Abstract
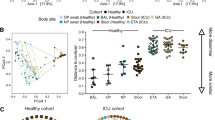
Ventilator-associated pneumonia (VAP) is a nosocomial infection contracted by ventilator patients in which bacteria colonize the upper digestive tract and contaminated secretions are released into the lower airway. This nosocomial infection increases the morbidity and mortality of the patients as well as the cost of treatment. Probiotic formulations have recently been proposed to prevent the colonization of these pathogenic bacteria. In this prospective observational study, we aimed to investigate the effects of probiotics on gut microbiota and their relation to clinical outcomes in mechanically ventilated patients. For this study, 35 patients were recruited (22 probiotic-treated and 13 without probiotic treatment) from a cohort of 169 patients. Patients in the probiotic group were given a dose of 6 capsules of a commercially available probiotic (VSL#3®:112.5 billion CFU/cap) in three divided doses for 10 days. Sampling was carried out after each dose to monitor the temporal change in the gut microbiota composition. To profile the microbiota, we used a 16S rRNA metagenomic approach, and differences among the groups were computed using multivariate statistical analyses. Differences in gut microbial diversity (Bray Curtis and Jaccard distance, p-value > 0.05) between the probiotic-treated group and the control group were not observed. Furthermore, treatment with probiotics resulted in the enrichment of Lactobacillus and Streptococcus in the gut microbiota of the probiotic-treated groups. Our results demonstrated that probiotics might lead to favorable alterations in gut microbiome characteristics. Future studies should focus on the appropriate dosages and frequency of probiotics, which can lead to improved clinical outcomes.











Similar content being viewed by others
Data Availability
Metadata and 16S rRNA gene sequences are submitted to the Sequence read archive (SRA) (NCBI) database https://www.ncbi.nlm.nih.gov/sra/. The submission IDs of the sequences are SUB11977043 (BioProject ID PRJNA874880) and SUB11991923 (PRJNA875248). Sequences will be available to the public immediately after acceptance of the article.
References
Rosenthal VD, Bijie H, Maki DG, Mehta Y, Apisarnthanarak A, Medeiros EA et al (2012) International Nosocomial Infection Control Consortium (INICC) report, data summary of 36 countries, for 2004-2009. Am J Infect Control 40(5):396–407
Ley RE, Peterson DA, Gordon JI (2006) Ecological and evolutionary forces shaping microbial diversity in the human intestine. Cell 124(4):837–848. https://doi.org/10.1016/j.cell.2006.02.017
McNaught CE, Woodcock NP, Anderson ADG, MacFie J (2005) A prospective randomized trial of probiotics in critically ill patients. Clin Nutr 24(2):211–219. https://doi.org/10.1016/j.clnu.2004.08.008
Shimizu K, Yamada T, Ogura H, Mohri T, Kiguchi T, Fujimi S et al (2018) Synbiotics modulate gut microbiota and reduce enteritis and ventilator-associated pneumonia in patients with sepsis: a randomized controlled trial. Crit Care 22(1):239. https://doi.org/10.1186/s13054-018-2167-x
Quigley EMM (2013) Gut bacteria in health and disease. Gastroenterol Hepatol 9(9):560–569
Compare D, Coccoli P, Rocco A, Nardone OM, De Maria S, Cartenì M et al (2012) Gut–liver axis: the impact of gut microbiota on non alcoholic fatty liver disease. Nutr Metab Cardiovasc Dis 22(6):471–476. https://doi.org/10.1016/j.numecd.2012.02.007
Shimizu K, Ogura H, Goto M, Asahara T, Nomoto K, Morotomi M et al (2006) Altered gut flora and environment in patients with severe SIRS. J Trauma 60(1):126–133. https://doi.org/10.1097/01.ta.0000197374.99755.fe
Shimizu K, Ogura H, Hamasaki T, Goto M, Tasaki O, Asahara T, Nomoto K, Morotomi M, Matsushima A, Kuwagata Y, Sugimoto H Altered gut flora are associated with septic complications and death in critically ill patients with systemic inflammatory response syndrome (2021) https://link.springer.com/article/10.1007%2Fs10620-010-1418-8
Clark JA, Coopersmith CM (2007) Intestinal crosstalk: a new paradigm for understanding the gut as the “motor” of critical illness. Shock 28(4):384–393. https://doi.org/10.1097/shk.0b013e31805569df
Zaborin A, Smith D, Garfield K, Quensen J, Shakhsheer B, Kade M et al (2014) Membership and behavior of ultra-low-diversity pathogen communities present in the gut of humans during prolonged critical illness. mBio 5(5):e01361–e01314. https://doi.org/10.1128/mBio.01361-14
Festi D, Schiumerini R, Birtolo C, Marzi L, Montrone L, Scaioli E et al (2011) Gut microbiota and its pathophysiology in disease paradigms. Dig Dis 29(6):518–524. https://doi.org/10.1159/000332975
Munukka E, Rintala A, Toivonen R, Nylund M, Yang B, Takanen A, Hänninen A, Vuopio J, Huovinen P, Jalkanen S, Pekkala S (2021) Faecalibacterium prausnitzii treatment improves hepatic health and reduces adipose tissue inflammation in high-fat fed mice. The ISME Journal. https://www.nature.com/articles/ismej201724 11(7):1667–1679
Machiels K, Joossens M, Sabino J, De Preter V, Arijs I, Eeckhaut V, Ballet V, Claes K, Van Immerseel F, Verbeke K, Ferrante M (2021) A decrease of the butyrate-producing species Roseburia hominis and Faecalibacterium prausnitzii defines dysbiosis in patients with ulcerative colitis. Gut 63(8):1275–1283 https://gut.bmj.com/content/63/8/1275.long
Bag S et al (2016) An improved method for high quality metagenomics DNA extraction from human and environmental samples. SciRep 6:26775. https://doi.org/10.1038/srep2677552
Amplicon PCR, Clean-Up PCR, Index PCR (2013) 16s metagenomic sequencing library preparation. Illumina, San Diego, CA, USA
Bioinformatics B (2011) FastQC: a quality control tool for high throughput sequence data. Babraham Institute, Cambridge, UK
Wick R. Porechop (2018). https://github.com/rrwick/Porechop/.
Wood DE, Lu J, Langmead B (2019) Improved metagenomic analysis with Kraken 2. Genome Biol 20(1):1–13. https://doi.org/10.1186/s13059-019-1891-0
Quast C et al (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:D590–D596. https://doi.org/10.1093/nar/gks1219
Lu J, Breitwieser FP, Thielen P, Salzberg SL (2017) Bracken: estimating species abundance in metagenomics data. PeerJ Computer Science 3:e104
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP (2016) DADA2: High-resolution sample inference from Illumina amplicon data. Nature methods 13(7):581–583. https://doi.org/10.1038/nmeth.3869
Team RC (2013) R: A language and environment for statistical computing
Hauswedell H, Singer J, Reinert K (2014) Lambda: the local aligner for massive biological data. Bioinformatics 30(17):i349–i355. https://doi.org/10.1093/bioinformatics/btu439
McMurdie PJ, Holmes S (2013) phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PloS one 8(4):e61217
Dixon P (2003) VEGAN, a package of R functions for community ecology. J Veg Sci 14(6):927–930
Almeida A, Mitchell AL, Boland M, Forster SC, Gloor GB, Tarkowska A, Lawley TD, Finn RD (2021) A new genomic blueprint of the human gut microbiota. Nature 568(7753):499–504 https://www.nature.com/articles/s41586-019-0965-1
Knight DJW, Gardiner D, Banks A, Snape SE, Weston VC, Bengmark S et al (2008) Effect of synbiotic therapy on the incidence of ventilator associated pneumonia in critically ill patients: a randomised, double-blind, placebo-controlled trial. Intensive Care Med 35(5):854. https://doi.org/10.1007/s00134-008-1368-1
Zeng J, Wang CT, Zhang FS, Qi F, Wang SF, Ma S et al (2016) Effect of probiotics on the incidence of ventilator-associated pneumonia in critically ill patients: a randomized controlled multicenter trial. Intensive Care Med 42(6):1018–1028. https://doi.org/10.1007/s00134-016-4303-x
Angus DC, Linde-Zwirble WT, Lidicker J, Clermont G, Carcillo J, Pinsky MR (2001) Epidemiology of severe sepsis in the United States: analysis of incidence, outcome, and associated costs of care. Crit Care Med 29(7):1303–1310. https://doi.org/10.1097/00003246-200107000-00002
Barraud D, Blard C, Hein F, Marçon O, Cravoisy A, Nace L et al (2010) Probiotics in the critically ill patient: a double blind, randomized, placebo-controlled trial. Intensive Care Med 36(9):1540–7. https://doi.org/10.1007/s00134-010-1927-0
Jain PK, McNaught CE, Anderson ADG, MacFie J, Mitchell CJ (2004) Influence of synbiotic containing Lactobacillus acidophilus La5, Bifidobacterium lactis Bb 12, Streptococcus thermophilus, Lactobacillus bulgaricus and oligofructose on gut barrier function and sepsis in critically ill patients: a randomised controlled trial. Clin Nutr Edinb Scotl 23(4):467–475. https://doi.org/10.1016/j.clnu.2003.12.002
Yamada T, Shimizu K, Ogura H, Asahara T, Nomoto K, Yamakawa K et al (2015) Rapid and sustained long-term decrease of fecal short-chain fatty acids in critically ill patients with systemic inflammatory response syndrome. JPEN J Parenter Enteral Nutr 39(5):569–577. https://doi.org/10.1177/0148607114529596
Bleichner G, Bléhaut H, Mentec H, Moyse D (1977) Saccharomyces boulardii prevents diarrhea in critically ill tube-fed patients. Intensive Care Med 23(5):517–523. https://doi.org/10.1007/s001340050367
Rongrungruang Y, Krajangwittaya D, Pholtawornkulchai K, Tiengrim S, Thamlikitkul V (2015) Randomized controlled study of probiotics containing Lactobacillus casei (Shirota strain) for prevention of ventilator-associated pneumonia. J Med Assoc Thail 98:253–259
Ferrie S, Daley M (2011) Lactobacillus GG as treatment for diarrhea during enteral feeding in critical illness: randomized controlled trial. JPEN J Parenter Enteral Nutr 35:43–49. https://doi.org/10.1177/0148607110370705
Manzanares W, Lemieux M, Langlois PL, Wischmeyer PE (2016) Probiotic and synbiotic therapy in critical illness: a systematic review and meta-analysis. Crit Care 20(1):262. https://doi.org/10.1186/s13054-016-1434-y
Batra P, Soni KD, Mathur P (2020) Efficacy of probiotics in the prevention of VAP in critically ill ICU patients: an updated systematic review and meta-analysis of randomized control trials. J Intensive Care 8(1):81. https://doi.org/10.1186/s40560-020-00487-8
de Toro L, Martín-Consuegra I, Sanchez-Casado M, Pérez-Pedrero Sánchez-Belmonte MJ, López-Reina Torrijos P, Sánchez-Rodriguez P, Raigal-Caño A et al (2014) The influence of symbiotics in multi-organ failure: randomised trial. Med Clin (Barc) 143(4):143–149. https://doi.org/10.1016/j.medcli.2013.09.046
Miquel S, Leclerc M, Martin R, Chain F, Lenoir M, Raguideau S, Hudault S, Bridonneau C, Northen T, Bowen B, Bermúdez-Humarán LG (2021) Identification of metabolic signatures linked to anti-inflammatory effects of Faecalibacterium prausnitzii. mBio 6(2):e00300–e00315 https://journals.asm.org/doi/10.1128/mBio.00300-15
Funding
AR received funding support from the Indian Council of Medical Research (ICMR), Govt. of India (Grant number 5/9/1208/2019-Nut, Dt. 16.09.2019). BD acknowledges Department of Biotechnology, Govt. of India (No. BT/PR30159/MED/15/188/2018) for financial support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Supplementary Information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Saikrishna, K., Talukdar, D., Das, S. et al. Study on Effects of Probiotics on Gut Microbiome and Clinical Course in Patients with Critical Care Illnesses. Microb Ecol 86, 1814–1828 (2023). https://doi.org/10.1007/s00248-023-02224-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-023-02224-8