Abstract
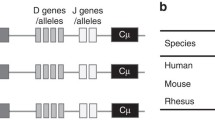
Protection and neutralization of a vast array of pathogens is accomplished by the tremendous diversity of the B cell receptor (BCR) repertoire. For jawed vertebrates, this diversity is initiated via the somatic recombination of immunoglobulin (Ig) germline elements. While it is clear that the number of these germline segments differs from species to species, the extent of cross-species sequence diversity remains largely uncharacterized. Here we use extensive computational and statistical methods to investigate the sequence diversity and evolutionary relationship between Ig variable (V), diversity (D), and joining (J) germline segments across nine commonly studied species ranging from zebrafish to human. Metrics such as guanine-cytosine (GC) content showed low redundancy across Ig germline genes within a given species. Other comparisons, including amino acid motifs, evolutionary selection, and sequence diversity, revealed species-specific properties. Additionally, we showed that the germline-encoded diversity differs across antibody (recombined V-D-J) repertoires of various B cell subsets. To facilitate future comparative immunogenomics analysis, we created VDJgermlines, an R package that contains the germline sequences from multiple species. Our study informs strategies for the humanization and engineering of therapeutic antibodies.






Similar content being viewed by others
References
(1980) Thermostability and aliphatic index of globular proteins. J Biochem. https://doi.org/10.1093/oxfordjournals.jbchem.a133168
Abbott RK, Lee JH, Menis S et al (2018) Precursor frequency and affinity determine B cell competitive fitness in germinal centers, tested with germline-targeting HIV vaccine immunogens. Immunity 48:133–146.e6. https://doi.org/10.1016/j.immuni.2017.11.023
Avnir Y, Watson CT, Glanville J, Peterson EC, Tallarico AS, Bennett AS, Qin K, Fu Y, Huang CY, Beigel JH, Breden F, Zhu Q, Marasco WA (2016) IGHV1-69 polymorphism modulates anti-influenza antibody repertoires, correlates with IGHV utilization shifts and varies by ethnicity. Sci Rep 6:20842. https://doi.org/10.1038/srep20842
Bolotin DA, Poslavsky S, Mitrophanov I, Shugay M, Mamedov IZ, Putintseva EV, Chudakov DM (2015) MiXCR: software for comprehensive adaptive immunity profiling. Nat Methods 12:380–381. https://doi.org/10.1038/nmeth.3364
Bouckaert R, Heled J, Kühnert D, Vaughan T, Wu CH, Xie D, Suchard MA, Rambaut A, Drummond AJ (2014) BEAST 2: a software platform for Bayesian evolutionary analysis. PLoS Comput Biol 10:e1003537. https://doi.org/10.1371/journal.pcbi.1003537
Boyd SD, Gaëta BA, Jackson KJ, Fire AZ, Marshall EL, Merker JD, Maniar JM, Zhang LN, Sahaf B, Jones CD, Simen BB, Hanczaruk B, Nguyen KD, Nadeau KC, Egholm M, Miklos DB, Zehnder JL, Collins AM (2010) Individual variation in the germline Ig gene repertoire inferred from variable region gene rearrangements. JI 184:6986–6992. https://doi.org/10.4049/jimmunol.1000445
Breden F, Luning Prak ET, Peters B, Rubelt F, Schramm CA, Busse CE, Vander Heiden JA, Christley S, Bukhari SAC, Thorogood A, Matsen IV FA, Wine Y, Laserson U, Klatzmann D, Douek DC, Lefranc MP, Collins AM, Bubela T, Kleinstein SH, Watson CT, Cowell LG, Scott JK, Kepler TB (2017) Reproducibility and reuse of adaptive immune receptor repertoire data. Front Immunol 8:1418. https://doi.org/10.3389/fimmu.2017.01418
Brown AJ, Snapkov I, Akbar R, Pavlović M, Miho E, Sandve GK, Greiff V (2019) Augmenting adaptive immunity: progress and challenges in the quantitative engineering and analysis of adaptive immune receptor repertoires. Mol Syst Des Eng 4:701–736. https://doi.org/10.1039/C9ME00071B
Chimge N-O, Pramanik S, Hu G, Lin Y, Gao R, Shen L, Li H (2005) Determination of gene organization in the human IGHV region on single chromosomes. Genes Immun 6:186–193. https://doi.org/10.1038/sj.gene.6364176
Colijn C, Plazzotta G (2017) A metric on phylogenetic tree shapes. Syst Biol 67:113–126. https://doi.org/10.1093/sysbio/syx046
Colless DH (1982) Phylogenetics: the theory and practice of phylogenetic systematics. Syst Zool 31:100–104. https://doi.org/10.2307/2413420
Collins AM, Wang Y, Roskin KM, Marquis CP, Jackson KJL (2015) The mouse antibody heavy chain repertoire is germline-focused and highly variable between inbred strains. Philos Trans R Soc B 370:20140236. https://doi.org/10.1098/rstb.2014.0236
Corcoran MM, Phad GE, Bernat NV, Stahl-Hennig C, Sumida N, Persson MAA, Martin M, Hedestam GBK (2016) Production of individualized V gene databases reveals high levels of immunoglobulin genetic diversity. Nat Commun 7:13642. https://doi.org/10.1038/ncomms13642
Drummond AJ, Rambaut A (2007) BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol Biol 7:214. https://doi.org/10.1186/1471-2148-7-214
Elhanati Y, Murugan A, Callan CG et al (2014) Quantifying selection in immune receptor repertoires. PNAS 111:9875–9880
Elhanati Y, Sethna Z, Marcou Q, Callan CG Jr, Mora T, Walczak AM (2015) Inferring processes underlying B-cell repertoire diversity. Philos Trans R Soc, B Biol Sci 370:20140243. https://doi.org/10.1098/rstb.2014.0243
Friedensohn S, Lindner JM, Cornacchione V, Iazeolla M, Miho E, Zingg A, Meng S, Traggiai E, Reddy ST (2018) Synthetic standards combined with error and bias correction improve the accuracy and quantitative resolution of antibody repertoire sequencing in human naïve and memory B cells. Front Immunol 9. https://doi.org/10.3389/fimmu.2018.01401
Gadala-Maria D, Yaari G, Uduman M, Kleinstein SH (2015) Automated analysis of high-throughput B-cell sequencing data reveals a high frequency of novel immunoglobulin V gene segment alleles. PNAS 112:E862–E870. https://doi.org/10.1073/pnas.1417683112
Georgiou G, Ippolito GC, Beausang J, Busse CE, Wardemann H, Quake SR (2014) The promise and challenge of high-throughput sequencing of the antibody repertoire. Nat Biotechnol 32:158–168. https://doi.org/10.1038/nbt.2782
Gidoni M, Snir O, Peres A, Polak P, Lindeman I, Mikocziova I, Sarna VK, Lundin KEA, Clouser C, Vigneault F, Collins AM, Sollid LM, Yaari G (2019) Mosaic deletion patterns of the human antibody heavy chain gene locus shown by Bayesian haplotyping. Nat Commun 10:628. https://doi.org/10.1038/s41467-019-08489-3
Glanville J, Zhai W, Berka J, Telman D, Huerta G, Mehta GR, Ni I, Mei L, Sundar PD, Day GMR, Cox D, Rajpal A, Pons J (2009) Precise determination of the diversity of a combinatorial antibody library gives insight into the human immunoglobulin repertoire. Proc Natl Acad Sci 106:20216–20221. https://doi.org/10.1073/pnas.0909775106
Greiff V, Miho E, Menzel U, Reddy ST (2015a) Bioinformatic and statistical analysis of adaptive immune repertoires. Trends Immunol 36:738–749. https://doi.org/10.1016/j.it.2015.09.006
Greiff V, Bhat P, Cook SC et al (2015b) A bioinformatic framework for immune repertoire diversity profiling enables detection of immunological status. Genome Medicine 7:49.
Greiff V, Menzel U, Miho E, Weber C, Riedel R, Cook S, Valai A, Lopes T, Radbruch A, Winkler TH, Reddy ST (2017a) Systems analysis reveals high genetic and antigen-driven predetermination of antibody repertoires throughout B Cell Development. Cell Reports 19(7):1467–1478
Greiff V, Weber CR, Palme J, Bodenhofer U, Miho E, Menzel U, Reddy ST (2017b) Learning the high-dimensional immunogenomic features that predict public and private antibody repertoires. J Immunol 199:2985–2997. https://doi.org/10.4049/jimmunol.1700594
Grenfell BT, Pybus OG, Gog JR et al (2004) Unifying the epidemiological and evolutionary dynamics of pathogens. Science 303:327–332. https://doi.org/10.1126/science.1090727
Guruprasad K, Reddy BVB, Pandit MW (1990) Correlation between stability of a protein and its dipeptide composition: a novel approach for predicting in vivo stability of a protein from its primary sequence. Protein Eng Des Sel 4:155–161. https://doi.org/10.1093/protein/4.2.155
Howe K, Clark MD, Torroja CF, Torrance J, Berthelot C, Muffato M, Collins JE, Humphray S, McLaren K, Matthews L, McLaren S, Sealy I, Caccamo M, Churcher C, Scott C, Barrett JC, Koch R, Rauch GJ, White S, Chow W, Kilian B, Quintais LT, Guerra-Assunção JA, Zhou Y, Gu Y, Yen J, Vogel JH, Eyre T, Redmond S, Banerjee R, Chi J, Fu B, Langley E, Maguire SF, Laird GK, Lloyd D, Kenyon E, Donaldson S, Sehra H, Almeida-King J, Loveland J, Trevanion S, Jones M, Quail M, Willey D, Hunt A, Burton J, Sims S, McLay K, Plumb B, Davis J, Clee C, Oliver K, Clark R, Riddle C, Elliott D, Threadgold G, Harden G, Ware D, Begum S, Mortimore B, Kerry G, Heath P, Phillimore B, Tracey A, Corby N, Dunn M, Johnson C, Wood J, Clark S, Pelan S, Griffiths G, Smith M, Glithero R, Howden P, Barker N, Lloyd C, Stevens C, Harley J, Holt K, Panagiotidis G, Lovell J, Beasley H, Henderson C, Gordon D, Auger K, Wright D, Collins J, Raisen C, Dyer L, Leung K, Robertson L, Ambridge K, Leongamornlert D, McGuire S, Gilderthorp R, Griffiths C, Manthravadi D, Nichol S, Barker G, Whitehead S, Kay M, Brown J, Murnane C, Gray E, Humphries M, Sycamore N, Barker D, Saunders D, Wallis J, Babbage A, Hammond S, Mashreghi-Mohammadi M, Barr L, Martin S, Wray P, Ellington A, Matthews N, Ellwood M, Woodmansey R, Clark G, Cooper JD, Tromans A, Grafham D, Skuce C, Pandian R, Andrews R, Harrison E, Kimberley A, Garnett J, Fosker N, Hall R, Garner P, Kelly D, Bird C, Palmer S, Gehring I, Berger A, Dooley CM, Ersan-Ürün Z, Eser C, Geiger H, Geisler M, Karotki L, Kirn A, Konantz J, Konantz M, Oberländer M, Rudolph-Geiger S, Teucke M, Lanz C, Raddatz G, Osoegawa K, Zhu B, Rapp A, Widaa S, Langford C, Yang F, Schuster SC, Carter NP, Harrow J, Ning Z, Herrero J, Searle SMJ, Enright A, Geisler R, Plasterk RHA, Lee C, Westerfield M, de Jong PJ, Zon LI, Postlethwait JH, Nüsslein-Volhard C, Hubbard TJP, Crollius HR, Rogers J, Stemple DL (2013) The zebrafish reference genome sequence and its relationship to the human genome. Nature 496:498–503. https://doi.org/10.1038/nature12111
Janeway CA, Murphy K (2011) Janeway’s immunobiology, 8th Revised edition. Taylor & Francis
Jardine JG, Kulp DW, Havenar-Daughton C, Sarkar A, Briney B, Sok D, Sesterhenn F, Ereno-Orbea J, Kalyuzhniy O, Deresa I, Hu X, Spencer S, Jones M, Georgeson E, Adachi Y, Kubitz M, deCamp AC, Julien JP, Wilson IA, Burton DR, Crotty S, Schief WR (2016) HIV-1 broadly neutralizing antibody precursor B cells revealed by germline-targeting immunogen. Science 351:1458–1463. https://doi.org/10.1126/science.aad9195
Johnston CM, Wood AL, Bolland DJ, Corcoran AE (2006) Complete sequence assembly and characterization of the C57BL/6 mouse Ig heavy chain V region. J Immunol 176:4221–4234. https://doi.org/10.4049/jimmunol.176.7.4221
Kidd MJ, Chen Z, Wang Y, Jackson KJ, Zhang L, Boyd SD, Fire AZ, Tanaka MM, Gaëta BA, Collins AM (2012) The inference of phased haplotypes for the immunoglobulin H chain V region gene loci by analysis of VDJ gene rearrangements. J Immunol 188:1333–1340. https://doi.org/10.4049/jimmunol.1102097
Kirik U, Greiff L, Levander F, Ohlin M (2017) Parallel antibody germline gene and haplotype analyses support the validity of immunoglobulin germline gene inference and discovery. Mol Immunol 87:12–22. https://doi.org/10.1016/j.molimm.2017.03.012
Kudla G, Lipinski L, Caffin F, Helwak A, Zylicz M (2006) High guanine and cytosine content increases mRNA levels in mammalian cells. PLoS Biol 4:e180. https://doi.org/10.1371/journal.pbio.0040180
Laserson U, Vigneault F, Gadala-Maria D, Yaari G, Uduman M, Vander Heiden JA, Kelton W, Taek Jung S, Liu Y, Laserson J, Chari R, Lee JH, Bachelet I, Hickey B, Lieberman-Aiden E, Hanczaruk B, Simen BB, Egholm M, Koller D, Georgiou G, Kleinstein SH, Church GM (2014) High-resolution antibody dynamics of vaccine-induced immune responses. PNAS 111:4928–4933. https://doi.org/10.1073/pnas.1323862111
Lavinder JJ, Hoi KH, Reddy ST, Wine Y, Georgiou G (2014) Systematic characterization and comparative analysis of the rabbit immunoglobulin repertoire. PLoS One 9:e101322. https://doi.org/10.1371/journal.pone.0101322
Lees W, Busse CE, Corcoran M, et al (2019) OGRDB: a reference database of inferred immune receptor genes. Nucleic Acids Res gkz822. https://doi.org/10.1093/nar/gkz822
Lefranc M-P, Giudicelli V, Ginestoux C, Bodmer J, Muller W, Bontrop R, Lemaitre M, Malik A, Barbie V, Chaume D (1999) IMGT, the international ImMunoGeneTics database. Nucleic Acids Res 27:209–212. https://doi.org/10.1093/nar/27.1.209
Letunic I, Bork P (2016) Interactive tree of life (iTOL) v3: an online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res 44:W242–W245. https://doi.org/10.1093/nar/gkw290
Levenshtein IV (1966) Binary codes capable of correcting deletions, insertions, and reversals. Soviet Phys 10:707–710
Lewitus E, Morlon H (2016) Characterizing and comparing phylogenies from their Laplacian spectrum. Syst Biol 65:495–507. https://doi.org/10.1093/sysbio/syv116
Methot SP, Di Noia JM (2017) Molecular mechanisms of somatic hypermutation and class switch recombination. In: Advances in Immunology. Elsevier, pp 37–87
Miho E, Yermanos A, Weber CR, Berger CT, Reddy ST, Greiff V (2018) Computational strategies for dissecting the high-dimensional complexity of adaptive immune repertoires. Front Immunol 9. https://doi.org/10.3389/fimmu.2018.00224
Nadel B, Feeney AJ (1997) Nucleotide deletion and P addition in V (D) J recombination: a determinant role of the coding-end sequence. Mol Cell Biol 17:3768–3778
Ohlin M, Scheepers C, Corcoran M, Lees WD, Busse CE, Bagnara D, Thörnqvist L, Bürckert JP, Jackson KJL, Ralph D, Schramm CA, Marthandan N, Breden F, Scott J, Matsen IV FA, Greiff V, Yaari G, Kleinstein SH, Christley S, Sherkow JS, Kossida S, Lefranc MP, van Zelm MC, Watson CT, Collins AM (2019) Inferred allelic variants of immunoglobulin receptor genes: a system for their evaluation, documentation, and naming. Front Immunol 10:435. https://doi.org/10.3389/fimmu.2019.00435
Olivieri DN, von Haeften B, Sánchez-Espinel C, Faro J, Gambón-Deza F (2014) Genomic V exons from whole genome shotgun data in reptiles. Immunogenetics 66:479–492. https://doi.org/10.1007/s00251-014-0784-3
Omer A, Shemesh O, Peres A, et al (2019) VDJbase: an adaptive immune receptor genotype and haplotype database. Nucleic Acids Res gkz872. https://doi.org/10.1093/nar/gkz872
Osorio D, Rondon-Villarreal P, Torres R (2015) Peptides: a package for data mining of antimicrobial peptides. R J 7:4–14
Palme J, Hochreiter S, Bodenhofer U (2015) KeBABS: an R package for kernel-based analysis of biological sequences. Bioinformatics btv176. https://doi.org/10.1093/bioinformatics/btv176
Pape KA, Maul RW, Dileepan T, Paustian AS, Gearhart PJ, Jenkins MK (2018) Naive B cells with high-avidity germline-encoded antigen receptors produce persistent IgM + and transient IgG + memory B cells. Immunity. 48:1135–1143.e4. https://doi.org/10.1016/j.immuni.2018.04.019
Parola C, Neumeier D, Reddy ST (2018) Integrating high-throughput screening and sequencing for monoclonal antibody discovery and engineering. Immunology 153:31–41. https://doi.org/10.1111/imm.12838
Pieper K, Tan J, Piccoli L, Foglierini M, Barbieri S, Chen Y, Silacci-Fregni C, Wolf T, Jarrossay D, Anderle M, Abdi A, Ndungu FM, Doumbo OK, Traore B, Tran TM, Jongo S, Zenklusen I, Crompton PD, Daubenberger C, Bull PC, Sallusto F, Lanzavecchia A (2017) Public antibodies to malaria antigens generated by two LAIR1 insertion modalities. Nature 548:597–601. https://doi.org/10.1038/nature23670
Ralph DK, Iv FAM (2016) Likelihood-based inference of B cell clonal families. PLoS Comput Biol 12:e1005086. https://doi.org/10.1371/journal.pcbi.1005086
Reddy ST, Ge X, Miklos AE, Hughes RA, Kang SH, Hoi KH, Chrysostomou C, Hunicke-Smith SP, Iverson BL, Tucker PW, Ellington AD, Georgiou G (2010) Monoclonal antibodies isolated without screening by analyzing the variable-gene repertoire of plasma cells. Nat Biotechnol 28:965–969. https://doi.org/10.1038/nbt.1673
Rolink A, Streb M, Melchers F (1991) The λ/λ ratio in surface immunoglobulin molecules on B lymphocytes differentiating from DHJH-rearranged murine pre-B cell clones in vitro. Eur J Immunol 21:2895–2898. https://doi.org/10.1002/eji.1830211137
Romo-González T, Vargas-Madrazo E (2005) Structural analysis of substitution patterns in alleles of human immunoglobulin VH genes. Mol Immunol 42:1085–1097. https://doi.org/10.1016/j.molimm.2004.11.004
Sackin MJ (1972) “Good” and “bad” phenograms. Syst Zool 21:225–226. https://doi.org/10.2307/2412292
Sasso EH, Buckner JH, Suzuki LA (1995) Ethnic differences of polymorphism of an immunoglobulin VH3 gene. J Clin Invest 96:1591–1600
Schiaffella E, Sehgal D, Anderson AO, Mage RG (1999) Gene conversion and hypermutation during diversification of VH sequences in developing splenic germinal centers of immunized rabbits. J Immunol 162:3984
Schwartz GW, Hershberg U (2013) Conserved variation: identifying patterns of stability and variability in BCR and TCR V genes with different diversity and richness metrics. Phys Biol 10:035005. https://doi.org/10.1088/1478-3975/10/3/035005
Sormanni P, Aprile FA, Vendruscolo M (2015) The CamSol method of rational design of protein mutants with enhanced solubility. J Mol Biol 427:478–490. https://doi.org/10.1016/j.jmb.2014.09.026
Sormanni P, Amery L, Ekizoglou S, Vendruscolo M, Popovic B (2017) Rapid and accurate in silico solubility screening of a monoclonal antibody library. Sci Rep 7:8200. https://doi.org/10.1038/s41598-017-07800-w
Stanfield RL, Wilson IA, Smider VV (2016) Conservation and diversity in the ultralong third heavy-chain complementarity-determining region of bovine antibodies. Sci Immunol 1:aaf7962. https://doi.org/10.1126/sciimmunol.aaf7962
Tonegawa S (1983) Somatic generation of antibody diversity. Nature 302:575–581. https://doi.org/10.1038/302575a0
van der Loo MPJ (2014) The stringdist package for approximate string matching. R J 6:111–122
Vollmers C, Sit RV, Weinstein JA, Dekker CL, Quake SR (2013) Genetic measurement of memory B-cell recall using antibody repertoire sequencing. Proc Natl Acad Sci 110:13463–13468. https://doi.org/10.1073/pnas.1312146110
Watson CT, Steinberg KM, Huddleston J, Warren RL, Malig M, Schein J, Willsey AJ, Joy JB, Scott JK, Graves TA, Wilson RK, Holt RA, Eichler EE, Breden F (2013) Complete haplotype sequence of the human immunoglobulin heavy-chain variable, diversity, and joining genes and characterization of allelic and copy-number variation. Am J Hum Genet 92:530–546. https://doi.org/10.1016/j.ajhg.2013.03.004
Watson CT, Steinberg KM, Graves TA, Warren RL, Malig M, Schein J, Wilson RK, Holt RA, Eichler EE, Breden F (2015) Sequencing of the human IG light chain loci from a hydatidiform mole BAC library reveals locus-specific signatures of genetic diversity. Genes Immun 16:24–34. https://doi.org/10.1038/gene.2014.56
Watson CT, Glanville J, Marasco WA (2017) The individual and population genetics of antibody immunity. Trends Immunol 38:459–470. https://doi.org/10.1016/j.it.2017.04.003
Weber J, Peng H, Rader C (2017) From rabbit antibody repertoires to rabbit monoclonal antibodies. Exp Mol Med 49:e305. https://doi.org/10.1038/emm.2017.23
Weinstein JA, Jiang N, White RA, Fisher DS, Quake SR (2009) High-throughput sequencing of the zebrafish antibody repertoire. Science 324:807–810. https://doi.org/10.1126/science.1170020
Wong A, You M (1985) Entropy and distance of random graphs with application to structural pattern recognition. EEE Transactions on Pattern Analysis and Machine Intelligence PAMI-7 599–609
Xu JL, Davis MM (2000) Diversity in the CDR3 region of VH is sufficient for most antibody specificities. Immunity 13:37–45. https://doi.org/10.1016/S1074-7613(00)00006-6
Yermanos A, Greiff V, Krautler NJ, Menzel U, Dounas A, Miho E, Oxenius A, Stadler T, Reddy ST (2017) Comparison of methods for phylogenetic B-cell lineage inference using time-resolved antibody repertoire simulations (AbSim). Bioinformatics 33:3938–3946. https://doi.org/10.1093/bioinformatics/btx533
Zhou JQ, Kleinstein SH (2019) Cutting edge: Ig H chains are sufficient to determine most B cell clonal relationships. JI 203:1687–1692. https://doi.org/10.4049/jimmunol.1900666
Zhou D, Leslie GA, Guo K, Gutman GA (1986) Expression of immunoglobulin lambda chains in the laboratory rat. Eur J Immunogenet 13:299–308. https://doi.org/10.1111/j.1744-313X.1986.tb01114.x
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Figure S1
Number and properties of the germline genes included in this study. (A) The number of IgH D, IgH J, IgK V, and IgL V genes for each species. Allelic germlines were only included for human IgH V genes. (B) Nucleotide sequence length of the IgH V genes. (C) Amino acid sequence length of the variable light chain genes. (D) Distributions of GC content for IgH V genes annotated either as pseudogenes or as functional alleles from human*01. (E) Distributions of protein instability indices for all IgH V genes and (F) IgK V genes. Dashed and solid lines indicate the median and quartiles, respectively. (PDF 665 kb)
Figure S2
Amino acid motifs improve species-separation of variable light chain genes. Mean amino acid frequencies for variable heavy, kappa, and lambda germline segments. The intensity corresponds to the mean occurrence (%) of a given amino acid across all variable gene segments for each species. (B) t-SNE plot projecting the amino acid frequencies for the individual variable heavy segments into two dimensions. Each triangle represents a single variable kappa chain and diamonds represent variable lambda chain. No allelic variants for human_02 were included in the analysis. (C) t-SNE plot projecting the k-mers (k=2,3,4) composition of each variable chain into two dimensions as in B. (PDF 62 kb)
Figure S3
Interspecies comparison between variable kappa and lambda chains. (A-B) The normalized Colless number, normalized Sackin index, normalized ladder size, and maximum distance from root were calculated for each inferred phylogenetic tree composed of all IgK V and IgL V genes per each species, respectively. (PDF 43 kb)
Figure S4
Clustering based on Laplacian spectra across variable heavy, kappa, and lambda chains. Heatmap displaying the pairwise Jensen Shannon index based on the Laplacian spectra from each variable gene phylogenetic tree. (PDF 36 kb)
Figure S5
Cross-species phylogenetic tree of variable heavy genes from human, macaque, platypus, and zebrafish. Sequences from each species are labeled with “H” (blue tip labels), “M” (red tip labels), “P” (black tip labels), and “Z” (green tip labels), respectively. (PDF 41 kb)
Figure S6
Pairwise distance from the variable kappa and lambda chains between species normalized by sequence length. (PDF 554 kb)
Table S1
(PDF 54 kb)
Table S2
(PDF 35 kb)
Rights and permissions
About this article
Cite this article
Yermanos, A., Dounas, A., Greiff, V. et al. Inter- and intraspecies comparison of phylogenetic fingerprints and sequence diversity of immunoglobulin variable genes. Immunogenetics 72, 279–294 (2020). https://doi.org/10.1007/s00251-020-01164-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-020-01164-8