Abstract
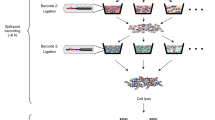
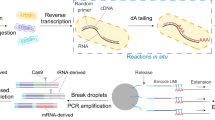
Microbes exhibit an extraordinary capacity to adapt their physiology to different environments using phenotypic heterogeneity. However, the majority of gene regulation studies are conducted in bulk reflecting only averaged gene expression levels across millions of cells. Bacterial single-cell RNA-seq (scRNA-seq) can overcome this by enabling whole transcriptome and unbiased analysis of microbes at the single-cell level. Here, we describe a detailed workflow of single-cell RNA-seq based on the multiple annealing and dC-tailing-based quantitative single-cell RNA-seq (MATQ-seq) protocol. Following adjustments to the original eukaryotic protocol, the workflow was applied to two major human pathogens Salmonella enterica serovar Typhimurium (henceforth Salmonella) and Pseudomonas aeruginosa (henceforth Pseudomonas). The development of bacterial scRNA-seq protocols offers promising avenues to explore the molecular programs underlying phenotypic heterogeneity on the transcriptome level in different settings such as infection, persistence, ecology, and biofilms.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Ackermann M (2015) A functional perspective on phenotypic heterogeneity in microorganisms. Nat Rev Microbiol 13(8):497–508. https://doi.org/10.1038/nrmicro3491
Beebout CJ, Eberly AR, Werby SH, Reasoner SA, Brannon JR, De S, Fitzgerald MJ, Huggins MM, Clayton DB, Cegelski L, Hadjifrangiskou M (2019) Respiratory heterogeneity shapes biofilm formation and host colonization in Uropathogenic Escherichia coli. MBio 10(2). https://doi.org/10.1128/mBio.02400-18
Yannarell SM, Velickovic D, Anderton CR, Shank EA (2021) Direct visualization of chemical cues and cellular phenotypes throughout Bacillus subtilis biofilms. mSystems 6(6):e0103821. https://doi.org/10.1128/mSystems.01038-21
Lyu Z, Yang A, Villanueva P, Singh A, Ling J (2021) Heterogeneous flagellar expression in single Salmonella cells promotes diversity in antibiotic tolerance. mBio 12:e0237421. https://doi.org/10.1128/mBio.02374-21
Jones EC, Uphoff S (2021) Single-molecule imaging of LexA degradation in Escherichia coli elucidates regulatory mechanisms and heterogeneity of the SOS response. Nat Microbiol 6(8):981–990. https://doi.org/10.1038/s41564-021-00930-y
Merida-Floriano A, Rowe WPM, Casadesus J (2021) Genome-wide identification and expression analysis of SOS response genes in Salmonella enterica Serovar Typhimurium. Cells 10(4):943. https://doi.org/10.3390/cells10040943
Roche B, Bumann D (2021) Single-cell reporters for pathogen responses to antimicrobial host attacks. Curr Opin Microbiol 59:16–23. https://doi.org/10.1016/j.mib.2020.07.013
Tanay A, Regev A (2017) Scaling single-cell genomics from phenomenology to mechanism. Nature 541(7637):331–338. https://doi.org/10.1038/nature21350
Imdahl F, Saliba AE (2020) Advances and challenges in single-cell RNA-seq of microbial communities. Curr Opin Microbiol 57:102–110. https://doi.org/10.1016/j.mib.2020.10.001
Avital G, Avraham R, Fan A, Hashimshony T, Hung DT, Yanai I (2017) scDual-Seq: mapping the gene regulatory program of Salmonella infection by host and pathogen single-cell RNA-sequencing. Genome Biol 18(1):200. https://doi.org/10.1186/s13059-017-1340-x
Betin V, Penaranda C, Bandyopadhyay N, Yang R, Abitua A, Bhattacharyya RP, Fan A, Avraham R, Livny J, Shoresh N, Hung DT (2019) Hybridization-based capture of pathogen mRNA enables paired host-pathogen transcriptional analysis. Sci Rep 9(1):19244. https://doi.org/10.1038/s41598-019-55633-6
Kuchina A, Brettner LM, Paleologu L, Roco CM, Rosenberg AB, Carignano A, Kibler R, Hirano M, RW DP, Seelig G (2021) Microbial single-cell RNA sequencing by split-pool barcoding. Science 371(6531):eaba5257. https://doi.org/10.1126/science.aba5257
Blattman SB, Jiang W, Oikonomou P, Tavazoie S (2020) Prokaryotic single-cell RNA sequencing by in situ combinatorial indexing. Nat Microbiol 5(10):1192–1201. https://doi.org/10.1038/s41564-020-0729-6
Imdahl F, Vafadarnejad E, Homberger C, Saliba AE, Vogel J (2020) Single-cell RNA-sequencing reports growth-condition-specific global transcriptomes of individual bacteria. Nat Microbiol 5(10):1202–1206. https://doi.org/10.1038/s41564-020-0774-1
Sheng K, Cao W, Niu Y, Deng Q, Zong C (2017) Effective detection of variation in single-cell transcriptomes using MATQ-seq. Nat Methods 14(3):267–270. https://doi.org/10.1038/nmeth.4145
Zong C, Lu S, Chapman AR, Xie XS (2012) Genome-wide detection of single-nucleotide and copy-number variations of a single human cell. Science 338(6114):1622–1626. https://doi.org/10.1126/science.1229164
Kroeger C, Colgan A, Srikumar S, Handler K, Sivasankaran SK, Hammarlof DL, Canals R, Grissom JE, Conway T, Hokamp K, Hinton JC (2013) An infection-relevant transcriptomic compendium for Salmonella enterica Serovar Typhimurium. Cell Host Microbe 14(6):683–695. https://doi.org/10.1016/j.chom.2013.11.010
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Homberger, C., Saliba, AE., Vogel, J. (2023). A MATQ-seq-Based Protocol for Single-Cell RNA-seq in Bacteria. In: Calogero, R.A., Benes, V. (eds) Single Cell Transcriptomics. Methods in Molecular Biology, vol 2584. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2756-3_4
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2756-3_4
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2755-6
Online ISBN: 978-1-0716-2756-3
eBook Packages: Springer Protocols