Abstract
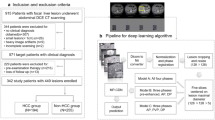
Dynamic contrast-enhanced magnetic resonance imaging provide not only the information on the morphological features of the lesions, but also the changes of the lesion’s blood perfusion. In this paper, we propose a tensor-based temporal data representation (TTD) model and a multi-channel fusion 3D convolutional neural network (MCF-3D CNN) to extract the temporal and spatial features of dynamic contrast enhanced-MR images (DCE-MR images). To evaluate the performance of the proposed methods, we established a DCE-MR image dataset for non-invasively assessing the differentiation of Hepatocellular carcinoma (HCC). The TTD model achieves the accuracy of 73.96% for non-invasive assessment of HCC differentiation via MCF-3D CNN. Meanwhile, the 3D CNN with TTD achieves accuracy, sensitivity and specificity of 95.17%, 96.33%, and 94.00%, respectively, in discriminating the HCC and cirrhosis. Compared with the normal data representation method, the proposed TTD method is more conducive for 3D CNN to extract temporal-spatial features of DCE-MR images.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Krizhevsky, A., Sutskever, I., Hinton, G.E.: ImageNet classification with deep convolutional neural networks. In: International Conference on Neural Information Processing Systems, pp. 1097–1105 (2012)
Liu, W., et al.: SSD: single shot MultiBox detector. In: Leibe, B., Matas, J., Sebe, N., Welling, M. (eds.) ECCV 2016. LNCS, vol. 9905, pp. 21–37. Springer, Cham (2016). https://doi.org/10.1007/978-3-319-46448-0_2
He, K., Gkioxari, G., Dollár, P., Girshick, R.B.: Mask R-CNN. CoRR, abs/1703.06870 (2017)
Ji, S., Xu, W., Yang, M., Yu, K.: 3D convolutional neural networks for human action recognition. IEEE Trans. Pattern Anal. Mach. Intell. 35, 221–231 (2013)
Litjens, G., et al.: A survey on deep learning in medical image analysis. Med. Image Anal. 42, 60 (2017)
Dou, Q., et al.: Automatic detection of cerebral microbleeds from mr images via 3D convolutional neural networks. IEEE Trans. Med. Imaging 35, 1182–1195 (2016)
Lemaître, G., Martí, R., Freixenet, J., Vilanova, J.C., Walker, P.M., Meriaudeau, F.: Computer-aided detection and diagnosis for prostate cancer based on mono and multi-parametric MRI: a review. Comput. Biol. Med. 60, 8 (2015)
Kamnitsas, K., et al.: Efficient multi-scale 3D CNN with fully connected CRF for accurate brain lesion segmentation. Med. Image Anal. 36, 61 (2017)
Yang, J.D., Roberts, L.R.: Hepatocellular carcinoma: a global view. Nat. Rev. Gastroenterol. Hepatol. 7, 448 (2010)
Sherman, M.: Hepatocellular carcinoma: screening and staging. Clin. Liver Dis. 15, 323–334 (2011)
Regimbeau, J.M., et al.: Risk factors for early death due to recurrence after liver resection for hepatocellular carcinoma: Results of a multicenter study †. J. Surg. Oncol. 85, 36–41 (2004)
Jiang, T., Zou, Y., Jie-Hua, X.U., Peng, L.R., Shan, H., Radiology, D.O.: Magnetic resonance imaging in predicting histopathological differentiation of hepatocellular carcinoma. J. Sun Yat-Sen Univ. Sci. 37, 903–911 (2016)
Fabijańska, A.: A novel approach for quantification of time-intensity curves in a DCE-MRI image series with an application to prostate cancer. Comput. Biol. Med. 73, 119–130 (2016)
Li, J., Fan, M., Zhang, J., Li, L.: Discriminating between benign and malignant breast tumors using 3D convolutional neural network in dynamic contrast enhanced-MR images. In: SPIE Medical Imaging, p. 1013808 (2017)
Zhou, W., et al.: Malignancy characterization of hepatocellular carcinomas based on texture analysis of contrast-enhanced MR images. J. Magn. Reson. Imaging JMRI 45, 1476–1484 (2017)
Lecun, Y., Bengio, Y., Hinton, G.: Deep learning. Nature 521, 436 (2015)
Glorot, X., Bordes, A., Bengio, Y.: Deep sparse rectifier neural networks. In: International Conference on Artificial Intelligence and Statistics, pp. 315–323 (2011)
Kingma, D.P., Ba, J.L.: Adam: a method for stochastic optimization. In: International Conference for Learning Representations (2015)
Srivastava, N., Hinton, G.E., Krizhevsky, A., Sutskever, I., Salakhutdinov, R.: Dropout: a simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 15, 1929–1958 (2014)
Chen, H., Yu, L., Dou, Q., Shi, L., Mok, V.C.T., Heng, P.A.: Automatic detection of cerebral microbleeds via deep learning based 3D feature representation. In: IEEE International Symposium on Biomedical Imaging, pp. 764–767 (2015)
Buda, M., Maki, A., Mazurowski, M.A.: A systematic study of the class imbalance problem in convolutional neural networks. CoRR, abs/1710.05381 (2017)
Shen, L., Lin, Z., Huang, Q.: Relay backpropagation for effective learning of deep convolutional neural networks. In: Leibe, B., Matas, J., Sebe, N., Welling, M. (eds.) ECCV 2016. LNCS, vol. 9911, pp. 467–482. Springer, Cham (2016). https://doi.org/10.1007/978-3-319-46478-7_29
Abadi, M., et al.: TensorFlow: large-scale machine learning on heterogeneous distributed systems. arXiv: Distributed, Parallel, and Cluster Computing (2016)
Acknowledgments
This work is supported in part by the grants from Beijing Natural Science Foundation (No.7184199), Capital’s Funds for Health Improvement and Research (No. 2018-2-2023), Research Foundation of Beijing Friendship Hospital, Capital Medical University (No. yyqdkt2017-25).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Nature Singapore Pte Ltd.
About this paper
Cite this paper
Jia, X., Xiao, Y., Yang, D., Yang, Z., Wang, X., Liu, Y. (2018). Temporal-Spatial Feature Learning of Dynamic Contrast Enhanced-MR Images via 3D Convolutional Neural Networks. In: Wang, Y., Jiang, Z., Peng, Y. (eds) Image and Graphics Technologies and Applications. IGTA 2018. Communications in Computer and Information Science, vol 875. Springer, Singapore. https://doi.org/10.1007/978-981-13-1702-6_38
Download citation
DOI: https://doi.org/10.1007/978-981-13-1702-6_38
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-13-1701-9
Online ISBN: 978-981-13-1702-6
eBook Packages: Computer ScienceComputer Science (R0)