Abstract
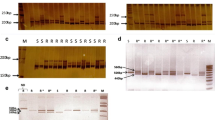
Inheritance and linkage studies were carried out with microsatellite [or simple sequence repeat (SSR)] markers in a F1 progeny including 101 individuals of a cross between Myrobalan plum (Prunus cerasifera Ehrh) clone P.2175 and the almond (Prunus dulcis Mill.)-peach (Prunus persica L. Batsch) hybrid clone GN22 [‘Garfi’ (G) almond × ‘Nemared’ (N) peach]. This three-way interspecific Prunus progeny was produced in order to associate high root-knot nematode (RKN) resistances from Myrobalan and peach with other favorable traits for Prunus rootstocks from plum, peach and almond. The RKN resistance genes, Ma from the Myrobalan plum clone P.2175 and R MiaNem from the ‘N’ peach, are each heterozygous in the parents P.2175 and GN22, respectively. Two hundred and seventy seven Prunus SSRs were tested for their polymorphism. One genetic map was constructed for each parent according to the ‘double pseudo-testcross’ analysis model. The Ma gene and 93 markers [two sequence characterized amplified regions (SCARs), 91 SSRs] were placed on the P.2175 Myrobalan map covering 524.8 cM. The R MiaNem gene, the Gr gene controlling the color of peach leaves, and 166 markers (one SCAR, 165 SSRs) were mapped to seven linkage groups instead of the expected eight in Prunus. Markers belonging to groups 6 and 8 in previous maps formed a single group in the GN22 map. A reciprocal translocation, already reported in a G × N F2, was detected near the Gr gene. By separating markers from linkage groups 6 and 8 from the GN22 map, it was possible to compare the eight homologous linkage groups between the two maps using the 68 SSR markers heterozygous in both parents (anchor loci). All but one of these 68 anchor markers are in the same order in the Myrobalan plum map and in the almond-peach map, as expected from the high level of synteny within Prunus. The Ma and R MiaNem genes confirmed their previous location in the Myrobalan linkage group 7 and in the GN22 linkage group 2, respectively. Using a GN22 F2 progeny of 78 individuals, a microsatellite map of linkage group 2 was also constructed and provided additional evidence for the telomeric position of R MiaNem in group 2 of the Prunus genome.



Similar content being viewed by others
References
Aranzana MJ, Garcia-Mas J, Carbo J, Arús P (2002) Development and variability analysis of microsatellite markers in peach. Plant Breed 121:87–92
Aranzana MJ, Pineda A, Cosson P, Ascasibar J, Dirlewanger E, Ascasibar J, Cipriani G, Ryder CD, Testolin R, Abbott A, King GJ, Iezzoni AF, Arùs P (2003) A set of simple-sequence repeat (SSR) markers covering the Prunus genome. Theor Appl Genet 106:819–825
Bergougnoux V, Claverie M, Bosselut N, Lecouls AC, Salesses G, Dirlewanger E, Esmenjaud D (2002) Marker-assisted selection of the Ma gene from Myrobalan plum for a complete-spectrum root-knot nematode (RKN) resistance in Prunus rootstocks. Acta Hortic 592:223–228
Blake MA (1937) Progress in peach breeding. Proc Am Soc Hortic Sci 35:49–53
Bliss FA, Arulsekar S, Foolad MR, Becerra V, Gillen AM, Warburton ML, Dandekar AM, Kocsisne GM, Mydin KK (2002) An expanded genetic map of Prunus based on an interspecific cross between almond and peach. Genome 45:520–529
Cantini C, Iezzoni AF, Lamboy WF, Bortizki M, Struss D (2001) DNA fingerprinting of tetraploid cherry germplasm using simple sequence repeats. J Am Soc Hortic Sci 126:205–209
Chaparro JX, Werner DJ, O’Malley D, Sederoff RR (1994) Targeted mapping and linkage analysis of morphological, isozyme, and RAPD markers in peach. Theor Appl Genet 87:805–815
Cho YG, Blair MW, Panaud O, McCouch SR (1996) Cloning and mapping of variety-specific rice genomic DNA sequences: amplified fragment length polymorphisms (AFLP) from silver-stained polyacrylamide gels. Genome 39:373–378
Cipriani G, Lot G, Huang WG, Marrazzo MT, Peterlunger E, Testolin R (1999) AC/GT and AG/CT microsatellite repeats in peach [Prunus persica (L.) Batsch]: isolation, characterization and cross-species amplification in Prunus. Theor Appl Genet 99:65–72
Claverie M, Bosselut N, Lecouls AC, Voisin R, Lafargue B, Poizat C, Kleinhentz M, Laigret F, Dirlewanger E, Esmenjaud D (2004) Location of independent root-knot nematode resistance genes in Prunus species. Theor Appl Genet 108:765–773
Dettori MT, Quarta R, Verde I (2001) A peach linkage map integrating RFLPs, SSRs, RAPDs and morphological markers. Genome 44:783–790
Dirlewanger E, Pronier V, Parvery C, Rothan C, Guy A, Monet R (1998) Genetic linkage map of peach (Prunus persica (L.) Batsch) using morphological and molecular markers. Theor Appl Genet 97:888–895
Dirlewanger E, Cosson P, Tavaud M, Aranzana MJ, Poizat C, Zanetto A, Arús P, Laigret F (2002) Development of microsatellite markers in peach [Prunus persica (L.) Batsch] and their use in genetic diversity analysis in peach and sweet cherry (Prunus avium L.). Theor Appl Genet 105:127–138
Downey SL, Iezzoni AF (2000) Polymorphic DNA markers in black cherry (Prunus serotina) are identified using sequences from sweet cherry, peach, and sour cherry. J Am Soc Hortic Sci 125:76–80
Esmenjaud D, Scotto La Massese C, Salesses G, Minot JC, Voisin R (1992) Method and criteria to evaluate resistance to Meloidogyne arenaria in Prunus cerasifera Ehrh. Fundam Appl Nematol 15:385–389
Esmenjaud D, Minot JC, Voisin R, Pinochet J, Salesses G (1994) Inter- and intraspecific resistance variability in Myrobalan plum, peach, and peach–almond rootstocks using 22 root-knot nematode populations. J Am Soc Hortic Sci 119:94–100
Esmenjaud D, Minot JC, Voisin R (1996a) Effect of durable inoculum pressure and high temperature on root-galling, nematode numbers and survival of Myrobalan plum genotypes (Prunus cerasifera) highly resistant to Meloidogyne spp. Fundam Appl Nematol 19:85–90
Esmenjaud D, Minot JC, Voisin R, Bonnet A, Salesses G (1996b) Inheritance of resistance to the root-knot nematode Meloidogyne arenaria in Myrobalan plum. Theor Appl Genet 92:873–879
Esmenjaud D, Minot JC, Voisin R, Pinochet J, Simard MH, Salesses G (1997) Differential response to root-knot nematodes in Prunus species and correlative genetic implications. J Nematol 29:370–380
Fargette M, Phillips MS, Block VC, Waugh R, Trudgill DL (1996) An RFLP study of relationships between species, populations, and resistance breaking lines of tropical Meloidogyne. Fundam Appl Nematol 19:193–200
Fernandez C, Pinochet J, Esmenjaud D, Salesses G, Felipe A (1994) Resistance among new Prunus rootstocks and selections to the root-knot nematodes in Spain and France. HortScience 29:1064–1067
Foolad MR, Arulsekar S, Becerra V, Bliss FA (1995) A genetic map of Prunus based on an interspecific cross between peach and almond. Theor Appl Genet 91:262–269
Garber ED (1972) Introduccion a la citogentica. Compaa Editorial Comercial, Mexico
Georgi L, Wang Y, Yvergniaux D, Ormsbee T, Iñimnigo M, Reighard G, Abbott A (2002) Construction of a BAC library and its application to the identification of simple sequence repeats in peach [Prunus persica (L.) Batsch]. Theor Appl Genet 105:1151–1158
Gómez Aparisi J, Carrera M, Felipe AJ, Socias i Company R (2001) ‘Garnem’, ‘Monegro’ y ‘Felinem’. Nuevos patrones híbridos almendro × melocotonero resistentes a nemátodos y de hoja roja para frutales de hueso. ITEA 97:282–287
Grattapaglia D, Sederoff R (1994) Genetic linkage maps of Eucalyptus grandis and E.urophylla using a pseudo-testcross mapping strategy and RAPD markers. Genetics 137:1121–1137
Graziano E, Arús P (2002) FITMAPS and SHOWMAP: two programs for graphical comparison and plotting of genetic maps. J Hered 93:225–227
Guo M, Lightfoot DA, Mok MC, Mok DWS (1991) Analyses of Phaseolus vulgaris L. and P. coccineus Lam. Hybrids by RFLP: preferential transmission of P. vulgaris alleles. Theor Appl Genet 81:703–709
Hurtado MA, Romero C, Vilanova S, Abbott AG, Llácer G, Badenes ML (2002) Genetic linkage maps of two apricot cultivars (Prunus armaniaca L.), and mapping of PPV (sharka) resistance. Theor Appl Genet 105:182–191
Janati A, Bergé JB, Triantaphyllou AC, Dalmasso A (1982) Nouvelles données sur l’utilisation des isoestérases pour l’identification des Meloidogyne. Rev Nématol 5:147–154
Jauregui B (1998) Localizacion de marcadores moleculares ligados a caracteres agronomicos en un cruzamiento interespecifico almendro × melocotonero. Dissertation, University of Barcelona
Jáuregui B, de Vicente MC, Messeguer R, Felipe A, Bonnet A, Salesses G, Arús P (2001) A reciprocal translocation between ‘Garfi’ almond and ‘Nemared’ peach. Theor Appl Genet 102:1169–1176
Joobeur T, Viruel MA, de Vicente MC, Jáuregui B, Ballester J, Dettori MT, Verde I, Truco MJ, Messeguer R, Batlle I, Quarta R, Dirlewanger E, Arús P (1998) Construction of a saturated linkage map for Prunus using an almond × peach F2 progeny. Theor Appl Genet 97:1034–1041
Joobeur T, Periam N, de Vicente MC, King GJ, Arús P (2000) Development of a second generation linkage map for almond using RAPD and SSR markers. Genome 43:649–655
Kester ED, Grassely C (1987) Almond rootstocks. In: Rom RC, Carlson RF (eds) Rootstocks for fruit crops. Wiley, New York, pp 265–293
Kianian SF, Quiros CF (1992) Generation of a Brassica oleracea composite RFLP map: linkage arrangements among various populations and evolutionary implications. Theor Appl Genet 84:544–554
Kochba J, Spiegel-Roy P (1975) Inheritance to the root-knot nematode (Meloidogyne javanica Chitwood) in bitter almond progenies. Euphytica 24:453–457
Lambert P, Hagen LS, Arús P, Audergon JM (2004) Genetic linkage maps of two apricot cultivars (Prunus armeniaca L.) compared with the almond ‘Texas’ × peach ‘Earlygold’ reference map for Prunus. Theor Appl Genet 108:1120–1130
Lander E, Green P, Abrahamson J, Barlow A, Daley M, Lincoln S, Newburg L (1987) MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181
Layne REC (1987) Peach rootstocks. In: Rom RC and Carlson RF (eds) Rootstocks for fruit crops. Willey, New York, pp 185–216
Lecouls AC, Salesses G, Minot JC, Voisin R, Bonnet A, Esmenjaud D (1997) Spectrum of the Ma genes for resistance to Meloidogyne spp. in Myrobalan plum. Theor Appl Genet 85:1325–2334
Lecouls AC, Rubio-Cabetas MJ, Minot JC, Voisin R, Bonnet A, Salesses G,Dirlewanger E, Esmenjaud D (1999) RAPD and SCAR markers linked to the Ma1 root-knot nematode resistance gene in Myrobalan plum (Prunus cerasifera Ehrh.). Theor Appl Genet 99:328–336
Lecouls AC, Bergougnoux V, Rubio-Cabetas MJ, Bosselut N, Voisin R, Poessel JL, Faurobert M, Bonnet A, Salesses G, Dirlewanger E, Esmenjaud D (2004) Marker-assisted selection of Prunus rootstocks for the wide-spectrum root-knot nematode resistance conferred by the Ma gene from Myrobalan plum (Prunus cerasifera). Mol Breed 13:113–124
Lopes MS, Sefc KM, Laimer M, Da Camara Machado A (2002) Identification of microsatellite loci in apricot. Mol Ecol Notes 2:24–26
Lu ZX, Sosinski B, Reighard GL, Baird WV, Abbott AG (1998) Construction of a genetic linkage map and identification of AFLP markers for resistance to root-knot nematodes in peach rootstocks. Genome 41:199–207
Lu ZX, Sossey-Alaoui GL, Reighard GL, Baird WV, Abbott AG (1999) Development and characterization of a codominant marker linked to root-knot nematode resistance, and its application to peach rootstock breeding. Theor Appl Genet 99:115–122
Minz G, Cohn E (1962) Susceptibility of peach rootstocks to root-knot nematodes. Plant Dis Rep 46:531–534
Mnejja M, Garcia-Mas J, Howad W, Badenes ML, Arús P (2004) Simple-sequence repeat (SSR) markers of Japanese plum (Prunus salicina Lindl.) are highly polymorphic and transferable to peach and almond. Mol Ecol Notes 4:163–166
Nyczepir AP (1991) Nematode management strategies in stone fruits in the United States. J Nematol 23:334–341
Rajapakse S, Belthoff LE, He G, Estager AE, Scorza R, Verde I, Ballard RE, Baird WV, Callahan A, Monet R, Abbott AG (1995) Genetic linkage mapping in peach using morphological, RFLP and RAPD markers. Theor Appl Genet 91:964–971
Ramming DW, Tanner O (1983) Nemared peach rootstock. HortScience 18:376
Rehder A (1954) Manual of cultivated trees and shrubs, 2nd edn. Dioscorides Press, Portland
Rubio-Cabetas MJ, Lecouls AC, Salesses G, Bonnet A, Minot JC, Voisin R, Esmenjaud D (1998) Evidence of a new gene for high resistance to Meloidogyne spp. in Myrobalan plum (Prunus cerasifera). Plant Breed 117:567–571
Rubio-Cabetas MJ, Minot JC, Voisin R, Esmenjaud D, Salesses G, Bonnet A (1999) Response of the Ma genes from Myrobalan plum to Meloidogyne hapla and M. mayaguensis. HortScience 34:1266–1268
Salesses G, Grasselly C, Renaud R, Claverie J (1992) Les porte-greffe des espèces fruitières à noyau du genre Prunus. In: Gallais A, Bannerot H (eds) Amélioration des espèces végétales cultivées: objectifs et critères de sélection. INRA Editions, Paris, pp 605–650
Salesses G, Dirlewanger E, Bonnet A, Lecouls AC, Esmenjaud D (1998) Interspecific hybridization and rootstock breeding for peach. Acta Hortic 465:209–217
Scotto La Massese C, Grasselly C, Minot JC, Voisin R (1984) Différence de comportement de 23 clones et hybrides de Prunus à l’égard de quatre espèces de Meloidogyne. Rev Nématol 7:265–270
Sharpe RH, Hesse CO, Lownsbery BF, Perry VG, Hansen CJ (1969) Breeding peaches for root-knot nematode resistance. J Am Soc Hortic Sci 94:209–212
Sherman WB, Lyrene PM, Hansche PE (1981) Breeding peach rootstocks resistant to root-knot nematodes. HortScience 64:523–524
Sosinski B, Gannavarapu M, Hager LD, Beck LE, King GJ, Ryder CD, Rajapakse S, Baird WV, Ballard RE, Abbott AG (2000) Characterization of microsatellite markers in peach [Prunus persica (L.) Batsch]. Theor Appl Genet 97:1034–1041
Testolin R, Marrazzo T, Cipriani G, Quarta R, Verde I, Dettori MT, Pancaldi M, Sansavini S (2000) Microsatellite DNA in peach (Prunus persica L. Batsch) and its use in fingerprinting and testing the genetic origin of cultivars. Genome 43:512–520
Viruel MA, Messeguer R, de Vicente MC, Garcia-Mas J, Puigdomènech P, Vargas F, Arús P (1995) A linkage map with RFLP and isozyme markers for almond. Theor Appl Genet 91:964–971
Wang D, Karle R, Brettin TS, Iezzoni AF (1998) Genetic linkage map in sour cherry using RFLP markers. Theor Appl Genet 97:1217–1224
Yamamoto T, Hayashi T (2002) New root-knot nematode resistance genes and their STS markers in peach. Sci Hortic 96:81–90
Yamamoto T, Shimada T, Imai T, Yaegaki H, Haji T, Matsuta N, Yamagushi M, Hayashi T (2001) Characterization of morphological traits based on a genetic linkage map in peach. Breed Sci 51:271–278
Yamamoto T, Mochida K, Imai T, Shi Z, Ogiwara I, Hayashi T (2002) Microsatellite markers in peach [Prunus persica (L.) Batsch] derived from an enriched genomic and cDNA libraries. Mol Ecol Notes 2:298–301
Acknowledgements
This work was partly funded by the Commission of the European Union via the FAIR Program of Research and Technological Development (Research project No. FAIR6-CT 98-4139; 1999–2003) and by the Conseil Regional d’Aquitaine (2000–2002). The authors thank the technical staff of the INRA ‘Domaine des Jarres’ experimental farm (Unité Expérimentale Arboricole) for cultivating the interspecific Myrobalan plum × almond-peach hybrids. The authors also wish to thank Patrick Lambert (INRA, Avignon, France) for the AMPA SSR primers.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by C. Möllers
Rights and permissions
About this article
Cite this article
Dirlewanger, E., Cosson, P., Howad, W. et al. Microsatellite genetic linkage maps of myrobalan plum and an almond-peach hybrid—location of root-knot nematode resistance genes. Theor Appl Genet 109, 827–838 (2004). https://doi.org/10.1007/s00122-004-1694-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-004-1694-9