergoes decarboxylation at pH 5.0 and carboxylation at pH 8.6. However, it is unknown whether the interaction of enzymes with substrates and products in the forward and reverse reactions can be exploited to improve the catalytic activity of reversible enzymes in the unfavorable direction. Here, we identify a V-shaped tunnel of 2,3-dihydroxybenzoic acid decarboxylase from Aspergillus oryzae (2,3-DHBD_Ao) through which the substrate travels in the enzyme, and demonstrate that the side chain conformation of a tyrosine residue controls the entry and exit of substrate/product during reversible reactions. Together with the kinetic studies of the mutants, it is clarified that interactions between substrate/product traveling through the enzyme tunnel in 2,3-DHBD_Ao are direction-dependent. These results enrich the understanding of the interactions of substrates/products with macromolecular reversible enzymes in different reaction directions, thereby demonstrating a possible path for engineering decarboxylases with higher carboxylation efficiency.
Key points
• The residue Trp23 of 2,3-DHBD_Ao served as a switch to control the entry and exit of catechol
• A V-shaped tunnel of 2,3-DHBD_Ao for decarboxylation and carboxylation reactions was identified
• The results provide a promising strategy for engineering decarboxylases with direction-dependent residues inside the substrate/product traveling tunnel of the enzyme
Graphical Abstract








Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article (and its supplementary information files).
References
Aleku GA, Roberts GW, Titchiner GR, Leys D (2021) Synthetic enzyme-catalyzed CO2 fixation reactions. Chemsuschem 14:1781–1804. https://doi.org/10.1002/cssc.202100159
Appel AM, Bercaw JE, Bocarsly AB, Dobbek H, DuBois DL, Dupuis M, Ferry JG, Fujita E, Hille R, Kenis PJ (2013) Frontiers, opportunities, and challenges in biochemical and chemical catalysis of CO2 fixation. Chem Rev 113:6621–6658. https://doi.org/10.1021/cr300463y
Baroja-Fernández E, Muñoz FJ, Li J, Bahaji A, Almagro G, Montero M, Etxeberria E, Hidalgo M, Sesma MT, Pozueta-Romero J (2012) Sucrose synthase activity in the sus1/sus2/sus3/sus4 Arabidopsis mutant is sufficient to support normal cellulose and starch production. PNAS 109:321–326. https://doi.org/10.1073/pnas.1117099109
Cannon WR, Singleton SF, Benkovic SJ (1996) A perspective on biological catalysis. Nat Struct Mol Biol 3:821–833. https://doi.org/10.1038/nsb1096-821
Fan Y, Feng J, Yang M, Tan X, Fan H, Guo M, Wang B, Xue S (2021) CO2 (aq) concentration–dependent CO2 fixation via carboxylation by decarboxylase. Green Chem 23:4403–4409. https://doi.org/10.1039/d1gc00825k
Gao X, Wu M, Zhang W, Li C, Guo RT, Dai Y, Liu W, Mao S, Lu F, Qin HM (2021) Structural basis of salicylic acid decarboxylase reveals a unique substrate recognition mode and access channel. J Agric Food Chem 69:11616–11625. https://doi.org/10.1021/acs.jafc.1c04091
Goto M, Hayashi H, Miyahara I, Hirotsu K, Yoshida M, Oikawa T (2006) Crystal structures of nonoxidative zinc-dependent 2,6-dihydroxybenzoate (γ-resorcylate) decarboxylase from Rhizobium sp. strain MTP-10005. J Biol Chem 281:34365–34373. https://doi.org/10.1074/jbc.m607270200
Ienaga S, Kosaka S, Honda Y, Ishii Y, Kirimura K (2013) p-Aminosalicylic acid production by enzymatic Kolbe-Schmitt reaction using salicylic acid decarboxylases improved through site-directed mutagenesis. Bull Chem Soc Jpn 86:628–634. https://doi.org/10.1246/bcsj.20130006
Kirimura K, Gunji H, Wakayama R, Hattori T, Ishii Y (2010) Enzymatic Kolbe-Schmitt reaction to form salicylic acid from phenol: enzymatic characterization and gene identification of a novel enzyme, Trichosporon moniliiforme salicylic acid decarboxylase. Biochem Biophys Res Commun 394:279–284. https://doi.org/10.1016/j.bbrc.2010.02.154
Kokkonen P, Bednar D, Pinto G, Prokop Z, Damborsky J (2019) Engineering enzyme access tunnels. Biotechnolo Adv 37:107386. https://doi.org/10.1016/j.biotechadv.2019.04.008
Kourist R, Guterl JK, Miyamoto K, Sieber V (2014) Enzymatic decarboxylation-an emerging reaction for chemicals production from renewable resources. ChemCatChem 6:689–701. https://doi.org/10.1002/cctc.201300881
Li T, Huo L, Pulley C, Liu A (2012) Decarboxylation mechanisms in biological system. Bioorg Chem 43:2–14. https://doi.org/10.1016/j.bioorg.2012.03.001
Maier JA, Martinez C, Kasavajhala K, Wickstrom L, Hauser KE, Simmerling C (2015) ff14SB: improving the accuracy of protein side chain and backbone parameters from ff99SB. J Chem Theory Comput 11:3696–3713. https://doi.org/10.1021/acs.jctc.5b00255
Mobley DL, Dill KA (2009) Binding of small-molecule ligands to proteins:“what you see” is not always “what you get.” Structure 17:489–498. https://doi.org/10.1016/j.str.2009.02.010
Müller M, Sprenger GA, Pohl M (2013) C-C bond formation using ThDP-dependent lyases. Curr Opin Chem Biol 17:261–270. https://doi.org/10.1016/j.cbpa.2013.02.017
Payne KA, Marshall SA, Fisher K, Cliff MJ, Cannas DM, Yan C, Heyes DJ, Parker DA, Larrosa I, Leys D (2019) Enzymatic carboxylation of 2-furoic acid yields 2,5-furandicarboxylic acid (FDCA). ACS Catal 9:2854–2865. https://doi.org/10.1021/acscatal.8b04862
Plasch K, Hofer G, Keller W, Hay S, Heyes DJ, Dennig A, Glueck SM, Faber K (2018) Pressurized CO2 as a carboxylating agent for the biocatalytic ortho-carboxylation of resorcinol. Green Chem 20:1754–1759. https://doi.org/10.1039/c8gc00008e
Saleh T, Kalodimos CG (2017) Enzymes at work are enzymes in motion. Science 355:247–248. https://doi.org/10.1126/science.aal4632
Schmidt NG, Eger E, Kroutil W (2016) Building bridges: biocatalytic C-C-bond formation toward multifunctional products. ACS Catal 6:4286–4311. https://doi.org/10.1021/acscatal.6b00758
Sheng X, Himo F (2021) Mechanisms of metal-dependent non-redox decarboxylases from quantum chemical calculations. Comput Struct Biotechnol J 19:3176–3186. https://doi.org/10.1016/j.csbj.2021.05.044
Sheng X, Patskovsky Y, Vladimirova A, Bonanno JB, Almo SC, Himo F, Raushel FM (2017) Mechanism and structure of γ-resorcylate decarboxylase. Biochemistry 57:3167–3175. https://doi.org/10.1021/acs.biochem.7b01213
Song M, Zhang X, Liu W, Feng J, Cui Y, Yao P, Wang M, Guo RT, Wu Q, Zhu D (2020) 2,3-Dihydroxybenzoic acid decarboxylase from Fusarium oxysporum: crystal structures and substrate recognition mechanism. ChemBioChem 21:2950–2956. https://doi.org/10.1002/cbic.202000244
Su KH, Wu CT, Lin SW, Mori S, Liu WM, Yang HC (2021) Calculation of CYP450 protein–ligand binding and dissociation free energy paths. J Chem Phys 155:025101. https://doi.org/10.1063/5.0046169
Van Der Spoel D, Lindahl E, Hess B, Groenhof G, Mark AE, Berendsen HJ (2005) GROMACS: fast, flexible, and free. J Comput Chem 26:1701–1718. https://doi.org/10.1002/jcc.20291
Wuensch C, Gross J, Steinkellner G, Lyskowski A, Gruber K, Glueck SM, Faber K (2014) Regioselective ortho-carboxylation of phenols catalyzed by benzoic acid decarboxylases: a biocatalytic equivalent to the Kolbe-Schmitt reaction. RSC Adv 4:9673–9679. https://doi.org/10.1039/c3ra47719c
Zhang X, Ren J, Yao P, Gong R, Wang M, Wu Q, Zhu D (2018) Biochemical characterization and substrate profiling of a reversible 2,3-dihydroxybenzoic acid decarboxylase for biocatalytic Kolbe-Schmitt reaction. Enzyme Microb Technol 113:37–43. https://doi.org/10.1016/j.enzmictec.2018.02.008
Acknowledgements
We thank Shijun Zhong from School of Bioengineering, Dalian University of Technology, for his comment and suggestion on the mechanism analysis. We cordially thank the staff of beamline BL17U1 and BL18U1 at the Shanghai Synchrotron Radiation Facility, People’s Republic of China, for assistance in synchrotron X-ray data collection.
Funding
This study was financially supported by the National Natural Science Foundation of China (No. 21877110) and National Key Research and Development Program (No. 2021YFC2103702).
Author information
Authors and Affiliations
Contributions
Y. F. designed and conducted most of the experiments, analyzed the results, and wrote the manuscript. S. W. conducted the theoretical calculations. Y. F. and X. S. analyzed the data and wrote the manuscript. J. S. and X. L. provided the useful suggestions for experiments. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval
This article does not contain any studies with animals performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Supplementary file2 The following video is available free of charge. (MP4 2.88 MB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Fan, Y., Wu, S., Shi, J. et al. The catalytic mechanism of direction-dependent interactions for 2,3-dihydroxybenzoate decarboxylase. Appl Microbiol Biotechnol 107, 7451–7462 (2023). https://doi.org/10.1007/s00253-023-12813-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-023-12813-9