Abstract
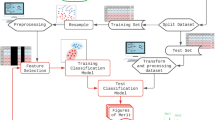
A new approach for the automatic detection of drug-induced organ toxicities based on Nuclear Magnetic Resonance Spectroscopy data from biofluids is presented in this paper. Spectral data from biofluids contain information on the concentration of various substances, but the combination of only a small subset of these cues is putatively useful for classification of new samples. We propose to divide the spectra into several short regions and train classifiers on them, using only a limited amount of information for class discrimination. These local experts are combined in an ensemble classification system and the subset of experts for the final classification is optimized automatically. Thus, only local experts for relevant spectral regions are used for the final ensemble classification. The proposed approach has been evaluated on a real data-set from industrial pharmacology, showing an improvement in classification accuracy and indicating relevant spectral regions for classification.
Parts of this work have been funded by a grant from Boehringer Ingelheim Pharma GmbH & Co. KG., Genomics group. The authors would like to thank the General Pharmacology Group of the company for providing the sample set.
Chapter PDF
Similar content being viewed by others
Keywords
- Nuclear Magnetic Resonance
- Nuclear Magnetic Resonance Spectrum
- Near Neighbor
- Local Expert
- Safety Pharmacology
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
References
Freeman, R.: Magnetic resonance in chemistry and medicine. Oxford University Press, New York (2003)
Lindon, J.C., et al.: Contemporary issues in toxicology the role of metabonomics in toxicology and its evaluation by the COMET project. Toxicology and Applied Pharmacology 187(3), 137–146 (2003)
Holmes, E., et al.: Development of a model for classification of toxin-induced lesions using 1H NMR spectroscopy of urine combined with pattern recognition. NMR in Biomedicine 11(4-5), 235–244 (1998)
Beckonert, O., et al.: NMR-based metabonomic toxicity classification: hierarchical cluster analysis and k-nearest-neighbour approaches. Analytica Chimica Acta 490, 3–15 (2003)
Fieno, T., Viswanathan, V., Tsoukalas, L.: Neural network methodology for 1H NMR spectroscopy classification. In: ICIIS 1999: Proc. Int. Conf. on Information Intelligence and Systems, pp. 80–85. IEEE Computer Society, Los Alamitos (1999)
Kittler, J., Hatef, M., Duin, R.P., Matas, J.: On combining classifiers. IEEE Transactions on Pattern Analysis and Machine Intelligence 20(3), 226–239 (1998)
Lienemann, K., Plötz, T., Fink, G.A.: On the application of SVM-Ensembles based on adapted random subspace sampling for automatic classification of NMR data. In: Haindl, M., Kittler, J., Roli, F. (eds.) MCS 2007. LNCS, vol. 4472, pp. 42–51. Springer, Heidelberg (2007)
Schölkopf, B., Smola, A.J.: Learning with Kernels. MIT Press, Cambridge (2002)
Ho, T.K.: The random subspace method for constructing decision forests. IEEE Transactions on Pattern Analysis and Machine Intelligence 20(8), 832–844 (1998)
Lienemann, K., Plötz, T., Pestel, S.: NMR-based urine analysis in rats: Prediction of proximal tubule kidney toxicity and phospholipidosis. Journal of Pharmacological and Toxicological Methods 58(1), 41–49 (2008)
Spraul, M., et al.: Automatic reduction of NMR spectroscopic data for statistical and pattern recognition classification of samples. Journal of Pharmaceutical & Biomedical Analysis 12, 1215–1225 (1994)
Torgrip, R.J.O., et al.: New methods of data partitioning based on pars peak alignment for improvedmultivariate biomarker/biopattern detection in 1H NMR spectroscopic metabolic profiling of urine. Metabolomics 2(1), 1–19 (2006)
Torgrip, R.J.O., Åberg, M., Karlberg, B., Jacobsson, S.P.: Peak alignment using reduced set mapping. Journal of Chemometrics 17, 573–582 (2003)
Skov, T., van den Berg, F., Tomasi, G., Bro, R.: Automated alignment of chromatographic data. Journal of Chemometrics 20(11-12), 484–497 (2006)
Kuncheva, L., Bezdek, J., Duin, R.: Decision templates for multiple classifier fusion: an experimental comparison. Pattern Recognition 34, 299–314 (2001)
Goldberg, D.E.: Genetic Algorithms in Search, Optimization and Machine Learning. Addison-Wesley Longman Publishing Co. (1989)
Matthews, B.W.: Comparison of the predicted and observed secondary structure of the T4 phage lysozyme. Biochimica et Biophysica Acta 405, 442–451 (1975)
Duda, R.O., Hart, P.E., Stork, D.G.: Pattern Classification, 2nd edn. John Wiley & Sons, Chichester (2001)
Breiman, L.: Random forests. Machine Learning 45(1), 5–32 (2001)
Barnes, R.J., et al.: Standard normal variate transformation and de-trending of near-infrared diffuse reflectance spectra. Applied Spectroscopy 43(5), 772–777 (1989)
Nord, L.I., Kenne, L., Jacobsson, S.: Multivariate analysis of 1H NMR spectra for saponins from quillaja saponaria molina. Anal. Chim. Acta 446, 197–207 (2001)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2008 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Lienemann, K., Plötz, T., Fink, G.A. (2008). Automatic Classification of NMR Spectra by Ensembles of Local Experts. In: da Vitoria Lobo, N., et al. Structural, Syntactic, and Statistical Pattern Recognition. SSPR /SPR 2008. Lecture Notes in Computer Science, vol 5342. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-540-89689-0_83
Download citation
DOI: https://doi.org/10.1007/978-3-540-89689-0_83
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-540-89688-3
Online ISBN: 978-3-540-89689-0
eBook Packages: Computer ScienceComputer Science (R0)