Abstract
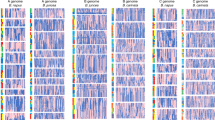
All Brassica species are derived from a common hexaploid ancestor, and this hexaploid ancestor has been further deduced to origin from a diploid species through a whole genome triplication event. The diploid ancestor has 7 chromosomes and resembles the karyotype of tPCK (translocation Proto-Calepineae Karyotype). The confirming evidences for the Brassicas’ tPCK ancestor are from below three aspects: (1) The reconstructed genomic segments of the three subgenomes of all Brassica species keep the genomic structure of tPCK; (2) The locations of extant centromeres and the traces of paleocentromeres on the genomes of Brassicas support its ancestral diploid genome as tPCK; (3) The phylogeny tree and evolution analysis based on the whole genome sequences of several sequenced Brassicaceae species find that the Brassicas are evolved from a tPCK genome, such as the tPCK species S. parvula. The determination of the shared diploid ancestor for all Brassica species lays an important foundation for the genetic studies of Brassica crops.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Burrell AM, Taylor KG, Williams RJ, Cantrell RT, Menz MA, Pepper AE (2011) A comparative genomic map for Caulanthus amplexicaulis and related species (Brassicaceae). Mol Ecol 20:784–798
Cheng F, Liu S, Wu J, Fang L, Sun S et al (2011) BRAD, the genetics and genomics database for Brassica plants. BMC Plant Biol 11:136
Cheng F, Wu J, Fang L, Wang X (2012a) Syntenic gene analysis between Brassica rapa and other Brassicaceae species. Front Plant Sci 3:198
Cheng F, Wu J, Fang L, Sun S, Liu B et al (2012b) Biased gene fractionation and dominant gene expression among the subgenomes of Brassica rapa. PLoS One 7:e36442
Cheng F, Mandáková T, Wu J, Xie Q, Lysak MA, Wang X (2013) Deciphering the diploid ancestral genome of the Mesohexaploid Brassica rapa. Plant Cell 25:1541–1554
Cheng F, Wu J, Wang X (2014) Genome triplication drove the diversification of Brassica plants. Hortic Res 1:14024
Dassanayake M, Oh DH, Haas JS, Hernandez A, Hong H et al (2011) The genome of the extremophile crucifer Thellungiella parvula. Nat Genet 43:913–918
Franzke A, Lysak MA, Al-Shehbaz IA, Koch MA, Mummenhoff K (2010) Cabbage family affairs: the evolutionary history of Brassicaceae. Trends Plant Sci 16:108–116
Hu TT, Pattyn P, Bakker EG, Cao J, Cheng JF et al (2011) The Arabidopsis lyrata genome sequence and the basis of rapid genome size change. Nat Genet 43:476–481
Initiative AG (2000) Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408:796–815
Koch MA, Haubold B, Mitchell-Olds T (2000) Comparative evolutionary analysis of chalcone synthase and alcohol dehydrogenase loci in Arabidopsis, Arabis, and related genera (Brassicaceae). Mol Biol Evol 17:1483–1498
Lagercrantz U, Lydiate DJ (1996) Comparative genome mapping in Brassica. Genetics 144:1903–1910
Li F, Hasegawa Y, Saito M, Shirasawa S, Fukushima A et al (2011) Extensive chromosome homoeology among Brassiceae species were revealed by comparative genetic mapping with high-density EST-based SNP markers in radish (Raphanus sativus L.). DNA Res 18:401–411
Liu S, Liu Y, Yang X, Tong C, Edwards D et al (2014) The Brassica oleracea genome reveals the asymmetrical evolution of polyploid genomes. Nat Commun (in press)
Lysak MA, Koch MA, Pecinka A, Schubert I (2005) Chromosome triplication found across the tribe Brassiceae. Genome Res 15:516–525
Lysak MA, Cheung K, Kitschke M, Bures P (2007) Ancestral chromosomal blocks are triplicated in Brassiceae species with varying chromosome number and genome size. Plant Physiol 145:402–410
Mandáková T, Lysak MA (2008) Chromosomal phylogeny and karyotype evolution in x = 7 crucifer species (Brassicaceae). Plant Cell 20:2559–2570
Nagaharu U (1935) Genome analysis in Brassica with special reference to the experimental formation of B. napus and peculiar mode of fertilication. Jpn J Bot 7:389–452
Nelson MN, Parkin IA, Lydiate DJ (2011) The mosaic of ancestral karyotype blocks in the Sinapis alba L. genome. Genome 54:33–41
Panjabi P, Jagannath A, Bisht NC, Padmaja KL, Sharma S et al (2008) Comparative mapping of Brassica juncea and Arabidopsis thaliana using Intron Polymorphism (IP) markers: homoeologous relationships, diversification and evolution of the A, B and C Brassica genomes. BMC Genom 9:113
Parkin IA, Gulden SM, Sharpe AG, Lukens L, Trick M et al (2005) Segmental structure of the Brassica napus genome based on comparative analysis with Arabidopsis thaliana. Genetics 171:765–781
Prakash S, Hinata K (1980) Taxonomy, cytogenetics and origin of crop Brassicas, a review. Oper Bot 55:1–57
Röbbelen G (1960) Beitra ge zur Analyse des Brassica-Genoms. Chromosoma 11:205–228
Schmutz J, Cannon SB, Schlueter J, Ma J, Mitros T et al (2010) Genome sequence of the palaeopolyploid soybean. Nature 463:178–183
Schnable PS, Ware D, Fulton RS, Stein JC, Wei F et al (2009) The B73 maize genome: complexity, diversity, and dynamics. Science 326:1112–1115
Schranz ME, Lysak MA, Mitchell-Olds T (2006) The ABC’s of comparative genomics in the Brassicaceae: building blocks of crucifer genomes. Trends Plant Sci 11:535–542
Sharma S, Padmaja KL, Gupta V, Paritosh K, Pradhan AK et al (2014) Two plastid DNA lineages–Rapa/Oleracea and Nigra–within the tribe Brassiceae can be best explained by reciprocal crosses at hexaploidy: evidence from divergence times of the plastid genomes and R-block genes of the A and B genomes of Brassica juncea. PLoS One 9:e93260
Shirasawa K, Oyama M, Hirakawa H, Sato S, Tabata S et al (2011) An EST-SSR linkage map of Raphanus sativus and comparative genomics of the Brassicaceae. DNA Res 18:221–232
Truco MJ, Hu J, Sadowski J, Quiros CF (1996) Inter- and infra-genomic homology of the Brassica genomes: implications for their origin and evolution. Theor Appl Genet 93:1225–1233
Wang X, Wang H, Wang J, Sun R, Wu J et al (2011a) The genome of the mesopolyploid crop species Brassica rapa. Nat Genet 43:1035–1039
Wang X, Wang H, Wang J, Sun R, Wu J et al (2011b) The genome of the mesopolyploid crop species Brassica rapa. Nat Genet 43:1035–1039
Wu HJ, Zhang Z, Wang JY, Oh DH, Dassanayake M et al (2012) Insights into salt tolerance from the genome of Thellungiella salsuginea. Proc Natl Acad Sci USA 109:12219–12224
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Cheng, F., Lysak, M.A., Mandáková, T., Wang, X. (2015). The Common Ancestral Genome of the Brassica Species. In: Wang, X., Kole, C. (eds) The Brassica rapa Genome. Compendium of Plant Genomes. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-662-47901-8_8
Download citation
DOI: https://doi.org/10.1007/978-3-662-47901-8_8
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-662-47900-1
Online ISBN: 978-3-662-47901-8
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)