Abstract
The mouse is a perfect model to study aging in mammals. It has a relatively short life span and genetic manipulations in this species are well established. Most interestingly, the mouse is a fantastic tool to produce stem cells. Forced expression of only four transcription factors (Oct3/4, Sox2, Klf4, and c-Myc) in murine and human somatic cells resets the expression of genes that are characteristic of differentiated cells and consequently induces the formation of pluripotent stem cells (iPSCs). This technology opens new and exciting possibilities in medical research, especially personalized cell therapies for treating human disease. To treat damaged tissues or repair organs in elderly patients, it will be necessary to establish iPSCs from their tissues. To determine the feasibility of using this technology with elderly patients, we asked whether it was indeed possible to establish iPSCs from the tissues of aged mice and to differentiate them to tissue cells. We succeeded in establishing iPSC clones using bone marrow (BM) from 21-month-old EGFP-C57BL/6 mice, which had been cultured for 4 days in the presence of granulocyte macrophage-colony stimulating factor (GM-CSF). Our iPSCs from aged mice (aged iPSCs) and those from mouse embryonic fibroblasts (MEFs) strongly expressed SSEA-1 and Pou5f1, and showed strong alkaline phosphatase (AP) activity. Our aged iPSCs made teratomas when injected into the back skin of syngeneic mice, and differentiated to tissue cells of three germ lines in vitro. Further experiments to make chimeric mice and germ line cells will determine whether the aged iPSCs possess the properties of much younger cells and are capable of regenerating aged mice.
Similar content being viewed by others
Keywords
- Pluripotent Stem Cell
- Leukemia Inhibitory Factor
- Aged Mouse
- Somatic Cell Nuclear Transfer
- Inner Cell Mass
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
21.1 Introduction
The mean life expectancy in developed countries has increased dramatically (Oeppen and Vaupel 2002). For instance, the mean life expectancy in Japan has increased from 50 to 80 years in no more than 6 decades. Socioeconomic factors, including the improvement of environmental conditions and medical care, may explain part of this increase in life span, but ample evidence suggests that genetic factors are also at play. Studies of twins and long-lived families have estimated that 20–30% of the variation in human life span is determined by genetic factors (Herskind et al. 1996; Mitchell et al. 2001; Hjelmborg et al. 2006).
The mouse is an ideal mammalian model for studying aging. It shares 99% of its genes with humans (Boguski 2002), has a short generation time, and its environment can be easily controlled (Yuan et al. 2011). Furthermore, the genetic resources for the mouse include hundreds of inbred strains and mutants and sophisticated genetic engineering technology for manipulating its genome (Paigen 1995; Peters et al. 2007).
Somatic cell nuclear transfer (SCNT) into oocytes and the cloning of entirely new animals from a single cell revolutionized the concepts of developmental biology. SCNT was first developed in Xenopus (Gurdon 1962), followed by “Dolly” the sheep (Wilmut et al. 1997) and then mice (Wakayama et al. 1998). This technique suggests that cellular factors affect chromatin structure and gene expression. The forced expression of only four transcription factors (Oct3/4, Sox2, Klf4, and c-Myc) in murine and human somatic cells can reset the genes of differentiated cells and induce pluripotent stem cells (iPSCs) (Takahashi and Yamanaka 2006; Takahashi et al. 2007; Wernig et al. 2007; Jaenisch and Young 2008; Yu, et al. 2007; Park et al. 2008; Lowry et al. 2008) (Fig. 21.1a). This technology opens exciting possibilities in medical research, especially personalized cell therapies for treating human diseases.
Methods to make pluripotent cells. A. Adults somatic cells are reprogrammed by Somatic cell nuclear transfer (SCNT) into oocytes. The forced expression of transcription factors in somatic cells can reset the genes of differentiated cells and induce pluripotent stem cells (iPSCs). B. Pluripotent stem cells from fetal and adult stem cells
Because of the human leukocyte antigen (HLA) barrier, almost all organ transplantation therapies require immunosuppressive drugs which cause several side effects. If it were possible to generate iPSCs from a patient’s own healthy cells and induce differentiation, we could avoid allogeneic organ transplantation. In developed countries, a large number of elderly people suffer from untreatable diseases. To treat damaged tissues or repair organs in elderly patients, it will be necessary to establish iPSCs from their tissues. To determine the feasibility of the application of this technology to elderly patients, we studied the possibilities to establish iPSCs from the tissues of aged mice and to redifferentiate them to a range of tissues. In the first section of this chapter, we will briefly summarize the molecular characteristics of iPSCs and then discuss the possibility of using this technique to rejuvenate aged damaged tissues.
21.2 Characteristics of Induced Pluripotent Stem Cells
Pluripotent stem cells exist in developing embryos and have been established in tissue culture. The first pluripotent cells were derived from a germ line tumor teratocarcinoma (Hogan 1976). Then, embryonic stem cells (ESCs) from the inner cell mass (ICM) of normal mouse embryos were developed, which made a great impact on biomedical research by permitting the creation of knockout mice (Evans and Kaufman 1981). Primordial germ cells isolated from embryonic day 8.5 embryos generated ES-like cells, termed embryonic germ cells (Surani 1999). From newborn or adult gonads, ES-like cells were generated and called spermatogonial stem cells, which are propagated in vitro in the presence of serum and leukemia inhibitory factor (LIF) (Kanatsu-Shinohara et al. 2004) (Fig. 21.1b). The pluripotent stem cells most commonly used in research are ESCs and iPSCs because they grow well in vitro. Many characteristics of ESCs and iPSCs are similar. ESCs or iPSCs are maintained on feeder fibroblasts in the presence of fetal bovine serum and LIF which support STAT3 (signal tranducer and activator of transcription 3) signaling. Alternatively, feeder layers can be omitted by the inclusion of LIF, BMP-4 and other small molecules that stimulate Wnt signaling (Ying et al. 2008).
Chromatin—chromosomal DNA as packaged with histones—provides the cellular context for gene expression. On the basis of histological evidence, ESCs and iPSCs exhibit euchromatin (lightly stained and sparsely packed chromatin), which is mostly devoid of more densely stained, compacted areas of the heterochromatin. The chromatin of ESCs or iPSCs is in an “open” state (Gaspar-Maia et al. 2011). Cells in the ICM of the mouse blastocyst at day 3.5, which are the source of ES cells, share the same open chromatin conformation as ES cells. Upon differentiation, heterochromatin appears heterogeneous and clustered in distinct blocks. DNA of ESCs and iPSCs is hypomethylated. Especially, demethylation of Oct4 and Nanog is widely used to monitor successful reprogramming (Mikkelsen et al. 2008). Histone-modifying enzymes that repress genes encoding lineage-specific developmental regulators have the most profound impact on ESC or iPSC states. Histone H3 tri-methylation and di-methylation on Lys9 (H3K9me3 and H3K9me2) are indicators of heterochromatin. They are low in iPSCs and ESCs (Hawkins et al. 2010), but upon differentiation, H3K9 is hypermethylated. In contrast, active markers such as H3K4me3 and acetylation of histones H3 and H4 are upregulated in ESCs and iPSCs (Fig. 21.2). The pluripotency regulatory network is also directly linked to the Jumonji (Jmj) family H3K9 demethylases Jmjd1a/KDM2A and Jmjd2c/KDM4B, which act on H3K9me2 and H3K9me3, respectively (Loh et al. 2007). Both genes lie downstream of Oct4 and are regulated positively through its action. Recent discoveries have generated substantial excitement, as they show that cytosines in mammalian cells can be hydroxymethylated to 5hmC (5-hydroxymethylcytosine), which is especially abundant in tissues such as ESCs and iPSCs. 5-Methylcytosine (5mC) can be hydroxylated by the ten-eleven translocation (TET) family of enzymes. Koh and colleagues (2011) reported that TET levels are high in pluripotent cells and decline during differentiation. 5hmC was especially enriched at the start sites of genes whose promoters bear dual histone 3 lysine 27 trimethylation (H3K27me3) and histone 3 lysine 4 trimethylation (H3K4me3) marks (Williams et al. 2011; Wu et al. 2011). Tet1 is required for establishing a defined genomic pattern of 5hmC, and also for initiating an enzymatic cascade to maintain CpG-rich gene promoters in a DNA hypo-methylated state (Wu et al. 2011).
21.3 Is It Possible to Generate iPSCs from Aged Mice?
It has been shown that cellular senescence induced by reprogramming impairs successful reprogramming to iPSCs. Banito and collaborators (2009) showed that expression of four reprogramming factors mentioned above triggers senescence via upregulating p53, p16 (INK4a), and p21(CIP1). Upregulation of these senescence-related proteins blocks proliferation of target fibroblasts and reduces the generation of iPSCs. Genetic inhibition of the Ink4a/Arf locus has a profound positive effect on the efficiency of iPS cell generation, increasing both the kinetics of reprogramming and the number of emerging iPS cell colonies. In murine cells, Arf, rather than Ink4a, is the main barrier to reprogramming by activation of p53 (encoded by Trp53) and p21 (encoded by Cdkn1a), whereas, in human fibroblasts, INK4a is more important in reprogramming than ARF (Li et al. 2009). p53 is a stress-response protein, which suppresses tumor formation by promoting senescence (permanent cell-cycle arrest). Several papers have shown that inactivation of p53 markedly increases the efficiency of iPSC production (Hong et al. 2009; Kawamura et al. 2009; Utikal et al. 2009; Marión et al. 2009). These results indicate that during the course of induction of iPSCs, cellular aging occurs and reduces the capacity to generate iPSCs.
It was reported that fibroblasts from the lung of a human fetus had a fixed, replicative life span of about 50 population doublings before replication stopped (Hayflick 1965). This finding was reinforced by the report that the replicative potential of fibroblasts cultured from skin decreased with the increasing age of human donors (Martin et al. 1970). Additional support for the putative relationship between donor age and replicative life span of fibroblasts came from the report that the number of possible divisions in vitro was directly related to the average in vivo lifetime of the species (Röhme 1981). Later, the limited in vitro lifetime was attributed to telomere shortening (Harley et al. 1990), which also contributed to organismal aging by limiting the proliferative capacity of adult stem cells (Blasco 2007; Flores et al. 2006). These results indicated that the efficiency of iPSCs production might be lower in aged cells than in young cells.
We investigated the establishment of iPSCs from aged mice (Cheng et al. 2011). We used BM-derived myeloid (BM-M) cells that were actively proliferating in the presence of the granulocyte macrophage-colony stimulating factor (GM-CSF) (Fig. 21.3). First, we calculated the efficiency of iPSC generation from aged mice. We compared MEF cells and BM-M cells obtained from either 2-month-old C57BL/6 or 23-month-old C57BL/6 mice. Colonies appeared approximately 15 days after virus transduction from both MEF and 2-month (2M)-old BM-M cells, although the number of colonies produced by MEF cells increased more rapidly than those from 2M-old BM-M cells (10 colonies from MEF versus 2 colonies from 2M-old BM-M cells at 15 days). As expected, 23-month-old BM-M cells produced fewer colonies and these appeared later (10 colonies from 2M-old BM-M cells versus 2 colonies from 23M-old BM-M cells 30 days after transduction; Fig. 21.4). These results indicate that the efficiency of establishing iPSCs from aged mice is indeed lower than that from young mice (Cheng et al. 2011). Then we tried to verify whether iPSC cell lines from aged mice could grow and differentiate as well as iPSCs from young mice. To this end, we planned to transplant differentiated iPSCs to damaged tissues of syngeneic C57BL/6 mice. In order to discriminate transplanted iPSCs from recipient syngeneic mice, we selected GFP-positive C57BL/6 mice. We succeeded in establishing iPSCs clones using BM of 21-month-old EGFP-C57BL/6 mice, which had been cultured for 4 days in the presence of GM-CSF. We selected two clones (1 and 2) and expanded them (Fig. 21.3).
21.4 Are Aged Somatic Cells Rejuvenated By iPSC Production?
There is an overall decline in tissue regenerative potential with age (Rando 2006). For instance, wound healing becomes slower with age in skin. This age-related decline of wound healing is mainly caused by the decrease of regenerative potential of stem cells in the skin, although immune cells affect these phenomena (Nishio et al. 2008). During aging, the somatic cells accumulate DNA lesions, caused by a variety of stimuli, although the host has defense systems. These include single- and double-strand DNA breaks, chromosomal translocations and single base mutations. Telomere shortening occurs as a consequence of somatic cell division (Wang et al. 2009). Most of these DNA mutations that accumulate during aging are irreversible, except telomere shortening (described later). Unlike acquired DNA mutations, epigenomic changes (DNA methylation and histone methylations, acetylation, and ubiquitination) are potentially reversible. Loci in CpG islands gained methylation with age, while loci outside of CpG islands lost methylation with age (Christensen et al. 2009). Maegawa and collaborators (2010) demonstrated a surprisingly high rate of hyper- and hypomethylation as a function of age in normal mouse small intestine tissues and a strong tissue specificity to the process. During aging, histone methylation also occurs and may affect gene expression and function. It has been shown that regulators of histone H3K4 trimethylation complex associate during the life span in C. elegans (Greer et al. 2010). In mammals, histone methylation affects the aging of tissue cells. In young murine oocytes, dimethylation of lysines 4, 9, 36 and 79 in histone 3 (H3K4me2, H3K9me2, H3K36me2, H3K79me2) and methylation of lysine 20 in histone H4 (H4K20me2) and trimethylation of lysine 9 in histone 3 (H3K9me3) were observed. However, in old murine oocytes, H3K4me2 and H3K9me2 increased and demethylation of H3K9me3, H3K36me2, H3K79me2, and H4K20me2 occurred (Manosalva and González 2010) (Fig. 21.5).
Prior to the discovery of iPSCs, several approaches were taken to determine whether aged somatic cells could be rejuvenated and whether differentiation was reversible. Early cell fusion experiments between somatic cells and ESCs demonstrated the reversibility of differentiation (Blau et al. 1985). It was shown that the life span of mice cloned from somatic cells (immature Sertoli cells) was significantly shorter than that of genotype- and sex-matched controls, most likely due to severe pneumonia and hepatic failure (Ogonuki et al. 2002). Immune responses (antibody production and phagocytosis) in cloned mice were lower than in controls. However, they reported that two of the 12 cloned mice in their experiment appeared to have normal life spans.
An important question is whether telomeres of aged iPSCs become shorter than those of young iPSCs. Because telomerase activity is upregulated in both human and mouse iPSCs (Takahashi et al. 2007; Stadtfeld et al. 2008), it was suggested that telomeres might be elongated in iPSCs compared to parental cells. Consistent with this idea, telomeres were found to be elongated in iPSCs taken from an elderly person (Dimos et al. 2008).
If aged iPSCs were rejuvenated, their gene expression profile and epigenetic state might be similar to those of iPSCs from MEFs. Reprogramming of somatic cells to pluripotency is accompanied by extensive remodeling of epigenetic marks, including DNA demethylation of key pluripotency genes such as Oct4 and Nanog. In somatic cells, the promoters of Oct4 and Nanog are highly methylated, reflecting their transcriptionally repressed state. The formation of iPSCs involves activation of these genes, and their demethylation is widely used to monitor successful reprogramming (Mikkelsen et al. 2008). In principle, demethylation could occur by a passive mechanism, such as the inhibition of DNA methyltransferase 1 (Dnmt1) during DNA replication, or by an active mechanism in which the methylated base is removed from nonreplicating DNA.
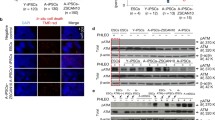
Our iPSCs from aged mice and MEF iPSCs strongly expressed SSEA-1 and Oct-4 (Fig. 21.6a) and Pou5f1, and they showed strong alkaline phosphatase (AP) activity. Gene expression of Nanog and Pou5f1 in aged iPSCs was as high as in MEF iPSCs and the promoters of Nanog and Pou5f1 were hypomethylated in both aged iPSCs and MEF iPSCs by ChIP assays using anti-H3-K27 antibodies (Cheng et al. 2011). However, our examination of gene expression and the epigenetic state of aged iPSCs is preliminary. Whole genome sequencing and extensive Chip assays will likely reveal differences between aged iPSCs and MEF iPSCs and show unique characteristic features of age-derived iPSCs. Recently, it was shown that the reprogramming process and subsequent culture of iPSCs in vitro can induce genetic and epigenetic abnormalities in these cells (Hussein et al. 2011; Gore et al. 2011; Lister et al. 2011; Mayshar et al. 2010; Laurent et al. 2011). It has been shown that copy number variation and point mutations frequently occur during iPSC generation.
21.5 Do iPSCs from Aged Mice Differentiate Normally?
We examined whether iPSCs from aged mice (aged iPSCs) demonstrate pluripotency similar to iPSCs from MEF (MEF iPSCs). We transplanted aged iPSCs (1 × 107) or MEF iPSCs to the dorsal flank of syngeneic C57BL/6 mice. Teratomas appeared after the injection of aged iPSCs and MEF iPSCs. After 21 days, we observed distinct tumors that were collected and fixed in OCT (Optimal Cutting Temperature) Compound (Sakura Finetechnical Co., Ltd., Japan) for frozen 5-μm sections.
We observed various tissues belonging to three germ layers (endoderm, mesoderm, and ectoderm) (Fig. 21.6b). They were positively stained by antibodies against alpha-smooth muscle actin (mesoderm), α-fetoprotein (endoderm) and neurofilament H (ectoderm). We detected GFP-positive cells from the transplantation of aged iPSCs. Our observation indicates that iPSCs can differentiate to tissue cells and make tissues in vivo (Cheng et al. 2011).
However, another group showed that iPSCs made by a retroviral approach failed to form detectable teratomas or formed teratomas that were subsequently immune rejected by T-cell infiltration and massive necrosis when they were transplanted to syngeneic mice. Further, they showed that iPSCs generated by an episomal approach were also rejected by CD4 T cells in syngeneic mice (Zhao et al. 2011). The differences between our results and their results could be explained if aged iPSCs were not immunogenic. However, when we injected MEF-derived iPSCs to syngeneic mice, we observed teratoma formation in syngeneic mice. Another possible explanation can be attributed to the choice of the injection site. We injected iPSCs subcutaneously into the dorsal flank of syngeneic C57BL/6 mice. They were also injected subcutaneously into the hind leg region of C57BL/6 mice. They demonstrated that several genes were upregulated in differentiated teratoma, which were rejected by CD4 T cells. The immunogenicity of iPSCs presents major problems for personalized cell therapy and must be investigated in greater detail using our aged iPSCs.
Next, we attempted to differentiate aged iPSCs to three germ cell lines in vitro. We cultured aged iPSCs by the hanging drop method for 8 days to induce embryoid bodies. They were then transferred to 24-well trays. When these cultures were stained with tissue-specific antibodies, we detected tissue cells belonging to all the three germ layers (Fig. 21.7) (Cheng et al. 2011). Currently, we are attempting to differentiate aged iPSCs in vitro to various tissues.
21.6 Future Directions
In order to use iPSCs for regenerative medicine, especially iPSCs from elderly patients, the cells must be differentiated properly. We would like to use the murine model to determine whether aged iPSCs can be developed in vivo as are those from young mice. Namely, can iPS technology reverse aging and produce young mice, even if the original cells are derived from aged mice? We have set up chimera experiments (Fig. 21.8), which will ultimately induce newborns fully made by iPSCs (germ line transmission). When we performed chimeric experiments using 8-cell aggregation methods, we obtained some chimeric mice (Fig. 21.8), although we have not yet achieved germ line transmission.
Similar questions about aging have been raised by SCNT technology (Kishigami et al 2008). Using in vitro senescent cow cells, the birth of six healthy cloned calves was reported. Nuclear transfer extended the replicative life span of senescent cells (zero to four population doublings remaining) to greater than 90 population doublings (Lanza et al. 2000). Thus, SCNT can restore at least cellular senescence. The first cloned mammal, Dolly, was found to have short telomeres that were comparable to the age of the cell donor (Shiels et al. 1999). However, later reports showed that murine clones grown over five generations did not have shorter telomeres (Wakayama et al. 2000). The first cloned mouse, Cumulina, died after 2 years and 7 months, which is slightly longer than the average life span of a mouse (Tamashiro et al. 2003). Although SCNT into oocytes and the cloning of animals from a single cell showed the possibility of the successful reprogramming of a somatic nucleus (Gurdon and Melton 2008), there are no reports of the application of SCNT using mouse cells that originated from old mice (e.g. 2 years old). Recently, Wakayama’s group succeeded in establishing nuclear transfer embryonic stem (ntES) cell lines from aged mice with an establishment rate of 10–25%, irrespective of sex or strain (Mizutani et al. 2008). Our current trial to produce germ line transmission from aged iPSCs (21-month-old C57BL/6 mice) and examine their life span will hopefully answer fundamental questions of aging and determine the usefulness of iPSCs for future regenerative medicine.
References
Banito A, Rashid ST, Acosta JC, Li S, Pereira CF, Geti I et al (2009) Senescence impairs successful reprogramming to pluripotent stem cells. Genes Dev 23(18):2134–2139
Blasco MA (2007) The epigenetic regulation of mammalian telomeres. Nat Rev Genet 8(4):299–309
Blau HM, Pavlath GK, Hardeman EC, Chiu CP, Silberstein L, Webster SG et al (1985) Plasticity of the differentiated state. Science 230(4727):758–766
Boguski MS (2002) Comparative genomics: the mouse that roared. Nature 420(6915):515–516
Cheng Z, Ito S, Nishio N, Xiao H, Zhang R, Suzuki H et al (2011) Establishment of induced pluripotent stem cells from aged mice using bone marrow-derived myeloid cells. J Mol Cell Biol 3(2):91–98
Christensen BC, Houseman EA, Marsit CJ, Zheng S, Wrensch MR, Wiemels JL et al (2009) Aging and environmental exposures alter tissue-specific DNA methylation dependent upon CpG island context. PLoS Genet 5(8):e1000602
Dimos JT, Rodolfa KT, Niakan KK, Weisenthal LM, Mitsumoto H, Chung W et al (2008) Induced pluripotent stem cells generated from patients with ALS can be differentiated into motor neurons. Science 321(5893):1218–1221
Evans MJ, Kaufman MH (1981) Establishment in culture of pluripotential cells from mouse embryos. Nature 292(5819):154–156
Flores I, Benetti R, Blasco MA (2006) Telomerase regulation and stem cell behaviour. Curr Opin Cell Biol 18(3):254–260
Gaspar-Maia A, Alajem A, Meshorer E, Ramalho-Santos M (2011) Open chromatin in pluripotency and reprogramming. Nat Rev Mol Cell Biol 12(1):36–47
Gore A, Li Z, Fung HL, Young JE, Agarwal S, Antosiewicz-Bourget J et al (2011) Somatic coding mutations in human induced pluripotent stem cells. Nature 471(7336):63–67
Greer EL, Maures TJ, Hauswirth AG, Green EM, Leeman DS, Maro GS et al (2010) Members of the H3K4 trimethylation complex regulate lifespan in a germline-dependent manner in C. elegans. Nature 466(7304):383–387
Gurdon JB (1962) The transplantation of nuclei between two species of Xenopus. Dev Biol 5:68–83
Gurdon JB, Melton DA (2008) Nuclear reprogramming in cells. Science 322(5909):1811–1815
Harley CB, Futcher AB, Greider CW (1990) Telomeres shorten during ageing of human fibroblasts. Nature 345(6274):458–460
Hawkins RD, Hon GC, Lee LK, Ngo Q, Lister R, Pelizzola M et al (2010) Distinct epigenomic landscapes of pluripotent and lineage-committed human cells. Cell Stem Cell 6(5):479–491
Hayflick L (1965) The limited in vitro lifetime of human diploid cell strains. Exp Cell Res 37:614–636
Herskind AM, McGue M, Holm NV, Sørensen TI, Harvald B, Vaupel JW (1996) The heritability of human longevity: a population-based study of 2872 Danish twin pairs born 1870–1900. Hum Genet 97(3):319–323
Hogan BL (1976) Changes in the behaviour of teratocarcinoma cells cultivated in vitro. Nature 263(5573):136–137
Hong H, Takahashi K, Ichisaka T, Aoi T, Kanagawa O, Nakagawa M et al (2009) Suppression of induced pluripotent stem cell generation by the p53–p21 pathway. Nature 460(7259):1132–1135
Hussein SM, Batada NN, Vuoristo S, Ching RW, Autio R, Närvä E et al (2011) Copy number variation and selection during reprogramming to pluripotency. Nature 471(7336):58–62
Jaenisch R, Young R (2008) Stem cells, the molecular circuitry of pluripotency and nuclear reprogramming. Cell 132(4):567–582
Kanatsu-Shinohara M, Inoue K, Lee J, Yoshimoto M, Ogonuki N, Miki H et al (2004) Generation of pluripotent stem cells from neonatal mouse testis. Cell 119(7):1001–1012
Kawamura T, Suzuki J, Wang YV, Menendez S, Morera LB, Raya A et al (2009) Linking the p53 tumour suppressor pathway to somatic cell reprogramming. Nature 460(7259):1140–1144
Kishigami S, Wakayama S, Hosoi Y, Iritani A, Wakayama T (2008) Somatic cell nuclear transfer: infinite reproduction of a unique diploid genome. Exp Cell Res 314(9):1945–1950
Koh KP, Yabuuchi A, Rao S, Huang Y, Cunniff K, Nardone J et al (2011) Tet1 and Tet2 regulate 5-hydroxymethylcytosine production and cell lineage specification in mouse embryonic stem cells. Cell Stem Cell 8(2):200–213
Lanza RP, Cibelli JB, Blackwell C, Cristofalo VJ, Francis MK, Baerlocher GM et al (2000) Extension of cell life-span and telomere length in animals cloned from senescent somatic cells. Science 288(5466):665–669
Laurent LC, Ulitsky I, Slavin I, Tran H, Schork A, Morey R et al (2011) Dynamic changes in the copy number of pluripotency and cell proliferation genes in human ESCs and iPSCs during reprogramming and time in culture. Cell Stem Cell 8(1):106–118
Li H, Collado M, Villasante A, Strati K, Ortega S, Cañamero M et al (2009) The Ink4/Arf locus is a barrier for iPS cell reprogramming. Nature 460(7259):1136–1139
Lister R, Pelizzola M, Kida YS, Hawkins RD, Nery JR, Hon G et al (2011) Hotspots of aberrant epigenomic reprogramming in human induced pluripotent stem cells. Nature 471(7336):68–73
Loh YH, Zhang W, Chen X, George J, Ng HH (2007) Jmjd1a and Jmjd2c histone H3 Lys 9 demethylases regulate self-renewal in embryonic stem cells. Genes Dev 21(20):2545–2557
Lowry WE, Richter L, Yachechko R, Pyle AD, Tchieu J, Sridharan R et al (2008) Generation of human induced pluripotent stem cells from dermal fibroblasts. Proc Natl Acad Sci USA 105(8):2883–2888
Maegawa S, Hinkal G, Kim HS, Shen L, Zhang L, Zhang J et al (2010) Widespread and tissue specific age-related DNA methylation changes in mice. Genome Res 20(3):332–340
Manosalva I, González A (2010) Aging changes the chromatin configuration and histone methylation of mouse oocytes at germinal vesicle stage. Theriogenology 74(9):1539–1547
Marión RM, Strati K, Li H, Murga M, Blanco R, Ortega S et al (2009) A p53-mediated DNA damage response limits reprogramming to ensure iPS cell genomic integrity. Nature 460(7259):1149–1153
Martin GM, Sprague CA, Epstein CJ (1970) Replicative life-span of cultivated human cells. Effects of donor’s age, tissue, and genotype. Lab Invest 23(1):86–92
Mayshar Y, Ben-David U, Lavon N, Biancotti JC, Yakir B, Clark AT et al (2010) Identification and classification of chromosomal aberrations in human induced pluripotent stem cells. Cell Stem Cell 7(4):521–531
Mikkelsen TS, Hanna J, Zhang X, Ku M, Wernig M, Schorderet P et al (2008) Dissecting direct reprogramming through integrative genomic analysis. Nature 454(7200):49–55
Mitchell BD, Hsueh WC, King TM, Pollin TI, Sorkin J, Agarwala R et al (2001) Heritability of life span in the Old Order Amish. Am J Med Genet 102(4):346–352
Mizutani E, Ono T, Li C, Maki-Suetsugu R, Wakayama T (2008) Propagation of senescent mice using nuclear transfer embryonic stem cell lines. Genesis 46(9):478–483
Nishio N, Okawa Y, Sakurai H, Isobe K (2008) Neutrophil depletion delays wound repair in aged mice. Age (Dordr) 30(1):11–19
Oeppen J, Vaupel JW (2002) Demography. Broken limits to life expectancy. Science 296(5570):1029–1031
Ogonuki N, Inoue K, Yamamoto Y, Noguchi Y, Tanemura K, Suzuki O et al (2002) Early death of mice cloned from somatic cells. Nat Genet 30(3):253–254
Paigen K. (1995) A miracle enough the power of mice. Nat Med 1:215–220
Park IH, Zhao R, West JA, Yabuuchi A, Huo H, Ince TA et al (2008) Reprogramming of human somatic cells to pluripotency with defined factors. Nature 451(7175):141–146
Peters LL, Robledo RF, Bult CJ, Churchill GA, Paigen BJ, Svenson KL (2007) The mouse as a model for human biology: a resource guide for complex trait analysis. Nat Rev Genet 8(1):58–69
Rando TA (2006) Stem cells, ageing and the quest for immortality. Nature 441(7097):1080–1086
Röhme D (1981) Evidence for a relationship between longevity of mammalian species and life spans of normal fibroblasts in vitro and erythrocytes in vivo. Proc Natl Acad Sci USA 78(8):5009–5013
Shiels PG, Kind AJ, Campbell KH, Waddington D, Wilmut I, Colman A et al (1999) Analysis of telomere lengths in cloned sheep. Nature 399(6734):316–317
Stadtfeld M, Maherali N, Breault DT, Hochedlinger K (2008) Defining molecular cornerstones during fibroblast to iPS cell reprogramming in mouse. Cell Stem Cell 2(3):230–240
Surani MA (1999) Reprogramming a somatic nucleus by trans-modification activity in germ cells. Semin Cell Dev Biol 10(3):273–277
Takahashi K, Tanabe K, Ohnuki M, Narita M, Ichisaka T, Tomoda K et al (2007) Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131(5):861–872
Takahashi K, Yamanaka S (2006) Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126(4):663–676
Tamashiro KL, Wakayama T, Yamazaki Y, Akutsu H, Woods SC, Kondo S et al (2003) Phenotype of cloned mice: development, behavior, and physiology. Exp Biol Med (Maywood) 228(10):1193–1200
Utikal J, Polo JM, Stadtfeld M, Maherali N, Kulalert W, Walsh RM et al (2009) Immortalization eliminates a roadblock during cellular reprogramming into iPS cells. Nature 460(7259):1145–1148
vB Hjelmborg J, Iachine I, Skytthe A, Vaupel JW, McGue M, Koskenvuo M et al (2006) Genetic influence on human lifespan and longevity. Hum Genet 119(3):312–321
Wakayama T, Perry AC, Zuccotti M, Johnson KR, Yanagimachi R (1998) Full-term development of mice from enucleated oocytes injected with cumulus cell nuclei. Nature 394(6691):369–374
Wakayama T, Shinkai Y, Tamashiro KL, Niida H, Blanchard DC, Blanchard RJ et al (2000) Cloning of mice to six generations. Nature 407(6802):318–319
Wang C, Jurk D, Maddick M, Nelson G, Martin-Ruiz C, von Zglinicki T (2009) DNA damage response and cellular senescence in tissues of aging mice. Aging Cell 8(3):311–323
Wernig M, Meissner A, Foreman R, Brambrink T, Ku M, Hochedlinger K et al (2007) In vitro reprogramming of fibroblasts into a pluripotent ES-cell-like state. Nature 448(7151):318–324
Williams K, Christensen J, Pedersen MT, Johansen JV, Cloos PA, Rappsilber J et al (2011) TET1 and hydroxymethylcytosine in transcription and DNA methylation fidelity. Nature 473(7347):343–348
Wilmut I, Schnieke AE, McWhir J, Kind AJ, Campbell KH (1997) Viable offspring derived from fetal and adult mammalian cells. Nature 385(6619):810–813
Wu H, D'Alessio AC, Ito S, Wang Z, Cui K, Zhao K et al (2011) Genome-wide analysis of 5-hydroxymethylcytosine distribution reveals its dual function in transcriptional regulation in mouse embryonic stem cells. Genes Dev 25(7):679–684
Ying QL, Wray J, Nichols J, Batlle-Morera L, Doble B, Woodgett J et al (2008) The ground state of embryonic stem cell self-renewal. Nature 453(7194):519–523
Yu J, Vodyanik MA, Smuga-Otto K, Antosiewicz-Bourget J, Frane JL, Tian S et al (2007) Induced pluripotent stem cell lines derived from human somatic cells. Science 318(5858):1917–1920
Yuan R, Peters LL, Paigen B (2011) Mice as a mammalian model for research on the genetics of aging. ILAR J 52(1):4–15
Zhao T, Zhang ZN, Rong Z, Xu Y (2011) Immunogenicity of induced pluripotent stem cells. Nature 474(7350):212–215
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2012 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Isobe, Ki., Cheng, Z., Ito, S., Nishio, N. (2012). Aging in the Mouse and Perspectives of Rejuvenation Through Induced Pluripotent Stem Cells (iPSCs). In: Kubiak, J. (eds) Mouse Development. Results and Problems in Cell Differentiation, vol 55. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-30406-4_21
Download citation
DOI: https://doi.org/10.1007/978-3-642-30406-4_21
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-30405-7
Online ISBN: 978-3-642-30406-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)