Abstract
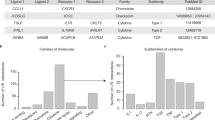
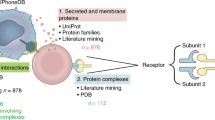
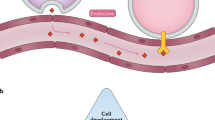
Cell–cell communication is crucial for development and tissue homeostasis in multicellular organisms. Single-cell transcriptomics has emerged as a revolutionary technique for dissecting cellular compositions and potential cell–cell communication events via ligand–receptor pairs. To provide a systematic characterization of intercellular communication, we developed a framework to map cell–cell communication events mediated by ligand–receptor interactions across different cell types using single-cell transcriptomics data. Our repository of ligands, receptors and their interactions is integrated with a computational approach to identify cell-type specific and biologically relevant interactions. Here, we summarize the structure and content of our repository and present a practical guide for inferring cell–cell communication networks from single-cell RNA sequencing data.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Camp JG et al (2017) Multilineage communication regulates human liver bud development from pluripotency. Nature 546:533–538
Vento-Tormo R et al (2018) Single-cell reconstruction of the early maternal-fetal interface in humans. Nature 563:347–353
Efremova M, Vento-Tormo M, Teichmann SA, Vento-Tormo R (2020) CellPhoneDB: inferring cell-cell communication from combined expression of multi-subunit ligand-receptor complexes. Nat Protoc 15:1484–1506
Satija R, Farrell JA, Gennert D, Schier AF, Regev A (2015) Spatial reconstruction of single-cell gene expression data. Nat Biotechnol 33:495–502
Wolf FA, Angerer P, Theis FJ (2018) SCANPY: large-scale single-cell gene expression data analysis. Genome Biol 19:15
Hie B, Cho H, DeMeo B, Bryson B, Berger B (2019) Geometric sketching compactly summarizes the single-cell transcriptomic landscape. Cell Syst 8:483–493
Acknowledgments
We thank Sarah Teichmann for useful guidance on the method and curation of protein–protein interactions; Gavin J. Wright, Laura Wood, and Gerard Graham for advice on protein–protein interactions; and Miquel Vento-Tormo and YDEVS members for their help with the webserver and the implementation of the code in github. M.E. is funded by a Barts Charity Lectureship (grant MGU045). The project was supported by Wellcome Sanger core funding (no. WT206194) and a Wellcome Strategic Support Science award (no. 211276/Z/18/Z).
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Efremova, M., Vento-Tormo, R. (2021). Inference of Ligand–Receptor Pairs from Single-Cell Transcriptomics Data. In: Turksen, K. (eds) Stem Cell Renewal and Cell-Cell Communication. Methods in Molecular Biology, vol 2346. Humana, New York, NY. https://doi.org/10.1007/7651_2020_343
Download citation
DOI: https://doi.org/10.1007/7651_2020_343
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1569-0
Online ISBN: 978-1-0716-1570-6
eBook Packages: Springer Protocols