Abstract
Vaccines are the most economical and potent substitute of available medicines to cure various bacterial and viral diseases. Earlier, killed or attenuated pathogens were employed for vaccine development. But in present era, the peptide vaccines are in much trend and are favoured over whole vaccines because of their superiority over conventional vaccines. These vaccines are either based on single proteins or on synthetic peptides including several B-cell and T-cell epitopes. However, the overall mechanism of action remains the same and works by prompting the immune system to activate the specific B-cell- and T-cell-mediated responses against the pathogen. Rino Rappuoli and others have contributed in this field by plotting the design of the most potent and fully computational approach for discovery of potential vaccine candidates which is popular as reverse vaccinology. This is quite an unambiguous advance for vaccine evolution where one begins with the genome information of the pathogen and ends up with the list of certain epitopes after application of multiple bioinformatics tools. This book chapter is an effort to bring this approach of reverse vaccinology into notice of readers using example of coronavirus.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
15.1 Introduction
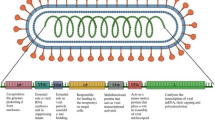
Coronaviruses are remarkably large, positive-stranded RNA viruses that are enveloped with the nucleocapsid having helical symmetry. The corona in coronavirus is a Latin word that means a “crown”, and it indicates to the typical presentation of virions underneath electron microscopy with a periphery of hefty, globular surface projections similar to that of a crown. Coronavirus is a pathogen associated with severe respiratory symptoms and was first identified from the nasal cavities of sufferers with the common cold in the early 1960s (de Groot et al. 2013; Brown et al. 2012). These were named human coronavirus OC43 and human coronavirus 229E. A total of 40 sequenced genomes of different strains of coronavirus are accessible from National Center for Biotechnology Information (NCBI), out of which 7 are pathogenic to humans. A coronavirus, i.e. SARS-CoV, was responsible for outbreak of severe acute respiratory syndrome (SARS) in the year 2003, whereas Middle East respiratory syndrome coronavirus (MERS-CoV) caused the most recent outbreak in 2012 causing acute respiratory disease in affected people with signs of fever, cough and difficulty in breathing. After first reported from Saudi Arabia in 2012, this novel virus has also dispersed to other countries like the United States and was known to have high death rate. MERS-CoV infections are highly communicable, and no explicit antiviral cure has been designed for it till date (Azhar et al. 2017).
It compelled us to apply the well-known reverse vaccinology (RV) approach on available proteome of coronavirus. RV approach has been successfully applied on many prokaryotes, but there are very few known applications on eukaryotes and viruses. So, it is worthwhile to explore the potential of this approach to identify potential vaccine candidates for coronavirus. RV basically does the in silico examination of the viral proteome to hunt antigenic and surface-exposed proteins. This approach was initially applied successfully to Neisseria meningitidis serogroup B (Kelly and Rappuoli 2005) against which none of the prevailing techniques could develop a vaccine. The present book chapter is intended to explore the potential of RV approach to select the probable vaccine candidates against coronavirus and validate the results using docking studies.
15.2 The Elementary Concept of Reverse Vaccinology
Undoubtedly, the traditional approaches for vaccine development are fortunate enough to efficiently resist the alarming pathogenic diseases of its time. However, the traditional approach suffers from certain limitations like it is very time-consuming, the pathogens which can’t be cultivated in the lab conditions are out of reach, and certain non-abundant proteins are not accessible using this approach (Rappuoli 2000). Consequently, a number of pathogenic diseases are left without any vaccine against them. All these limitations are conquered by reverse vaccinology approach utilizing genome sequence information which ultimately is translated into proteins. Hence all the proteins expressed by the genome are accessible irrespective of their abundance, conditions in which they expressed. The credit of fame of reverse vaccinology should go to the advancements in the sequencing strategies worldwide. Accordingly, improvement in the sequencing technologies has flooded the genome databases with huge amount of data which can be computationally undertaken to reveal the various crucial aspects of the virulence factors of the concerned pathogen. Reverse vaccinology is based on same approach of computationally analysing the genome of pathogen and proceeds step by step to ultimately identify the highly antigenic, secreted proteins with high epitope densities. The best epitopes are selected as potential vaccine candidates (Pizza et al. 2000). This approach has brought the unapproachable pathogens of interest in spotlight and is evolving as the most reassuring tool for precise selection of vaccine candidates and brought the use of peptide vaccines in trend (Sette and Rappuoli 2010; Kanampalliwar et al. 2013).
15.3 Successful Applications of Reverse Vaccinology
Bexsero is the first universal serogroup B meningococcal vaccine developed using RV, and it has currently earned positive judgement from the European Medicines Agency (Gabutti 2014). Whether it is discovery of pili in gram-positive pathogens which were thought to not have any pili or the sighting of factor G-binding protein in meningococcus (Alessandro and Rino 2010), the reverse vaccinology steals all the credits from other conventional approaches. Most of the applications of RV are against prokaryotes and very few against eukaryotes and viruses because of complexity of their genome. Corynebacterium urealyticum (Guimarães et al. 2015), Mycobacterium tuberculosis (Monterrubio-López et al. 2015), H. pylori (Naz et al. 2015), Acinetobacter baumannii (Chiang et al. 2015), Rickettsia prowazekii (Caro-Gomez et al. 2014), Neospora caninum (Goodswen et al. 2014) and Brucella melitensis (Vishnu et al. 2017) are the examples of some pathogens that are recently approached using this in silico technique in order to spot some epitopes having potential of being a vaccine candidate. Herpesviridae (Bruno et al. 2015) and hepatitis C virus (HCV) (Kolesanova et al. 2015) are the examples of the viruses that are addressed using this approach.
15.4 Workflow of Reverse Vaccinology (with Example of Coronavirus)
15.4.1 Retrieval of Proteome of Different Strains of Coronavirus from NCBI
The proteome of different strains of the coronavirus of interest was downloaded from NCBI’s ftp site (ftp://ftp.ncbi.nlm.nih.gov/genomes/Viruses/; NCBI Resource Coordinators 2017). The proteome information is available for download in many formats including FASTA format for different sequenced viruses. Strains pathogenic to humans were selected for further analysis. Among them a single strain was selected as the seed genome on the basis of literature. Sequence similarity searches using Blastp (http://blast.ncbi.nlm.nih.gov/blast, http://ugene.unipro.ru/) were performed to reveal the orthologs in different strains (Altschul et al. 1990; Okonechnikov et al. 2012; Golosova et al. 2014). Multiple sequence alignment (MSA) was done via ClustalW, and the phylogenetic tree was constructed using NJ method from Unipro UGENE 1.16.1 bioinformatics toolkit (Okonechnikov et al. 2012).
15.4.2 Analysis of Secondary Structure of Proteins from Seed Genome
Analysis of secondary structure of the proteins of seed genome was done by means of ExPASy portal. The aim is to forecast the solvent accessibility, instability index, theoretical pI, molecular weight, grand average of hydropathicity (GRAVY), aliphatic index, number of charged residues, extinction coefficient etc. (http://web.expasy.org/protparam/; Gasteiger et al. 2005).
15.4.3 Subcellular Localization Predictions and Count of Transmembrane Helices
Virus-mPLoc was used to identify the localization of proteins of virus in the infected cells of host (http://www.csbio.sjtu.edu.cn/bioinf/virus-multi/; Hong-Bin Shen and Kuo-Chin Chou 2010). This information is important to understand the destructive role and mechanism of the viral proteins in causing the disease. In total six different subcellular locations, namely, host cytoplasm, viral capsid, host plasma membrane, host nucleus, host endoplasmic reticulum and secreted proteins, were covered. These predictions could help in formulation of better therapeutic options against the virus. As per the protocol of RV, secreted and membrane proteins are of special interest, therefore, filtered for further analysis. To predict the number of transmembrane helices TMHMM Server v. 2.0 (http://www.cbs.dtu.dk/services/TMHMM/; Krogh et al. 2001) was used.
15.4.4 Signal Peptides
Signal peptides are known to impact the immune responses and possess high epitope densities. Moreover, most of the known vaccine candidates also possess signal peptides. Hence, it is worthwhile to predict signal peptides in proteins prior to epitope predictions. Signal-BLAST web server is used to predict the signal peptides without any false predictions (http://sigpep.services.came.sbg.ac.at/signalblast.html; Frank and Sippl 2008). The prediction options include best sensitivity, balanced prediction, best specificity and detect cleavage site only. We choose to make the predictions using each option, and the proteins predicted as signal peptide by all the four options were preferred for further investigation.
15.4.5 Adhesion Probability
The most appropriate targets as vaccine candidates are those which possess the adhesion-like properties because they not only mediate the adhesion of pathogen’s proteins with cells of host but also facilitate transmission of virus. Adhesions are known to be crucial for virulence and are located on surface which makes them promptly approachable to antibodies. The stand-alone SPAAN with a sensitivity of 89% and specificity of 100% was used to carry out the adhesion probability predictions, and the proteins with having adhesion probabilities higher than or equal to 0.4 were selected (Sachdeva et al. 2004).
15.4.6 BetaWrap Motifs
BetaWrap motifs are dominant in virulence factors of the pathogens. If the proteins are predicted to possess such motifs, then they are appropriate to be taken under reverse vaccinology studies. BetaWrap server is the only online web server to make such predictions. The proteins having P-value lower than 0.1 were anticipated to contain BetaWraps (http://groups.csail.mit.edu/cb/betawrap/betawrap.html; Bradley et al. 2001).
15.4.7 Antigenicity Predictions
For added identification of the antigenic likely of the proteins, they were subjected to VaxiJen server version 2.0. It is basically an empirical method to hunt antigenic proteins. So, if the proteins are not found antigenic using other sequence-based methods, then they can be identified using this method. This step confirms the antigenicity of proteins selected using above-mentioned steps (http://www.ddg-pharmfac.net/vaxijen/VaxiJen/VaxiJen.html; Doytchinova and Flower 2007).
15.4.8 Allergenicity Predictions
For being a probable vaccine candidate, the protein should not exhibit the characteristics of an allergen as they trigger the type-1 hypersensitivity reactions causing allergy. Therefore, to escape out such possibilities, the proteins were also subjected to allergenicity predictions using Allertop (http://www.pharmfac.net/allertop; Dimitrov et al. 2014) and AlgPred tools (http://www.imtech.res.in/raghava/algpred/submission.html; Saha and Raghava 2006a, b).
15.4.9 Similarity with Host Proteins
To check whether the filtered proteins possess any similarity to host proteins or not, the standard Blastp (http://blast.ncbi.nlm.nih.gov/blast) searches were performed. In case of sequence similarity, there is a feasibility of generation of immune responses against own cells.
15.4.10 Epitope Mapping
Predicting the epitopes binding to MHC class I is the main decisive phase of the RV to carry out valid vaccine predictions. The epitopes showing their affinity for T-cells were first selected via IEDB (http://tools.immuneepitope.org/mhci/), ProPred-I (http://www.imtech.res.in/raghava/propred1/; Singh and Raghava 2003), BIMAS (http://www-bimas.cit.nih.gov/molbio/hla_bind/; Parker et al. 1994) and NetCTL tools (http://www.cbs.dtu.dk/services/NetCTL/; Larsen et al. 2005). For the epitope to be included in the hit list, it must be predicted by any three of these four mentioned tools. For making the predictions of B-cell epitopes, BepiPred (http://www.cbs.dtu.dk/services/BepiPred/; Larsen et al. 2006) and ABCPred tools (http://www.imtech.res.in/raghava/abcpred/ABC_submission.html; Saha and Raghava 2006a, b) were used. The overlapping B-cell and T-cell epitopes were identified.
15.4.11 Docking of the Predicted Epitopes with HLA-A*0201
The predicted epitopes were docked with receptor that is HLA-A*0201 using ClusPro (http://cluspro.bu.edu/login.php; Kozakov et al. 2017) that is an automated protein-protein docking web server. The literature searches provided the information of conserved residues of the receptor site. The default parameters were used for docking (Comeau et al. 2004a, b; Kozakov et al. 2006).
15.5 Results and Discussion
15.5.1 Retrieval of Proteome from NCBI
A total of 40 different sequenced strains of coronavirus are available at NCBI. Among them 7 strains are pathogenic to humans. Various information regarding source, host and collection of these strains are presented in Table 15.1 and 15.2. This information can be obtained from NCBI’s genome database, the Virus Pathogen Database and Analysis Resource and Genomes OnLine Database (Liolios et al. 2006; Pickett et al. 2012). The MERS strain is taken as seed genome as it is the most prevalent and disastrous strain among others. Its proteome consists of total 11 proteins as shown in Table 15.3. The results of sequence similarity to reveal orthologs using Blastp are shown in Table 15.4. The sequences with greater than 30% identity score are considered as homologs. The phylogenetic tree is depicted in Fig. 15.1 and the MERS-CoV, taken as seed genome, found clustered with different Bat coronaviruses.
15.5.2 Analysis of Secondary Structure
The results of analysis of secondary structure of the proteome using ExPASy tools are shown in the Table 15.5. From the analysis of charge on the residues and pH values, it is concluded that six of the proteins are basic and positively charged unlike allergens which are acidic in nature. However, five proteins are acidic and show negative charge. The negative GRAVY score of five proteins justify them to be of hydrophilic nature with majority of the residues positioned towards the surface. For the rest of six proteins, the GRAVY score is positive; it means that these are hydrophobic proteins. The proteins with less than 40 value of instability index are quite stable than those with higher values. All the proteins are having the molecular weight less than 110 kDa except 3 (YP_009047202.1, YP_009047203.1 and YP_009047204.1). This exhibits the effectiveness of lightweight proteins as targets as they can be easily purified because of their low molecular weights. The protein YP_009047204.1 is reported as a spike glycoprotein. It is acidic with prominent negative charge, with negative GRAVY score which suggests its hydrophilicity and presence on surface. However the envelope protein YP_009047209.1 and membrane protein YP_009047210.1 are basic and hydrophobic.
15.5.3 Subcellular Localization Predictions
Figure 15.2 depicts the subcellular localization of proteins of the seed genome, i.e. MERS-CoV. Only one protein was predicted to be localized in host cytoplasm, four in host membrane, two in both host cell membrane and endoplasmic reticulum (ER) while two in only ER, and two are left unrecognized. The known spike protein is predicted to be localized in host ER. From these results we decided to pick the proteins which are located in host membrane or were predicted to be localized in both host membrane and ER. The two are known envelop protein and membrane protein from bibliographic studies, and along with that, the known spike protein was also included in the filtered results. Out of the filtered proteins, only two (YP_009047210.1 and YP_009047208.1) contain more than two transmembrane helices, therefore filtered out. The results of transmembrane helices prediction are tabulated in Table 15.6. Figure 15.3 depicts the subcellular localization of proteins of all the four selected genomes using Virus-mPLoc prediction tool.
15.5.4 Signal Peptides
The proteins that are predicted to possess the signal peptides by Signal-BLAST web server are YP_009047204.1 and YP_009047205.1. The results of Signal-BLAST web server are tabulated in the Table 15.7.
15.5.5 Adhesion Probability
This step takes into account the concept of adhesion-based virulence. Adhesions cause pathogen recognition and initiation of inflammatory responses by the host. SPAAN predicted 2 (YP_009047204.1 and YP_009047205.1) out of 11 proteins of MERS strain as adhesive (Table 15.8).
15.5.6 BetaWrap
Only one protein (YP_009047204.1) was predicted to contain BetaWrap motifs within it (Table 15.8). Hence, it is considered virulent and might be responsible for initializing the infection in the host.
15.5.7 VaxiJen 2.0
A total of 9 out of 11 proteins of MERS strain were predicted antigenic (prediction values greater than 0.4). The protein with accession number YP_009047206.1 and YP_009047208.1 were among the filtered proteins, however, not predicted antigenic, therefore filtered out. As a result, only four proteins (YP_009047204.1, YP_009047205.1, YP_009047207.1 and YP_009047209.1) were kept for further analyses.
15.5.8 AlgPred and Allertop
None of the 11 proteins of MERS-CoV possessed any clue of allergenicity as per prediction results from AlgPred and Allertop tools; it means that no vigorous immune responses will be mounted if the epitopes from these proteins will be adopted as vaccine candidates.
15.5.9 Similarity with Host Proteome
None of the protein of MERS strain shows similarity with the proteins of host that demonstrates that the epitopes from these proteins can safely elicit the required immune response without the hazard of autoimmunity.
15.5.10 Epitope Mapping
In total 12 different 9-mer epitopes with potential to bind to receptors of both B-cell and T-cell were predicted. The list of the predicted epitopes can be found in the Table 15.9 and are specific for MERS-CoV strain. All these epitopes displayed no conservancy with proteins of other human and non-human pathogenic strains.
15.5.11 Docking Analysis
Docking permits to reveal the binding energy or potency of connection among epitopes and the receptor in appropriate orientation. The ClusPro docking server was used to dock the predicted 90 epitopes against HLA-A*0201. The structure of the receptor was available from PDB and was optimized before docking to free it from the complexed self-peptide (4U6Y, Resolution 1.47 Å, Bouvier et al. 1998). PEPstr (Peptide Tertiary Structure Prediction Server; Kaur et al. 2007) was used to derive the tertiary structure of the predicted peptides.
Figure 15.4 depicts the quaternary structure of the receptor HLA-A*0201 with its conserved active site known to form complex with the peptides (Bouvier et al. 1998). The binding energy results obtained after performing docking analysis are listed in Table 15.9.
The 9-mer epitope VVCAITLLV at site 21 of protein YP_009047209.1 docked to the receptor with smallest amount of binding energy (−951.7) and 12 hydrogen bonds. The next epitope in the list was also from the same protein YP_009047209.1 at site 27, i.e. TLLVCMAFL. The predicted structure of the top 5 potent epitopes on the basis of docking energy and the snapshots of docking results are displayed in Figs. 15.5, 15.6, 15.7, 15.8 and 15.9.
(a) 3D Structure of the 9-mer epitope starting from 21(VVCAITLLV) position of protein YP_009047209.1 (b) Docking results of epitope “VVCAITLLV” with A chain of HLA-A*0201 using ClusPro. (c) The snapshot representing the epitope docked in the pocket of molecular surface of the receptor (all the structures are visualized using Chimera 1.10.1)
(a) 3D Structure of the 9-mer epitope starting from 27(TLLVCMAFL) position of protein YP_009047209.1. (b) Docking results of epitope “TLLVCMAFL” with A chain of HLA-A*0201 using ClusPro. (c) The snapshot representing the epitope docked in the pocket of molecular surface of the receptor (all the structures are visualized using Chimera 1.10.1)
(a) 3D Structure of the 9-mer epitope starting from 716(GLVNSSLFV) position of protein YP_009047204.1. (b) Docking results of epitope “GLVNSSLFV” with A chain of HLA-A*0201 using ClusPro. (c) The snapshot representing the epitope docked in the pocket of molecular surface of the receptor (all the structures are visualized using Chimera 1.10.1)
(a) 3D Structure of the 9-mer epitope starting from 18(YVDVGPDSV) position of protein YP_009047204.1. (b) Docking results of epitope “YVDVGPDSV” with A chain of HLA-A*0201 using ClusPro. (c) The snapshot representing the epitope docked in the pocket of molecular surface of the receptor (all the structures are visualized using Chimera 1.10.1)
(a) 3D Structure of the 9-mer epitope starting from 160(KMGRFFNHT) position of protein YP_009047204.1. (b) Docking results of epitope “KMGRFFNHT” with A chain of HLA-A*0201 using ClusPro. (c) The snapshot representing the epitope docked in the pocket of molecular surface of the receptor (all the structures are visualized using Chimera 1.10.1)
The most chief restriction for developing a safe and sound vaccine against any of the virus is to identify the protective antigens. The present study is an effort of application of reverse vaccinology approach to investigate a choice of coronavirus proteomes to identify possible vaccine targets. This technique has demonstrated to be a competent way to forecast 12 different epitopes from the selected seed genome. These epitopes are from spike glycoprotein, NS3 protein, NS4B protein and envelope protein. Unfortunately none of the epitope is found conserved in other strains, and all are specific to MERS-CoV. The docking analysis studies revealed perfect binding between HLA-A*0201 receptor and epitopes. The conserved residues of the receptor site are also involved in H-bonding with epitope residues. Further, the selected antigenic epitopes must be validated using in vitro and in vivo studies to confirm their potential as vaccine candidates.
References
Alessandro S, Rino R (2010) Review: reverse vaccinology: developing vaccines in the era of genomics. Immunity 33:530–541. https://doi.org/10.1016/j.immuni.2010.09.017
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Azhar EI, Lanini S, Ippolito G, Zumla A (2017) The Middle East respiratory syndrome coronavirus – a continuing risk to global health security. Adv Exp Med Biol 972:49–60. https://doi.org/10.1007/5584_2016_133.PMID:27966107
Bouvier M, Guo HC, Smith KJ, Wiley DC (1998) Crystal structures of HLA-A*0201 complexed with antigenic peptides with either the amino- or carboxyl-terminal group substituted by a methyl group. Proteins Struct Funct Genet 33(1):97–106. PMID: 9741848
Bradley P, Cowen L, Menke M, King J, Berger B (2001) BETAWRAP: successful prediction of parallel beta -helices from primary sequence reveals an association with many microbial pathogens. Proc Natl Acad Sci U S A 98(26):14819–14824. https://doi.org/10.1073/pnas.251267298
Brown JK, Fauquet CM, Briddon RW, Zerbini M, Moriones E, Navas-Castillo J, King AM, Adams MJ, Carstens EB, Lefkowitz EJ (2012) Geminiviridae. In: Virus taxonomy—ninth report of the International Committee on Taxonomy of Viruses. Elsevier Science, Burlington, pp 351–373
Bruno L, Cortese M, Rappuoli R, Merola M (2015) Lessons from Reverse Vaccinology for viral vaccine design. Curr Opin Virol 11:89–97. https://doi.org/10.1016/j.coviro.2015.03.001
Caro-Gomez E, Gazi M, Goez Y, Valbuena G (2014) Discovery of novel cross-protective Rickettsia prowazekii T-cell antigens using a combined reverse vaccinology and in vivo screening approach. Vaccine 32(39):4968–4976. https://doi.org/10.1016/j.vaccine.2014.06.089
Chiang MH, Sung WC, Lien SP, Chen YZ, Lo AF, Huang JH, Kuo SC, Chong P (2015) Identification of novel vaccine candidates against Acinetobacter baumannii using reverse vaccinology. Hum Vaccin Immunother 11(4):1065–1073. https://doi.org/10.1080/21645515.2015.1010910
Comeau SR, Gatchell DW, Vajda S, Camacho CJ (2004a) ClusPro: a fully automated algorithm for protein-protein docking. Nucleic Acids Res 32(suppl_2):W96–W99. https://doi.org/10.1093/nar/gkh354
Comeau SR, Gatchell DW, Vajda S, Camacho CJ (2004b) ClusPro: an automated docking and discrimination method for the prediction of protein complexes. Bioinformatics 20(1):45–50. https://doi.org/10.1093/bioinformatics/btg371
De Groot RJ, Baker SC, Baric RS et al (2013) Middle East respiratory syndrome coronavirus (MERS-CoV): announcement of the coronavirus study group. J Virol 87(14):7790–7792. https://doi.org/10.1128/JVI.01244-13
Dimitrov I, Bangov I, Flower DR, Doytchinova I (2014) AllerTOP v.2-a server for in silico prediction of allergens. J Mol Model BioMed Central Ltd 20(6):227. DOI: https://doi.org/10.1007/s00894-014-2278-5
Doytchinova IA, Flower DR (2007) VaxiJen: a server for prediction of protective antigens, tumour antigens and subunit vaccines. BMC Bioinf 8(1):4. https://doi.org/10.1186/1471-2105-8-4
Frank K, Sippl MJ (2008) High performance signal peptide prediction based on sequence alignment techniques. Bioinformatics 24:2172–2176. https://doi.org/10.1093/bioinformatics/btn422
Gabutti G (2014) Meningococcus B: control of two outbreaks by vaccination. J Prev Med Hyg 55(2):35–41. PMCID: PMC4718325
Gasteiger E, Hoogland C, Gattiker A, Duvaud S, Wilkins MR, Appel RD et al (2005) Protein identification and analysis tools on the ExPASy server. Proteomics Protoc Handb:571–607. https://doi.org/10.1385/1-59259-890-0:571
Golosova O, Henderson R, Vaskin Y, Gabrielian A, Grekhov G, Nagarajan V, Oler AJ, Quiñones M, Hurt D, Fursov M, Huyen Y (2014) Unipro UGENE NGS pipelines and components for variant calling, RNA-seq and ChIP-seq data analyses. Peer J 2:e644. https://doi.org/10.7717/peerj.644
Goodswen SJ, Kennedy PJ, Ellis JT (2014) Discovering a vaccine against neosporosis using computers: is it feasible? Trends Parasitol 30(8):401–411. https://doi.org/10.1016/j.pt.2014.06.004
Guimarães L, Soares S, Trost E, Blom J, Ramos R, Silva A, Barh D, Azevedo V (2015) Genome informatics and vaccine targets in Corynebacterium urealyticum using two whole genomes, comparative genomics, and reverse vaccinology. BMC Genomics 16(Suppl 5):S7. https://doi.org/10.1186/1471-2164-16-S5-S7
Kanampalliwar AM, Soni R, Tiwari A, Giridhar A (2013) Reverse vaccinology: basics and applications. J Vaccin Vaccin 4(6):194–198. https://doi.org/10.4172/2157-7560.1000194
Kaur H, Garg A, Raghava GPS (2007) PEPstr: a de novo method for tertiary structure prediction of small bioactive peptides. Protein Pept Lett 14:626–630. https://doi.org/10.2174/092986607781483859
Kelly DF, Rappuoli R (2005) Reverse vaccinology and vaccines for serogroup B Neisseria meningitidis. Adv Exp Med Biol 568:217–223
Kolesanova EF, Sobolev BN, Moysa AA, Egorova EA, Archakov AI (2015) Way to the peptide vaccine against hepatitis C. Biomed Khim 61(2):254–264. https://doi.org/10.18097/PBMC20156102254
Kozakov D, Brenke R, Comeau SR, Vajda S (2006) PIPER: an FFT-based protein docking program with pair wise potentials. Proteins 65(2):392–406. https://doi.org/10.1002/prot.21117
Kozakov D, Hall DR, Xia B, Porter KA, Padhorny D, Yueh C, Beglov D, Vajda S (2017) The ClusPro web server for protein-protein docking. Nat Protoc 12(2):255–278
Krogh A, Larsson B, von Heijne G, Sonnhammer EL (2001) Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J Mol Biol 305(3):567–580. https://doi.org/10.1006/jmbi.2000.4315
Larsen MV, Lundegaard C, Lamberth K, Buus S, Brunak S, Lund O, Nielsen M (2005) An integrative approach to CTL epitope prediction, a combined algorithm integrating MHC-I binding, TAP transport efficiency, and proteasomal cleavage prediction. J Immunol 35:2295–2303. https://doi.org/10.1002/eji.200425811
Larsen JEP, Lund O, Nielsen M (2006) Improved method for predicting linear B-cell epitopes. Immunome Res 2(1):2. https://doi.org/10.1186/1745-7580-2-2
Liolios K, Tavernarakis N, Hugenholtz P, Kyrpides NC (2006) The genomes on line database (GOLD) v.2: a monitor of genome projects worldwide. Nucleic Acids Res 34:D332–D334. https://doi.org/10.1093/nar/gkj145
Monterrubio-López GP, González-Y-Merchand JA, Ribas-Aparicio RM (2015) Identification of novel potential vaccine candidates against tuberculosis based on reverse vaccinology. Biomed Res Int 2015:483150. https://doi.org/10.1155/2015/483150
Naz A, Awan FM, Obaid A, Muhammad SA, Paracha RZ, Ahmad J, Ali A (2015) Identification of putative vaccine candidates against Helicobacter pylori exploiting exoproteome and secretome: a reverse vaccinology based approach. Infect Genet Evol 32:280–291. https://doi.org/10.1016/j.meegid.2015.03.027
NCBI Resource Coordinators (2017) Database resources of the national center for biotechnology information. Nucleic Acids Res 45(D1):D12–D17. https://doi.org/10.1093/nar/gkw1071
Okonechnikov K, Golosova O, Fursov M, UGENE team (2012) Unipro UGENE: a unified bioinformatics toolkit. Bioinformatics 28:1166–1167. https://doi.org/10.1093/bioinformatics/bts091
Parker KC, Bednarek MA, Coligan JE (1994) Scheme for ranking potential HLA-A2 binding peptides based on independent binding of individual peptide side-chains. J Immunol 152(1):163–75. PMID: 8254189
Pickett BE, Greer DS, Zhang Y, Stewart L, Zhou L, Sun G et al (2012) Virus pathogen database and analysis resource (ViPR): a comprehensive bioinformatics database and analysis resource for the coronavirus research community. Viruses 11:3209–3226. https://doi.org/10.3390/v4113209
Pizza M, Scarlato V, Masignani V et al (2000) Identification of vaccine candidates against serogroup B meningococcus by whole-genome sequencing. Science 287:1816–1820. https://doi.org/10.1126/science.287.5459.1816
Rappuoli R (2000) Reverse vaccinology. Curr Opin Microbiol 3:445–450. https://doi.org/10.1016/S1369-5274(00)00119-3
Sachdeva G, Kumar K, Preti J, Ramachandran S (2004) SPAAN: a software program for prediction of adhesins and adhesin-like proteins using neural networks. Bioinformatics 21:483–491. https://doi.org/10.1093/bioinformatics/bti028
Saha S, Raghava GPS (2006a) AlgPred: prediction of allergenic proteins and mapping of IgE epitopes. Nucleic Acids Res 34:W202–W209. https://doi.org/10.1093/nar/gkl343
Saha S, Raghava GPS (2006b) Prediction of continuous B-cell epitopes in an antigen using recurrent neural network. Proteins 65(1):40–48. https://doi.org/10.1002/prot.21078
Sette A, Rappuoli R (2010) Reverse vaccinology: developing vaccines in the era of genomics. Immunity 33(4):530–541. https://doi.org/10.1016/j.immuni.2010.09.017
Shen H-B, Chou K-C (2010) Virus-mPLoc: a fusion classifier for viral protein subcellular location prediction by incorporating multiple sites. J Biomol Struct Dyn 28:175–186. https://doi.org/10.1080/07391102.2010.10507351
Singh H, Raghava GPS (2003) ProPred1: prediction of promiscuous MHC class-I binding sites. Bioinformatics 19(8):1009–1014. https://doi.org/10.1093/bioinformatics/btg108
Vishnu US, Sankarasubramanian J, Gunasekaran P, Rajendhran J (2017) Identification of potential antigens from non-classically secreted proteins and designing novel multitope peptide vaccine candidate against Brucella melitensis through reverse vaccinology and immunoinformatics approach. Infect Genet Evol 55:151–158
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Dangi, M., Kumari, R., Singh, B., Chhillar, A.K. (2018). Advanced In Silico Tools for Designing of Antigenic Epitope as Potential Vaccine Candidates Against Coronavirus. In: Shanker, A. (eds) Bioinformatics: Sequences, Structures, Phylogeny . Springer, Singapore. https://doi.org/10.1007/978-981-13-1562-6_15
Download citation
DOI: https://doi.org/10.1007/978-981-13-1562-6_15
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-13-1561-9
Online ISBN: 978-981-13-1562-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)