Abstract
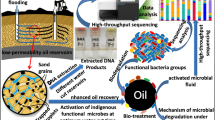
The role of indigenous microbial communities in residual oil extraction following a recovery process is not well understood. This study investigated the dynamics of resident microbial communities in oil-field simulating sand pack bioreactors after the polymer flooding stage resumed with waterflooding and explored their contribution to the oil extraction process. The microbial community succession was studied through high-throughput sequencing of 16S rRNA genes. The results revealed alternating dominance of minority populations, including Dietzia sps., Acinetobacter sps., Soehngenia sps., and Paracoccus sps., in each bioreactor following the flooding process. Additionally, the post-polymer waterflooding stage led to higher oil recovery, with hydroxyethylcellulose, tragacanth gum, and partially hydrolyzed polyacrylamide polymer-treated bioreactors yielding additional recovery of 4.36%, 5.39%, and 3.90% residual oil in place, respectively. The dominant microbial communities were previously reported to synthesize biosurfactants and emulsifiers, as well as degrade and utilize hydrocarbons, indicating their role in aiding the recovery process. However, the correlation analysis of the most abundant taxa showed that some species were more positively correlated with the oil recovery process, while others acted as competitors for the carbon source. The study also found that higher biomass favored the plugging of high permeability zones in the reservoir, facilitating the dislodging of crude oil in new channels. In conclusion, this study suggests that microbial populations significantly shift upon polymer treatment and contribute synergistically to the oil recovery process depending on the characteristics of the polymers injected.
Key points
• Post-polymer flooded microbial ecology shows unique indigenous microbial consortia.
• Injected polymers are observed to act as enrichment substrates by resident communities.
• The first study to show successive oil recovery stage post-polymer flood without external influence.








Similar content being viewed by others
Data availability
All data generated during this study are available by the corresponding author upon reasonable request.
References
Agrawal A, An D, Cavallaro A, Voordouw G (2014) Souring in low-temperature surface facilities of two high-temperature Argentinian oil fields. Appl Microbiol Biotechnol 98(18):8017–8029. https://doi.org/10.1007/s00253-014-5843-z
Arenskötter M, Bröker D, Steinbüchel A (2004) Biology of the metabolically diverse genus Gordonia. Appl Environ Microbiol 70(6):3195–3204. https://doi.org/10.1128/AEM.70.6.3195-3204.2004
Chen Z-Y, Feng Q-X, Liu R-L (2001) Development and application of thermophilic microorganism species in oil recovery. Acta Petrolei Sinica 22(6):59–62. https://doi.org/10.7623/syxb200106013
Chen M-Y, Wu S-H, Lin G-H, Lu C-P, Lin Y-T, Chang W-C, Tsay S-S (2004) Rubrobacter taiwanensis sp. nov., a novel thermophilic, radiation-resistant species isolated from hot springs. Int J Syst Evol Microbiol 54(5):1849–1855. https://doi.org/10.1099/ijs.0.63109-0
Cheng L, Shi S, Li Q, Chen J, Zhang H, Lu Y (2014) Progressive degradation of crude oil n-alkanes coupled to methane production under mesophilic and thermophilic conditions. PLoS ONE 9(11):e113253. https://doi.org/10.1371/journal.pone.0113253
Cui K, Sun S, Xiao M, Liu T, Xu Q, Dong H, Wang D, Gong Y, Sha T, Hou J (2018) Microbial mineralization of montmorillonite in low-permeability oil reservoirs for microbial enhanced oil recovery. Appl Environ Microbiol 84(14):e00176–18. https://doi.org/10.1128/AEM.00176-18
Dong X, Greening C, Brüls T, Conrad R, Guo K, Blaskowski S, Kaschani F, Kaiser M, Laban NA, Meckenstock RU (2018) Fermentative Spirochaetes mediate necromass recycling in anoxic hydrocarbon-contaminated habitats. ISME J 12(8):2039–2050. https://doi.org/10.1038/s41396-018-0148-3
Gao C (2013) Viscosity of partially hydrolyzed polyacrylamide under shearing and heat. J Pet Explor Prod Technol 3(3):203–206. https://doi.org/10.1007/s13202-013-0051-4
Gao P, Li G, Le J, Liu X, Liu F, Ma T (2018) Succession of microbial communities and changes of incremental oil in a post-polymer flooded reservoir with nutrient stimulation. Appl Microbiol Biotechnol 102(4):2007–2017. https://doi.org/10.1007/s00253-018-8766-2
Gassara F, Suri N, Stanislav P, Voordouw G (2015) Microbially enhanced oil recovery by sequential injection of light hydrocarbon and nitrate in low-and high-pressure bioreactors. Environ Sci Technol 49(20):12594–12601. https://doi.org/10.1021/acs.est.5b03879
Gassara F, Suri N, Voordouw G (2017) Nitrate-mediated microbially enhanced oil recovery (N-MEOR) from model upflow bioreactors. J Hazard Mater 324:94–99. https://doi.org/10.1016/j.jhazmat.2015.12.039
Geetha S, Banat IM, Joshi SJ (2018) Biosurfactants: production and potential applications in microbial enhanced oil recovery (MEOR). Biocatal Agric Biotechnol 14:23–32. https://doi.org/10.1016/j.bcab.2018.01.010
Gittel A, Sørensen KB, Skovhus TL, Ingvorsen K, Schramm A (2009) Prokaryotic community structure and sulfate reducer activity in water from high-temperature oil reservoirs with and without nitrate treatment. Appl Environ Microbiol 75(22):7086–7096. https://doi.org/10.1128/AEM.01123-09
Goodyear S, Mead B, Woods C (1995) A novel recovery mechanism for polymer flooding in gravity-dominated viscous oil reservoirs. SPE Reservoir Eng 10(04):259–265. https://doi.org/10.2118/27772-PA
Hania WB, Bouanane-Darenfed A, Cayol J-L, Ollivier B, Fardeau M-L (2016) Reclassification of Anaerobaculum mobile, Anaerobaculum thermoterrenum, Anaerobaculum hydrogeniformans as Acetomicrobium mobile comb. nov., Acetomicrobium thermoterrenum comb. nov. and Acetomicrobium hydrogeniformans comb. nov., respectively, and emendation of the genus Acetomicrobium. Int J Syst Evol Microbiol 66(3):1506–1509. https://doi.org/10.1099/ijsem.0.000910
Ke C-Y, Sun W-J, Li Y-B, Hui J-F, Lu G-M, Zheng X-Y, Zhang Q-Z, Zhang X-L (2018) Polymer-assisted microbial-enhanced oil recovery. Energy Fuels 32(5):5885–5892. https://doi.org/10.1021/acs.energyfuels.8b00812
Ke C-Y, Lu G-M, Wei Y-L, Sun W-J, Hui J-F, Zheng X-Y, Zhang Q-Z, Zhang X-L (2019) Biodegradation of crude oil by Chelatococcus daeguensis HB-4 and its potential for microbial enhanced oil recovery (MEOR) in heavy oil reservoirs. Bioresour Technol 287:121442. https://doi.org/10.1016/j.biortech.2019.121442
Kouřilová X, Schwarzerová J, Pernicová I, Sedlář K, Mrázová K, Krzyžánek V, Nebesářová J, Obruča S (2021) The first insight into polyhydroxyalkanoates accumulation in multi-extremophilic Rubrobacter xylanophilus and Rubrobacter spartanus. Microorganisms 9(5):909. https://doi.org/10.3390/microorganisms9050909
Le J, Liu F, Zhang J, Bai L, Wang R, Liu X, Hou Z, Lin W (2014) A field test of activation indigenous microorganism for microbial enhanced oil recovery in reservoir after polymer flooding. Acta Petrolei Sinica 35(1):99–106. https://doi.org/10.7623/syxb201401011
Li G, Gao P, Wu Y, Tian H, Dai X, Wang Y, Cui Q, Zhang H, Pan X, Dong H (2014) Microbial abundance and community composition influence production performance in a low-temperature petroleum reservoir. Environ Sci Technol 48(9):5336–5344. https://doi.org/10.1021/es500239w
Li X-X, Liu J-F, Zhou L, Mbadinga SM, Yang S-Z, Gu J-D, Mu B-Z (2017) Diversity and composition of sulfate-reducing microbial communities based on genomic DNA and RNA transcription in production water of high temperature and corrosive oil reservoir. Front Microbiol 8:1011. https://doi.org/10.3389/fmicb.2017.01011
Nakano M, Kihara M, Iehata S, Tanaka R, Maeda H, Yoshikawa T (2011) Wax ester-like compounds as biosurfactants produced by Dietzia maris from n-alkane as a sole carbon source. J Basic Microbiol 51(5):490–498. https://doi.org/10.1002/jobm.201000420
Nasr-El-Din H, Hawkins B, Green K (1991) Viscosity behavior of alkaline, surfactant, polyacrylamide solutions used for enhanced oil recovery. In: SPE International Symposium on Oilfield Chemistry, Society of Petroleum Engineers, SPE-21028. https://doi.org/10.2118/21028-MS
Nazina TN, Shestakova NM, Semenova EM, Korshunova AV, Kostrukova NK, Tourova TP, Min L, Feng Q, Poltaraus AB (2017) Diversity of metabolically active bacteria in water-flooded high-temperature heavy oil reservoir. Front Microbiol 8:707. https://doi.org/10.3389/fmicb.2017.00707
Nazina TN, Bidzhieva SK, Grouzdev DS, Sokolova DS, Tourova TP, Parshina SN, Avtukh AN, Poltaraus AB, Talybly AK (2020) Soehngenia longivitae sp. nov., a fermenting bacterium isolated from a petroleum reservoir in Azerbaijan, and emended description of the genus soehngenia. Microorganisms. 8(12):1967. https://doi.org/10.3390/microorganisms8121967
Peihui H, Fengrong S, Mei S (2001) Microbial for studies on the microorganisms using petroleum hydrocarbons as sole carbon source. In: SPE Asia Pacific Improved Oil Recovery Conference, Kuala Lumpur, Malaysia, SPE-72128-MS. https://doi.org/10.2118/72128-MS
Quero GM, Cassin D, Botter M, Perini L, Luna GM (2015) Patterns of benthic bacterial diversity in coastal areas contaminated by heavy metals, polycyclic aromatic hydrocarbons (PAHs) and polychlorinated biphenyls (PCBs). Front Microbiol 6:1053. https://doi.org/10.3389/fmicb.2015.01053
Reis R, Pacheco G, Pereira A, Freire D (2013) Biosurfactants: production and applications. In: Rolando C, Francisca R (eds) Biodegradation - Life of Science, InTech, London UK, pp 31–61. https://doi.org/10.5772/56144
Rellegadla S, Jain S, Agrawal A (2021) A holistic approach to determine the enhanced oil recovery potential of hydroxyethylcellulose, tragacanth gum and carboxymethylcellulose. J Mol Liq 341:117334. https://doi.org/10.1016/j.molliq.2021.117334
Ren G, Wang J, Qu L, Li W, Hu M, Bian L, Zhang Y, Le J, Dou X, Chen X (2020) Compositions and co-occurrence patterns of bacterial communities associated with polymer-and ASP-flooded petroleum reservoir blocks. Front Microbiol 11:3011. https://doi.org/10.3389/fmicb.2020.580363
Rosenberg E, Barkay T, Navon-Venezia S, Ron E (1999) Role of Acinetobacter bioemulsans in petroleum degradation. In: Fass R, Flashner Y, Reuveny S (eds) Novel Approaches for Bioremediation of Organic Pollution. Springer, Boston, MA, pp 171–180. https://doi.org/10.1007/978-1-4615-4749-5_17
Safdel M, Anbaz MA, Daryasafar A, Jamialahmadi M (2017) Microbial enhanced oil recovery, a critical review on worldwide implemented field trials in different countries. Renew Sustain Energy Rev 74:159–172. https://doi.org/10.1016/j.rser.2017.02.045
Schippers A, Schumann P, Spröer C (2005) Nocardioides oleivorans sp. nov., a novel crude-oil-degrading bacterium. Int J Syst Evol Microbiol 55(4):1501–1504. https://doi.org/10.1099/ijs.0.63500-0
Sharma PK, Munir RI, Blunt W, Dartiailh C, Cheng J, Charles TC, Levin DB (2017) Synthesis and physical properties of polyhydroxyalkanoate polymers with different monomer compositions by recombinant Pseudomonas putida LS46 expressing a novel PHA synthase (PhaC116) enzyme. Appl Sci 7(3):242. https://doi.org/10.3390/app7030242
She Y-H, Zhang F, Xia J-J, Kong S-Q, Wang Z-L, Shu F-C, Hu J-M (2011) Investigation of biosurfactant-producing indigenous microorganisms that enhance residue oil recovery in an oil reservoir after polymer flooding. Appl Biochem Biotechnol 163(2):223–234. https://doi.org/10.1007/s12010-010-9032-y
She H, Kong D, Li Y, Hu Z, Guo H (2019) Recent advance of microbial enhanced oil recovery (MEOR) in China. Geofluids 2019(1):1871392. https://doi.org/10.1155/2019/1871392
Sokolova DS, Semenova EM, Grouzdev DS, Bidzhieva SK, Babich TL, Loiko NG, Ershov AP, Kadnikov VV, Beletsky AV, Mardanov AV (2021) Sulfidogenic microbial communities of the uzen high-temperature oil field in Kazakhstan. Microorganisms 9(9):1818. https://doi.org/10.3390/microorganisms9091818
Souza EC, Vessoni-Penna TC, de Souza Oliveira RP (2014) Biosurfactant-enhanced hydrocarbon bioremediation: an overview. Int Biodeterior Biodegrad 89:88–94. https://doi.org/10.1016/j.ibiod.2014.01.007
Sun W, Dong Y, Gao P, Fu M, Ta K, Li J (2015) Microbial communities inhabiting oil-contaminated soils from two major oilfields in Northern China: implications for active petroleum-degrading capacity. J Microbiol 53(6):371–378. https://doi.org/10.1007/s12275-015-5023-6
Sun J-Q, Xu L, Liu X-Y, Zhao G-F, Cai H, Nie Y, Wu X-L (2018) Functional genetic diversity and culturability of petroleum-degrading bacteria isolated from oil-contaminated soils. Front Microbiol 9:1332. https://doi.org/10.3389/fmicb.2018.01332
Teramoto M, Suzuki M, Hatmanti A, Harayama S (2010) The potential of Cycloclasticus and Altererythrobacter strains for use in bioremediation of petroleum-aromatic-contaminated tropical marine environments. J Biosci Bioeng 110(1):48–52. https://doi.org/10.1016/j.jbiosc.2009.12.008
Vasileva-Tonkova E, Gesheva V (2005) Glycolipids produced by Antarctic Nocardioides sp. during growth on n-paraffin. Process Biochem. 40(7):2387–2391. https://doi.org/10.1016/j.procbio.2004.09.018
Vila J, Nieto JM, Mertens J, Springael D, Grifoll M (2010) Microbial community structure of a heavy fuel oil-degrading marine consortium: linking microbial dynamics with polycyclic aromatic hydrocarbon utilization. FEMS Microbiol Ecol 73(2):349–362. https://doi.org/10.1111/j.1574-6941.2010.00902.x
Wang X-B, Chi C-Q, Nie Y, Tang Y-Q, Tan Y, Wu G, Wu X-L (2011) Degradation of petroleum hydrocarbons (C6–C40) and crude oil by a novel Dietzia strain. Bioresour Technol 102(17):7755–7761. https://doi.org/10.1016/j.biortech.2011.06.009
Wang X-B, Nie Y, Tang Y-Q, Wu G, Wu X-L (2013) n-Alkane chain length alters Dietzia sp. strain DQ12-45-1b biosurfactant production and cell surface activity. Appl Environ Microbiol 79(1):400–402. https://doi.org/10.1128/AEM.02497-12
Wang W, Cai B, Shao Z (2014) Oil degradation and biosurfactant production by the deep sea bacterium Dietzia maris As-13-3. Front Microbiol 5:711. https://doi.org/10.3389/fmicb.2014.00711
Wang H-Z, Lv X-M, Yi Y, Zheng D, Gou M, Nie Y, Hu B, Nobu MK, Narihiro T, Tang Y-Q (2019) Using DNA-based stable isotope probing to reveal novel propionate-and acetate-oxidizing bacteria in propionate-fed mesophilic anaerobic chemostats. Sci Rep 9(1):1–12. https://doi.org/10.1038/s41598-019-53849-0
Yamada T, Imachi H, Ohashi A, Harada H, Hanada S, Kamagata Y, Sekiguchi Y (2007) Bellilinea caldifistulae gen. nov., sp. nov. and Longilinea arvoryzae gen. nov., sp. nov., strictly anaerobic, filamentous bacteria of the phylum Chloroflexi isolated from methanogenic propionate-degrading consortia. Int J Syst Evol Microbiol 57(10):2299–2306. https://doi.org/10.1099/ijs.0.65098-0
Zhang F, She Y-H, Chai L-J, Banat IM, Zhang X-T, Shu F-C, Wang Z-L, Yu L-J, Hou D-J (2012) Microbial diversity in long-term water-flooded oil reservoirs with different in situ temperatures in China. Sci Rep 2(1):1–10. https://doi.org/10.1038/srep00760
Zhu H, Carlson HK, Coates JD (2013) Applicability of anaerobic nitrate-dependent Fe (II) oxidation to microbial enhanced oil recovery (MEOR). Environ Sci Technol 47(15):8970–8977. https://doi.org/10.1021/es401838b
Funding
This work was supported by the IMPRINT2 SERB Department of Science and Technology project (Award no: IMP/2018/000589), the Department of Biotechnology project (Award no: BT/PR25132/NER/95/1034/2017), and ONGC. SR acknowledges the support of a Senior Research Fellowship from CSIR India (File no. 09/1131(0028)/2019-EMR-I). This work was also supported by departmental facilities that were generated by the DST FIST grant (SR/FST/LSI-676/2016) to the Department of Microbiology, Central University of Rajasthan.
Author information
Authors and Affiliations
Contributions
SR and AA conceived and designed the study. SR performed research and wrote the original draft. SR, AA, and GP analyzed the data, and GP and SJ performed writing – review and editing.
Corresponding author
Ethics declarations
Ethical approval
This work does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
253_2023_12673_MOESM1_ESM.pdf
Supplementary file 1 The raw reads of microbial communities analysis were deposited in the National Center for Biotechnology Information (BioProject ID: PRJNA796728, http://www.ncbi.nlm.nih.gov/bioproject/796728). All data generated or analyzed during this study are included in this published article and its supplementary information files. The experimental procedures, polymer flooding ORSBs recovery data, sequencing data (rarefaction curves, diversity indices, relative abundances), and procedures including crude oil standard preparation are in the Supplementary Information. (PDF 2781 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Rellegadla, S., Prajapat, G., Jain, S. et al. Microbial communities succession post to polymer flood demonstrate a role in enhanced oil recovery. Appl Microbiol Biotechnol 107, 5531–5544 (2023). https://doi.org/10.1007/s00253-023-12673-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-023-12673-3