Abstract
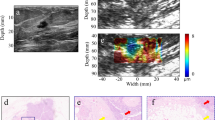
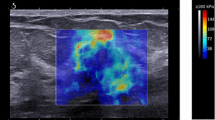
Breast cancer is the most common cancer among women and the second highest cause of cancer-related death. Diagnostic magnetic resonance imaging (MRI) is recommended to screen high-risk patients. Strain-Encoded (SENC) can improve MRI’s specificity by detecting and differentiating masses according to their stiffness. Previous phantom and ex-vivo studies have utilized SENC to detect cancerous masses. However, SENC required a 30% compression of the tissue, which may not be feasible for in-vivo imaging. In this work, we use finite element method simulations and phantom experiments to determine the minimum compression required to detect and classify masses. Results show that SENC is capable of detecting stiff masses at compression level of 7%, though higher compression is needed in order to differentiate between normal tissue and benign or malignant masses. With on-line SENC calculations implemented on the scanner console, we propose to start with small compressions for maximum patient comfort, then progress to larger compressions if any masses are detected.
Chapter PDF
Similar content being viewed by others
Keywords
- Dynamic Mechanical Analyzer
- Breast Magnetic Resonance Imaging
- Finite Element Method Simulation
- Phantom Experiment
- Malignant Mass
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
References
American Cancer society, Cancer Facts and Figures 2009 (2009), http://www.cancer.org
Bluemke, D.A., Gatsonis, C., Chen, M.H., DeAngelis, G.A., DeBruhl, N., Harms, S., Heywang-Kobrunner, S.H., Hylton, N., Kuhl, C.K., Lehman, C., et al.: Magnetic resonance imaging of the breast prior to biopsy. Jama 292(22), 2735 (2004)
Samani, A., Zubovits, J., Plewes, D.: Elastic moduli of normal and pathological human breast tissues: an inversion-technique-based investigation of 169 samples. Physics in Medicine and Biology 52(6), 1565–1576 (2007)
Osman, N., Sampath, S., Atalar, E., Prince, J.: Imaging longitudinal cardiac strain on short-axis images using strain-encoded MRI. Magnetic Resonance in Medicine 46(2), 324–334 (2001)
Osman, N.: Detecting stiff masses using strain-encoded (SENC) imaging. Magnetic Resonance in Medicine 49(3), 606–608 (2003)
Fahmy, A., Krieger, A., Osman, N.: An integrated system for real-time detection of stiff masses with a single compression. IEEE Transactions on Biomedical Engineering 53(7), 1286–1293 (2006)
Harouni, A.A., Hossain, J., Jacobs, M.A., Osman, N.F.: Improved Hardware for Higher Spatial Resolution Strain-Encoded (SENC) MRI. Academic Radiology 18, 705–715 (2011)
Harouni, A.A., El Khouli, R.H., Hossain, J., Bluemke, D.A., Osman, N.F., Jacobs, M.A.: Enhancing mass detection and classification in breast tissue using Strain-ENCoded (SENC) breast MRI with histological validation. Submitted to Journal of Magnetic Resonance Imaging (March 2011)
Yousef, T.A., Osman, N.F.: Effect of Noise and Slice Profile on Strain Quantifications of Strain Encoding (SENC) MRI. In: Sachse, F.B., Seemann, G. (eds.) FIHM 2007. LNCS, vol. 4466, pp. 50–59. Springer, Heidelberg (2007)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2011 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Harouni, A.A., Jacobs, M.A., Osman, N.F. (2011). Finding the Optimal Compression Level for Strain-Encoded (SENC) Breast MRI; Simulations and Phantom Experiments. In: Fichtinger, G., Martel, A., Peters, T. (eds) Medical Image Computing and Computer-Assisted Intervention – MICCAI 2011. MICCAI 2011. Lecture Notes in Computer Science, vol 6891. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-23623-5_56
Download citation
DOI: https://doi.org/10.1007/978-3-642-23623-5_56
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-23622-8
Online ISBN: 978-3-642-23623-5
eBook Packages: Computer ScienceComputer Science (R0)