Abstract
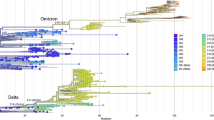
Omicron is a relatively new form of COVID-19 that has created an unavoidable and life-threatening situation to the entire world since late 2021. Absence of appropriate vaccination, medication, the epidemiological cycle has become more complex. This study primarily concentrates on the analysis of genome sequence for COVID-19 variants. To conduct such analysis, two datasets are collected from Kaggle and GISAID. Using these datasets, the globally existing genome sequences are identified and insights regarding the countries that are carrying significantly higher genome sequence count are provided. This investigation analyzes the worldwide virus variants and further identifies that the United States and United Kingdom are the countries where proper inspection should be provided because of the genome sequence count. An adequate idea regarding the mutations of the Omicron virus is also considered in this study. To address this issue, recent genome sequence data ranging from February, 2022 to 10th March, 2022 is analyzed to understand how the latest arrival, Omicron, is perturbing the world. This study emphasizes on the constant surveillance of genome sequences among all the countries which in turn will benefit the health care professionals and frontline healthcare workers as well as the Governments can take necessary policies and precautions to combat such pandemic.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
G.C. Calafiore, C. Novara, C. Possieri, A time-varying SIRD model for the COVID-19 contagion in Italy. Annu. Rev. Control. 50, 361–372 (2020). https://doi.org/10.1016/j.arcontrol.2020.10.005
E. Callaway, The coronavirus is mutating—does it matter? Nature 585(7824), 174–177 (2020). https://doi.org/10.1038/d41586-020-02544-6
J. Chu, A statistical analysis of the novel coronavirus (COVID-19) in Italy and Spain. PLoS ONE 16(3), e0249037 (2021). https://doi.org/10.1371/journal.pone.0249037
S.J. Fong, G. Li, N. Dey, R. Gonzalez-Crespo, E. Herrera-Viedma, Finding an accurate early forecasting model from small dataset: a case of 2019-nCoV novel coronavirus outbreak. Int. J. Interact. Multimedia Artif. Intell. 6(1), 132 (2020). https://doi.org/10.9781/ijimai.2020.02.002
E. Gambhir, R. Jain, A. Gupta, U. Tomer, Regression analysis of COVID-19 using machine learning algorithms, in 2020 International Conference on Smart Electronics and Communication (ICOSEC) (2020). https://doi.org/10.1109/icosec49089.2020.9215356
GISAID—initiative. GISAID. https://www.gisaid.org/
D.W. Hawman, K. Meade-White, C. Clancy, J. Archer, T. Hinkley, S.S. Leventhal, D. Rao, A. Stamper, M., R. Rosenke, K. Krieger, S. Randall, A.P. Khandhar, L. Hao, T.Y. Hsiang, A.L. Greninger, M. Gale, P. Berglund, D.H. Fuller, J.H. Erasmus, et al.: Replicating RNA platform enables rapid response to the SARS-CoV-2 Omicron variant and elicits enhanced protection in naïve hamsters compared to ancestral vaccine. Preprint at bioRxiv (2022). https://doi.org/10.1101/2022.01.31.478520
J.A. Hay, S.M. Kissler, J.R. Fauver, C. Mack, C.G. Tai, R.M. Samant, S. Connelly, D.J. Anderson, G. Khullar, M. MacKay, M. Patel, S. Kelly, A. Manhertz, I. Eiter, D. Salgado, T. Baker, B. Howard, J.T. Dudley, C.E. Mason, Y.H. Grad et al., Viral dynamics and duration of PCR positivity of the SARS-CoV-2 Omicron variant. medRxiv Preprint (2022). https://doi.org/10.1101/2022.01.13.22269257
P. Hemarajata, SARS-CoV-2 Sequencing Data: The Devil Is in the Genomic Detail (ASM.Org., 2020). https://asm.org/Articles/2020/October/SARS-CoV-2-Sequencing-Data-The-Devil-Is-in-the-Gen. Accessed 28 Oct 2020
C.J. Houldcroft, M.A. Beale, J. Breuer, Clinical and biological insights from viral genome sequencing. Nat. Rev. Microbiol. 15(3), 183–192 (2017). https://doi.org/10.1038/nrmicro.2016.182
J.J. Huamán-Saavedra, The Omicron variant of SARS-CoV-2. Rev. Méd. Trujillo 17(1), 3–4 (2022). https://doi.org/10.17268/rmt.2022.v17i1.4256
A.L. Lundberg, R. Lorenzo-Redondo, E.A. Ozer, C.A. Hawkins, J.F. Hultquist, S.B. Welch, P.V. Prasad, J.F. Oehmke, C.J. Achenbach, R.L. Murphy, J.I. White, R.J. Havey, L.A. Post, Has Omicron changed the evolution of the pandemic? JMIR Publ. Health Surveill. 8(1), e35763 (2022). https://doi.org/10.2196/35763
S. Mallapaty, COVID-19: how Omicron overtook delta in three charts. Nature (2022). https://doi.org/10.1038/d41586-022-00632-3
T. Nyberg, M. Ferguson, S.G. Nash, H.H. Webster, S. Flaxman, N. Andrews, W. Hinsley, J.L. Bernal, M. Kall, S. Bhatt, P. Blomquist, A. Zaidi, E. Volz, N.A. Aziz, K. Harman, S. Funk, S. Abbott, R. Hope, A. Charlett, S. Thelwall et al., Comparative analysis of the risks of hospitalisation and death associated with SARS-CoV-2 Omicron (B.1.1.529) and delta (B.1.617.2) variants in England: a cohort study. Lancet (2022). https://doi.org/10.1016/s0140-6736(22)00462-7
Omicron daily cases by country (COVID-19 variant). Kaggle. https://www.kaggle.com/datasets/yamqwe/omicron-covid19-variant-daily-cases. Accessed 16 Feb 2022
A.J. Pollard, E.M. Bijker, A guide to vaccinology: from basic principles to new developments. Nat. Rev. Immunol. 21, 83–100 (2021). https://doi.org/10.1038/s41577-020-00479-7
Why genomic sequencing is crucial in COVID-19 response: WHO | Regional Office for Africa. https://www.afro.who.int/news/why-genomic-sequencing-crucial-covid-19-response. Accessed 18 Mar 2022
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this paper
Cite this paper
Dutta, S., Bandyopadhyay, S.K., Janarthanan, M., Bose, P. (2023). A Novel SARS-COV-2 Variant Omicron Disseminating Evaluation. In: Reddy, A.B., Nagini, S., Balas, V.E., Raju, K.S. (eds) Proceedings of Third International Conference on Advances in Computer Engineering and Communication Systems. Lecture Notes in Networks and Systems, vol 612. Springer, Singapore. https://doi.org/10.1007/978-981-19-9228-5_5
Download citation
DOI: https://doi.org/10.1007/978-981-19-9228-5_5
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-19-9227-8
Online ISBN: 978-981-19-9228-5
eBook Packages: Intelligent Technologies and RoboticsIntelligent Technologies and Robotics (R0)