Abstract
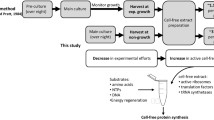
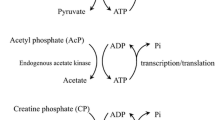
With the rapid development of cell-free biotechnology, more and more cell-free protein synthesis (CFPS) systems have been established and optimized for protein expression in vitro. Here, we aim to improve the productivity of a newly developed Streptomyces-based CFPS system. Protein translation in CFPS systems depends on the entire endogenous translation system from cell lysates. However, lysates might lack such translation-related elements, limiting the efficiency of protein translation and therefore the productivity of CFPS systems. To address this limitation, we sought to add protein translation related factors to CFPS reactions. By doing this, the protein yield of EGFP was significantly improved up to approximately 400 μg/mL. In this chapter, we mainly describe the preparation of Streptomyces cell extracts, expression and purification of nine translation related factors, and optimization of the Streptomyces-based CFPS system for enhanced protein expression.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Min SE, Lee KH, Park SW et al (2016) Cell-free production and streamlined assay of cytosol-penetrating antibodies. Biotechnol Bioeng 113:2107–2112
Schwarz D, Junge F, Durst F et al (2007) Preparative scale expression of membrane proteins in Escherichia coli-based continuous exchange cell-free systems. Nat Protoc 2:2945–2957
Li J, Lawton TJ, Kostecki JS et al (2016) Cell-free protein synthesis enables high yielding synthesis of an active multicopper oxidase. Biotechnol J 11:212–218
Martin RW, Des Soye BJ, Kwon YC et al (2018) Cell-free protein synthesis from genomically recoded bacteria enables multisite incorporation of noncanonical amino acids. Nat Commun 9:1203
Zhuang L, Huang S, Liu WQ et al (2020) Total in vitro biosynthesis of the nonribosomal macrolactone peptide valinomycin. Metab Eng 60:37–44
Carlson ED, Gan R, Hodgman CE et al (2012) Cell-free protein synthesis: applications come of age. Biotechnol Adv 30:1185–1194
Liu WQ, Zhang L, Chen M et al (2019) Cell-free protein synthesis: recent advances in bacterial extract sources and expanded applications. Biochem Eng J 141:182–189
Silverman AD, Karim AS, Jewett MC (2020) Cell-free gene expression: an expanded repertoire of applications. Nat Rev Genet 21:151–170
Li J, Wang H, Kwon YC et al (2017) Establishing a high yielding Streptomyces-based cell-free protein synthesis system. Biotechnol Bioeng 114:1343–1353
Li J, Wang H, Jewett MC (2018) Expanding the palette of Streptomyces-based cell-free protein synthesis systems with enhanced yields. Biochem Eng J 130:29–33
Moore SJ, Lai HE, Needham H et al (2017) Streptomyces venezuelae TX-TL - a next generation cell-free synthetic biology tool. Biotechnol J 12:1600678
Pospiech A, Neumann B (1995) A versatile quick-prep of genomic DNA from gram-positive bacteria. Trends Genet 11:217–218
Xu H, Liu WQ, Li J (2020) Translation related factors improve the productivity of a Streptomyces-based cell-free protein synthesis system. ACS Synth Biol 9:1221–1224
Acknowledgments
This work was supported by grants from the National Natural Science Foundation of China (No. 31971348 and No. 31800720) and the Natural Science Foundation of Shanghai (19ZR1477200).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Xu, H., Liu, WQ., Li, J. (2022). A Streptomyces-Based Cell-Free Protein Synthesis System for High-Level Protein Expression. In: Karim, A.S., Jewett, M.C. (eds) Cell-Free Gene Expression. Methods in Molecular Biology, vol 2433. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1998-8_5
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1998-8_5
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1997-1
Online ISBN: 978-1-0716-1998-8
eBook Packages: Springer Protocols