Abstract
Selecting proper and efficient glycopeptide enrichment approaches are essential for mass spectrometry-based glycoproteomics since glycopeptides are usually with microheterogeneity and low abundance in most biological samples. Herein, we introduced a cotton hydrophilic interaction liquid chromatography (HILIC) approach for large-scale glycopeptide enrichment with 80% acetonitrile/1% trifluoroacetic acid as the optimal sample loading buffer. The comparison of cotton HILIC with Venusil HILIC and mixed anion-exchange (MAX) approaches indicated that cotton HILIC was superior in overall glycopeptide enrichment, whereas Venusil HILIC preferred in complex glycan structures and MAX performed better with high mannose glycans. Exploration of capacity and recovery rate of cotton HILIC illustrated that 5mg cotton packed in a 200μL tip achieved a reasonable glycopeptide enrichment performance (~6% recovery) from ~0.5mg peptides. In conclusion, cotton HILIC can be used as an optional glycopeptide enrichment approach in glycosylation analysis with its specific merit.
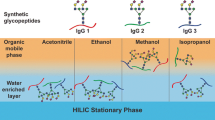
Graphical abstract





Similar content being viewed by others
Data availability
The mass spectrometry data and results have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository [36] with the dataset identifier PXD035078 (username, reviewer_pxd035078@ebi.ac.uk; password, 4gDBebpX).
References
Sprovieri P, Martino G. The role of the carbohydrates in plasmatic membrane. Physiol Res. 2018;67(1):1–11.
Moremen K, Tiemeyer M, Nairn A. Vertebrate protein glycosylation: diversity, synthesis and function. Nat Rev Mol Cell Biol. 2012;13:448–62.
Fournet M, Bonte F, Desmouliere A. Glycation damage: a possible hub for major pathophysiological disorders and aging. Aging Dis. 2018;9(5):880–900.
Yu A, Zhao J, Peng W, Banazadeh A, Williamson SD, Goli M, et al. Advances in mass spectrometry-based glycoproteomics. Electrophoresis. 2018;39(24):3104–22.
Bondt A, Rombouts Y, Selman MH, Hensbergen PJ, Reiding KR, Hazes JM, et al. Immunoglobulin G (IgG) fab glycosylation analysis using a new mass spectrometric high-throughput profiling method reveals pregnancy-associated changes. Mol Cell Proteomics. 2014;13(11):3029–39.
Wieczorek M, Braicu EI, Oliveira-Ferrer L, Sehouli J, Blanchard V. Immunoglobulin G subclass-specific glycosylation changes in primary epithelial ovarian cancer. Front Immunol. 2020;11:654.
Li J, Zhao T, Li J, Shen J, Jia L, Zhu B, et al. Precision N-glycoproteomics reveals elevated LacdiNAc as a novel signature of intrahepatic cholangiocarcinoma. Mol Oncol. 2022;16(11):2135–52.
Kratz EM, Kaluza A, Zimmer M, Ferens-Sieczkowska M. The analysis of sialylation, N-glycan branching, and expression of O-glycans in seminal plasma of infertile men. Dis Markers. 2015;2015:941871.
Cao L, Qu Y, Zhang Z, Wang Z, Prytkova I, Wu S. Intact glycopeptide characterization using mass spectrometry. Expert Rev Proteomics. 2016;13(5):513–22.
Woo CM, Iavarone AT, Spiciarich DR, Palaniappan KK, Bertozzi CR. Isotope-targeted glycoproteomics (IsoTaG): a mass-independent platform for intact N- and O-glycopeptide discovery and analysis. Nat Methods. 2015;12(6):561–7.
Zhu Z, Desaire H. Carbohydrates on proteins: site-specific glycosylation analysis by mass spectrometry. Annu Rev Anal Chem (Palo Alto, Calif). 2015;8:463–83.
Dell A, Morris HR. Glycoprotein structure determination by mass spectrometry. Science (New York, NY). 2001;291(5512):2351–6.
Ongay S, Boichenko A, Govorukhina N, Bischoff R. Glycopeptide enrichment and separation for protein glycosylation analysis. J Sep Sci. 2012;35(18):2341–72.
Liu Y, Fu D, Yu L, Xiao Y, Peng X, Liang X. Oxidized dextran facilitated synthesis of a silica-based concanavalin a material for lectin affinity enrichment of glycoproteins/glycopeptides. J Chromatogr A. 2016;1455:147–55.
Li Y, Shah P, De Marzo AM, Van Eyk JE, Li Q, Chan DW, et al. Identification of glycoproteins containing specific glycans using a lectin-chemical method. Anal Chem. 2015;87(9):4683–7.
Zhang L, Jiang H, Yao J, Wang Y, Fang C, Yang P, et al. Highly specific enrichment of N-linked glycopeptides based on hydrazide functionalized soluble nanopolymers. Chem Commun (Camb). 2014;50(8):1027–9.
Liu Y, Ren L, Liu Z. A unique boronic acid functionalized monolithic capillary for specific capture, separation and immobilization of cis-diol biomolecules. Chem Commun (Camb). 2011;47(17):5067–9.
Wang Y, Liu M, Xie L, Fang C, Xiong H, Lu H. Highly efficient enrichment method for glycopeptide analyses: using specific and nonspecific nanoparticles synergistically. Anal Chem. 2014;86(4):2057–64.
Yang W, Shah P, Hu Y, Toghi Eshghi S, Sun S, Liu Y, et al. Comparison of enrichment methods for intact N- and O-linked glycopeptides using strong anion exchange and hydrophilic interaction liquid chromatography. Anal Chem. 2017;89(21):11193–7.
Shen J, Jia L, Dang L, Su Y, Zhang J, Xu Y, et al. StrucGP: de novo structural sequencing of site-specific N-glycan on glycoproteins using a modularization strategy. Nat Methods. 2021;18(8):921–9.
Dong X, Qin H, Mao J, Yu D, Li X, Shen A, et al. In-depth analysis of glycoprotein sialylation in serum using a dual-functional material with superior hydrophilicity and switchable surface charge. Anal Chem. 2017;89(7):3966–72.
Li J, Liu J, Liu Z, Tan Y, Liu X, Wang F. Detecting proteins glycosylation by a homogeneous reaction system with zwitterionic gold nanoclusters. Anal Chem. 2017;89(8):4339–43.
Pan Y, Ma C, Tong W, Fan C, Zhang Q, Zhang W, et al. Preparation of sequence-controlled triblock copolymer-grafted silica microparticles by sequential-ATRP for highly efficient glycopeptides enrichment. Anal Chem. 2015;87(1):656–62.
Wang Y, Wang J, Gao M, Zhang X. An ultra hydrophilic dendrimer-modified magnetic graphene with a polydopamine coating for the selective enrichment of glycopeptides. J Mater Chem B. 2015;3(44):8711–6.
Taraji M, Haddad PR, Amos RIJ, Talebi M, Szucs R, Dolan JW, et al. Chemometric-assisted method development in hydrophilic interaction liquid chromatography: a review. Anal Chim Acta. 2018;1000:20–40.
Selman MH, Hemayatkar M, Deelder AM, Wuhrer M. Cotton HILIC SPE microtips for microscale purification and enrichment of glycans and glycopeptides. Anal Chem. 2011;83(7):2492–9.
Liu L, Jin S, Mei P, Zhou P. Preparation of cotton wool modified with boric acid functionalized titania for selective enrichment of glycopeptides. Talanta. 2019;203:58–64.
Han J, Chen Q, Jin W, Zou M, Lu Y, Liu Y, et al. Purification of N- and O-glycans and their derivatives from biological samples by the absorbent cotton hydrophilic chromatographic column. J Chromatogr A. 2020;1620:461001.
Xin M, Xu Y, You S, Li C, Zhu B, Shen J, et al. Precision structural interpretation of site-specific N-glycans in seminal plasma. J Proteome Res. 2022;21(7):1664–74.
Xin M, You S, Xu Y, We S, Zhu B, Shen J, Wu J, Li C, Chen Z, Su Y, Shi J, Sun S. Precision mapping of glycosite-specific glycans reveals distinctive N-glycosylation on human spermatozoa. Mol Cell Proteomics. 2021;21(4):100214.
Sun S, Shah P, Eshghi ST, Yang W, Trikannad N, Yang S, et al. Comprehensive analysis of protein glycosylation by solid-phase extraction of N-linked glycans and glycosite-containing peptides. Nat Biotechnol. 2016;34(1):84–8.
Deutsch EW, Mendoza L, Shteynberg D, Slagel J, Sun Z, Moritz RL. Trans-proteomic pipeline, a standardized data processing pipeline for large-scale reproducible proteomics informatics. Proteomics Clin Appl. 2015;9(7-8):745–54.
Bern M, Kil YJ, Becker C. Byonic: advanced peptide and protein identification software. Curr Protoc Bioinformatics. 2012;40(1):13.20.1–13.20.14.
Zhu J, Wang F, Chen R, Cheng K, Xu B, Guo Z, et al. Centrifugation assisted microreactor enables facile integration of trypsin digestion, hydrophilic interaction chromatography enrichment, and on-column deglycosylation for rapid and sensitive N-glycoproteome analysis. Anal Chem. 2012;84(11):5146–53.
Sha Q, Wu Y, Wang C, Sun B, Zhang Z, Zhang L, et al. Cellulose microspheres-filled pipet tips for purification and enrichment of glycans and glycopeptides. J Chromatogr A. 2018;1569:8–16.
Vizcaino JA, Cote RG, Csordas A, Dianes JA, Fabregat A, Foster JM, et al. The PRoteomics IDEntifications (PRIDE) database and associated tools: status in 2013. Nucleic Acids Res. 2013;41(Database issue):D1063–9.
Funding
This work was supported by the National Key R&D Program of China (2019YFA0905200), the Natural Sciences Foundation of Shaanxi Province (Grant No. 2021JQ-447), and the National Natural Science Foundation of China (Grant No. 91853123, 81773180, 81800655, and 21705127).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethics approval
Mice-related experiments were conducted in compliance with ethical regulations and approved by the ethics committee of Northwest University, China. Human seminal plasma-related experiments were approved by the ethics committee of Xi’an Fourth Hospital and Northwest University.
Source of biological materials
Three mice, strain C57BL/6, male, aged 3 months, were obtained from the Central Animal Breeding House of Xi’an Jiaotong University, Xi’an, Shaanxi. Seminal plasma from six healthy men were collected in Xi’an Fourth Hospital. Semen was collected through masturbation in sterile containers after 3–7 days of sexual abstinence according to the criteria of the WHO laboratory manual. Then seminal plasma was obtained by centrifugation of semen.
Statement on animal welfare
Mice were anesthetized with intraperitoneal injection of chloral hydrate, and brain tissue was removed from each mouse and cut up in PBS buffer following internationally recognized guidelines.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xin, M., You, S., Wu, J. et al. Evaluation of absorbent cotton for glycopeptide enrichment. Anal Bioanal Chem 414, 8245–8253 (2022). https://doi.org/10.1007/s00216-022-04353-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-022-04353-4