Abstract
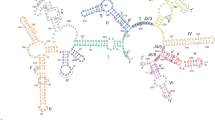
Splicing of group II introns is essential for the metabolism of many organisms (Michel et al. 1989; Michel and Ferat 1995). These ubiquitous introns play a critical role in the processing of mitochondrial genes from plants, fungi, and yeast. Group II introns and RNA molecules resembling them are abundant in euglena and other lower eukaryotes, and they have even been identified in prokaryotes. It has been proposed that, through reactions analogous to the reverse of splicing, excised introns can migrate and introduce themselves into new genomes that may not even ordinarily contain introns (Lambowitz and Belfort 1993; Schmidt et al. 1994). Thus, in addition to their function in RNA splicing, group II introns have the capability for involvement in other biochemical transformations. The apparent complexity of their structure and active-site chemistry has fueled interest in the mechanism of group II intron catalysis. This chapter attempts to describe recent work on group II intron chemistry and its foundation in structural features of the folded RNA.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Preview
Unable to display preview. Download preview PDF.
Similar content being viewed by others
References
Abramovitz D, Pyle AM (1996) Catalytic role of 2′-hydroxyl groups within a group II intron active site. Science 271: 1410–1413
Allain FH-T, Varani G (1995) Divalent metal ion binding to a conserved wobble pair defining the upstream site of cleavage of group I self-splicing introns. Nucl Acids Res 23: 341–350
Altura R, Rymond B, Seraphin B, Rosbash M (1989) Sequence requirements for branch formation in a group II self-splicing intron. Nucl Acids Res 17: 335–354
Augustin S, Müller MW, Schweyen RJ (1990) Reverse self-splicing of group II intron RNAs in vitro. Nature 343: 383–386
Bachl J, Schmelzer C (1990) Effect of deletions at structural domains of group II intron bIl on self-splicing in vitro. J Mol Biol 212: 113–125
Beebe JA, Fierke CA (1994) A kinetic mechanism for cleavage of pre-tRNA Asp catalyzed by the RNA component of Bacillus subtilis ribonuclease P. Biochemistry 33: 10294–10304
Bevilacqua PC, Turner DH (1991) Comparison of binding of mixed ribose-deoxyribose analogues of CUCU to a ribozyme and to GGAGAA by equilibrium dialysis: evidence for ribozyme specific interactions with 2’-OH groups. Biochemistry 30: 10632–10640
Boulanger SC, Belcher SM, Schmidt U, Dib-Hajj SD, Schmidt T, Perlman PS (1995) Studies of point mutants define three essential paired nucleotides in the domain 5 substructure of a group II intron. Mol Cell Biol 15: 4479–4488
Cech TR (1986) The generality of self-splicing RNA: relationship to nuclear mRNA splicing. Cell 44: 207–210
Cech TR (1993) Structure and mechanism of the large catalytic RNAs: group I and group II introns and ribonuclease P. The RNA world. Cold Spring Harbor Press, Cold Spring Harbor, pp 239–270
Cech TR, Herschlag D, Piccirilli JA, Pyle AM (1992) RNA catalysis by a group I ribozyme; developing a model for transition state stabilization. J Biol Chem 267: 17479–17482
Chanfreau G, Jacquier A (1993) Interaction of intronic boundaries is required for the second splicing step efficiency of a group II intron. EMBO 12: 5173–5180
Chanfreau G, Jacquier A (1994) Catalytic site components common to both splicing steps of a group II intron. Science 266: 1383–1387
Chanfreau G, Jacquier A (1996) An RNA conformational change between the two chemical steps of group II self-splicing. EMBO J (jiadan)
Chin K, Pyle AM (1995) Branch-point attack in group II introns is a highly reversible transesterification, providing a possible proof-reading mechanism for 5’-splice site selection. RNA 1: 391–406
Copertino DW, Shigeoka S, Hallick RB (1992) Chloroplast group III twintron excision utilizing multiple 5’- and 3’-splice sites. EMBO J 11: 5041–5050
Costa M, Michel F (1995) Frequent use of the same tertiary motif by self-folding RNAs. EMBO 14: 1276–1285
Daniels D, Michels WJ, Pyle AM (1996) Two competing pathways for self-splicing by group II introns; a quantitative analysis of in-vitro reaction rates and products. J Mol Biol 256: 31–99
Dib-Hajj SD, Boulanger SC, Hebbar SK, Peebles CL, Franzen JS, Perlman PS (1993) Domain 5 interacts with domain 6 and influences the second transesterification reaction of group II intron self-splicing. Nucl Acids Res 21: 1797–1804
Fabrizio P, Abelson J (1990) Two domains of yeast U6 small nuclear RNA required for both steps of nuclear precursor mRNA splicing. Science 250: 404–409
Franzen JS, Zhang M, Peebles CL (1993) Kinetic analysis of the 5’-splice junction hydrolysis of a group II intron promoted by domain 5. Nucl Acids Res 21: 627–634
Freier SM, Kierzek R, Jaeger JA, Sugimoto N, Caruthers MA, Neilson T, Turner DH (1986) Improved free energy parameters for predictions of RNA duplex stability. Proc Natl Acad Sci USA 83: 9373–9377
Griffin EA, Qin Z-F, Michels WA, Pyle AM (1995) Group II intron ribozymes that cleave DNA and RNA linkages with similar efficiency, and lack contacts with substrate 2′-hydroxyl groups. Chemistry + Biology 2: 761–770
Halbreich A, Pajot P, Foucher M, Grandchamp C, Slonimski P (1980) A pathway of cytochrome b mRNA processing in yeast mitochondria: specific splicing steps and an intron-derived circular RNA. Cell 19: 321–329
Harris ME, Pace NR (1995) Identification of phosphates involved in catalysis by the ribozyme RNase P RNA. RNA 1: 210–218
Harris-Kerr CL, Zhang M, Peebles CL (1993) The phylogenetically predicted base-pairing interaction between α and α is required for group II splicing in vitro. Proc Natl Acad Sci USA 90: 10658–10662
Hensgens LA, Amberg AC, Roosendaal E, van der Horst G, van der Veen R, van Ommen GJB, Grivell LA (1982) Variation, transcription, and circular RNAs of the mitochondrial gene for subunit 1 of cytochrome c oxidase. J Mol Biol 164: 35–58
Herschlag D (1991) Implications of ribozyme kinetics for targeting the cleavage of specific RNA molecules in vivo: more isn’t always better. Proc Natl Acad Sci USA 88: 6921–6925
Herschlag D, Cech TR (1990a) Catalysis of RNA cleavage by the Tetrahymena thermophila ribozyme 1. Kinetic description of an RNA substrate complementary to the active site. Biochemistry 29: 10159–10171
Herschlag D, Cech TR (1990b) DNA cleavage catalyzed by the ribozyme from Tetrahymena. Nature 344: 405–409
Herschlag D, Piccirilli JA, Cech TR (1991) Ribozyme-catalyzed and nonenzymatic reactions of phosphate esters: rate effects upon substitution of sulfur for a nonbridging phosphoryl oxygen atom. Biochemistry 30: 4844–4854
Herschlag D, Eckstein F, Cech TR (1993) Contributions of 2′-hydroxyl groups of an RNA substrate to binding and catalysis by the Tetrahymena ribozyme. An energetic picture of an active site composed of RNA. Biochemistry 32: 8299–8311
Heus HA, Pardi A (1991) Structural features that give rise to the unusual stability of RNA hairpins containing GNRA loops. Science 253: 191–194
Hornig H, Aebi M, Weissman C (1986) Effect of mutations at the lariat branch acceptor site on β-globin pre-mRNA splicing in vitro. Nature 324: 589–591
Jacquier A, Jacquesson-Breuleux N (1991) Splice site selection and role of the lariat in a group II intron. J Mol Biol 219: 415–428
Jacquier A, Michel F (1987) Multiple exon-binding sites in class II self-splicing introns. Cell 50: 17–29
Jacquier A, Michel F (1990) Base-pairing interactions involving the 5′- and 3′-terminal nucleotides of group II self-splicing introns. J Mol Biol 213: 437–447
Jacquier A, Rosbash M (1986) Efficient trans-splicing of a yeast mitochondrial RNA Group II intron implicates a strong 5’-exon-intron interaction. Science 234: 1099–1104
Jarrell KA, Dietrich RC, Perlman PS (1988) Group II intron domain 5 facilitates a trans-splicing reaction. Mol Cell Biol 8: 2361–2366
Koch JL, Boulanger SC, Dib-Hajj SD, Hebbar SK, Perlman PS (1992) Group II Introns deleted for multiple substructures retain self-splicing activity. Mol Cell Biol 12: 1950–1958
Kwakman JH, Konings DA, Hogeweg P, Pel HJ, Grivell LA (1990) Structural analysis of a group II intron by chemical modifications and minimal energy calculations. J Biomol Struct Dyn 8: 413–430
Kwakman JHJM, Konings D, Pel HJ, Grivell LA (1989) Structure-function relationships in a self-splicing group II intron: a large part of domain II of the mitochondrial intron aI5 is not essential for splicing. Nucl Acids Res 17: 4205–4216
Lambowitz AM, Belfort M (1993) Introns as mobile genetic elements. Annu Rev Biochem 62: 587–622
Lambowitz AM, Perlman PS (1990) Involvement of aminoacyl-tRNA synthetases and other proteins in group I and group II intron splicing. TIBS 15: 440–444
Liu Q, Pyle AM (1995) The role of branch-point nucleotide identity in splicing by group II introns (in preparation)
Madhani HD, Guthrie C (1992) A novel base-pairing interaction between U2 and U6 snRNAs suggests a mechanism for the catalytic activation of the spliceosome. Cell 71: 803–817
Major F, Turcotte M, Gautheret D, Lapalme G, Fillion E, Cedergren R (1991) The combination of symbolic and numerical computation for three-dimensional modeling of RNA. Science 253: 1255–1260
Maschhoff KL, Padgett RA (1993) The stereochemical course of the first step of premRNA splicing. Nucl Acids Res 21: 5456–5462
Michel F, Ferat J-L (1995) Structure and activities of group II introns. Annu Rev Biochem 64: 435–461
Michel F, Jacquier A, Dujon B (1982) Comparison of fungal mitochondrial introns reveals extensive homologies in RNA secondary structure. Biochimie 64: 867–881
Michel F, Umesono K, Ozeki H (1989) Comparative and functional anatomy of group II catalytic introns — a review. Gene 82: 5–30
Michels WJ, Pyle AM (1995) Conversion of a group II intron into a new multiple-turnover ribozyme that selectively cleaves oligonucleotides: elucidation of reaction mechanism and structure/function relationships. Biochemistry 34: 2965–2977
Michels WJ, Liu Q, Pyle AM (1995) A family of highly efficient group II intron ribozymes. (in preparation)
Moore MJ, Sharp PA (1992) Site-specific modification of pre-mRNA: the 2’-hydroxyl groups at the splice sites. Science 256: 992–997
Moore MJ, Query CC, Sharp PA (1993) Splicing of precursors to messenger RNAs by the spliceosome. The RNA World. Cold Spring Harbor Press, Cold Spring Harbor, pp 303–357
Mörl M, Schmelzer C (1990) Integration of group II intron bIl into a foreign RNA by reversal of the self-splicing reaction in vitro. Cell 60: 629–636
Mörl M, Niemer I, Schmelzer C (1992) New reactions catalyzed by a group II intron ribozyme with RNA and DNA substrates. Cell 70: 803–810
Müller MW, Stocker P, Hetzer M, Schweyen RJ (1991) Fate of the junction phosphate in alternating forward and reverse self-splicing reaction of group II intron RNA. J Mol Biol 222: 145–150
Müller MW, Hetzer M, Schweyen RJ (1993) Group II intron RNA catalysis of progressive nucleotide insertion: a model for RNA editing. Nature 261: 1035–1038
Ott G, Arnold L, Limmer S (1993) Proton NMR studies of manganese ion binding to tRNA-derived acceptor arm duplexes. Nucl Acids Res 21: 5859–5864
Pace NR, Smith D (1990) Ribonuclease P: function and variation. J Biol Chem 265: 3587–3590
Padgett RA, Konarska MM, Grabowski PJ, Hardy SF, Sharp PA (1984) Lariat RNAs as intermediates and products in the splicing of messenger RNA precursors. Science 225: 898–903
Padgett RA, Podar M, Boulanger SC, Perlman PS (1994) The stereochemical course of group II intron self-splicing. Science 266: 1685–1688
Pan T, Long DM, Uhlenbeck OC (1993) Divalent metal ions in RNA folding and catalysis. The RNA world. Cold Spring Harbor Press, Cold Spring Harbor, pp 271–302
Peebles CL, Perlman PS, Mecklenburg KL, Petrillo ML, Tabor JH, Jarrell KA, Cheng H-L (1986) A self-splicing RNA excises an intron lariat. Cell 44: 213–223
Peebles CL, Benatan EJ, Jarrell KA, Perlman PS (1987) Group II intron self-splicing: development of alternative reaction conditions and identification of a predicted intermediate. Cold Spring Harbor Symp Quant Biol 52: 223–232
Peebles CL, Belcher SM, Zhang M, Dietrich RC, Perlman PC (1993) Mutation of the conserved first nucleotide of a group II intron from yeast mitochondrial DNA reduces the rate but allows accurate splicing. J Biol Chem 268: 11929–11938
Peebles CL, Zhang M, Perlman PS, Franzen JF (1995) Identification of a catalytically critical trinucleotide in domain 5 of a group II intron. Proc Natl Acad Sci USA 92: 4422–4426
Perreault J-P, Altman S (1992) Important 2′-hydroxyl groups in model substrates for M1 RNA, the catalytic RNA subunit of RNase P from E. coli. J Mol Biol 226: 399–409
Pley HM, Flaherty KM, McKay DB (1994) Model for an RNA tertiary interaction from the structure of an intermolecular complex between a GAAA tetraloop and an RNA helix. Nature 372: 111–113
Podar M, Perlman PS, Padgett RA (1995a) Stereochemical selectivity of group II intron splicing, reverse-splicing and hydrolysis reactions. Mol Cell Biol 15: 4466–4478
Podar M, Dib-Hajj S, Perlman PS (1995b) A UV-induced Mgt+-dependent cross-link traps an active form of domain 3 of a self-splicing group II intron. RNA 1: 828–840
Pyle AM (1993) Ribozymes: a distinct class of metalloenzymes. Science 261: 709–714
Pyle AM (1995) The role of metal ions in ribozymes. Metal ions in biological systems, vol 32. Marcel Dekker, New York, pp 479–520
Pyle AM, Cech TR (1991) Ribozyme recognition of RNA by tertiary interactions with specific ribose 2’-OH groups. Nature 350: 628–631
Pyle AM, Green JB (1994) Building a kinetic framework for group II intron ribozyme activity: quantitation of interdomain binding and reaction rate. Biochemistry 33: 2716–2725
Pyle AM, Green JB (1995) RNA folding. Curr Opin Struct Biol 5: 303–310
Pyle AM, Murphy FL, Cech TR (1992) Ribozyme substrate binding site in the catalytic core of the Tetrahymena ribozyme. Nature 358: 123–128
Schmelzer C, Schweyen RJ (1986) Self-splicing of group II introns in vitro: mapping of the branch point and mutational inhibition of lariat formation. Cell 46: 557–565
Schmelzer C, Schmidt C, Schweyen RJ (1982) Identification of splicing signals in introns of yeast mitochondrial split genes: mutational alterations in intron bIl and secondary structures in related introns. Nucl Acids Res 10: 6797–6808
Schmidt U, Kosack M, Stahl U (1987) Lariat RNA of a group II intron in a filamentous fungus. Curr Genet 12: 291–295
Schmidt U, Sägebarth R, Schmelzer C, Stahl U (1993) Self-splicing of a Podospora anserina group IIA intron in vitro. J Mol Biol 231: 559–568
Schmidt WM, Schweyen RJ, Wolf K, Mueller MW (1994) Transposable group II introns in fission and budding yeast. Site-specific genomic instabilities and formation of group II IVS p1DNAs. J Mol Biol 243: 157–166
Sharp PA (1985) On the origin of RNA splicing and introns. Cell 42: 397–400
Smith D, Pace NR (1993) Multiple magnesium ions in the ribonuclease P reaction mechanism. Biochemistry 32: 5273–5281
Smith D, Burgin AB, Haas ES, Pace NR (1992) Influence of metal ions on the ribonuclease P reaction; distinguishing binding from catalysis. J Biol Chem 267: 2429–2436
Steitz TA, Steitz JA (1993) A general two-metal ion mechanism for catalytic RNA. Proc Natl Acad Sci USA 90: 6498–6502
Strobel SA, Cech TR (1993) Tertiary interactions with the internal guide sequence mediate docking of the P1 helix into the catalytic core of the Tetrahymena ribozyme. Biochemistry 32: 13593–13604
Suchy M, Schmelzer C (1991) Restoration of the self-splicing activity of a defective group II intron by a small trans-acting RNA. J Mol Biol 222: 179–187
Sullenger BA, Cech TR (1994) Ribozyme-mediated repair of defective mRNA by targeted trans-splicing. Nature 371: 619–622
Van der Veen R, Amberg AC, van der Horst G, Bonen L, Tabak HF, Grivell LA (1986) Excised group II introns in yeast mitochondria are lariats and can be formed by self-splicing in vitro. Cell 44: 225–234
Van der Veen R, Amberg AC, Grivell LA (1987a) Self-splicing of a group II intron in yeast mitochondria: dependence on 5’exon sequences. EMBO J 6: 1079–1084
Van der Veen R, Kwakman JHJM, Grivell LA (1987b) Mutations at the lariat acceptor site allow self-splicing of a group II intron without lariat formation. EMBO J 6: 3827–3831
Van Dyck E, Jank B, Ragnini A, Schweyen RJ, Duyckaerts S, Sluse F, Foury F (1995) Overexpression of a novel member of the mitochondrial carrier family rescues defects in both DNA and RNA metabolism in yeast mitochondria. Mol Gen Genet 246: 426–436
Waldherr M, Ragnin A, Jank B, Teply R, Wiesenberger G, Schweyen RJ (1993) A multitude of suppressors of group II intron-splicing defects in yeast. Curr Genet 24: 301–306
Weeks KM, Crothers DM (1993) Major groove accessibility of RNA. Science 261: 1574–1577
Wiesenberger G, Waldherr M, Schweyen RJ (1992) The nuclear gene MRS2 is essential for the excission of group II introns from yeast mitochondrial transcripts in vivo. J Biol Chem 267: 6963–6969
Wissinger B, Brennicke A, Schuster W (1992) Regenerating good sense: RNA editing and trans-splicing in plant mitochondria. Trends Genet 8: 322–328
Yu Y-T, Maroney PA, Darynkiewicz E, Nilsen TW (1995) U6 snRNA function in nuclear pre-mRNA splicing: a phosphorothioate interference analysis of the U6 phosphate backbone. RNA 1: 46–54
Zaug AI, Grosshans CA, Cech TR (1988) Sequence-specific endoribonuclease activity of the Tetrahymena ribozyme: enhanced cleavage of certain oligonucleotide substrates that form mismatched ribozyme-substrate complexes. Biochemistry 27: 8924–8931
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 1996 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Pyle, A.M. (1996). Catalytic Reaction Mechanisms and Structural Features of Group II Intron Ribozymes. In: Eckstein, F., Lilley, D.M.J. (eds) Catalytic RNA. Nucleic Acids and Molecular Biology, vol 10. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-61202-2_5
Download citation
DOI: https://doi.org/10.1007/978-3-642-61202-2_5
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-540-62679-4
Online ISBN: 978-3-642-61202-2
eBook Packages: Springer Book Archive