Abstract
The last few years have seen an explosion of experimental and computational methods for investigating RNA structures of entire transcriptomes in vivo. Very recent experimental protocols now also allow trans RNA–RNA interactions to be probed in a transcriptome-wide manner. All of the experimental strategies require comprehensive computational pipelines for analysing the raw data and converting it back into actual RNA structure features or trans RNA–RNA interactions. The overall performance of these methods thus strongly depends on the experimental and the computational protocols employed. In order to get the best out of both worlds, both aspects need to be optimised simultaneously. This review introduced the methods and proposes ideas how they could be further improved.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
- RNA secondary structures
- trans RNA–RNA interactions
- RNA structure prediction
- RNA–RNA interaction prediction
- Transcriptomes
- In vivo RNA structure probing
- In vivo probing of trans RNA–RNA interaction
- RNA structurome
- RNA interactome
1 Introduction
The remarkable chemical properties of RNA allow transcripts in vivo to directly interact with themselves (via so-called RNA structure) or in trans with other transcripts, DNA and proteins. Many known RNA functions are expressed in terms of RNA structure. Substantial insight into the potential functional roles of any RNA can already be gained by studying its so-called RNA secondary structure, i.e. the set of base-paired sequence positions that form base pairs via hydrogen bonds (the consensus base pairs are {G, C}, {G, U} and {A, U}). Obviously, the functional roles of any RNA can be encoded not only via RNA structure features, but also via sequence signals such as the sequence of codons defining a contiguous open-reading frame at messenger-RNA (mRNA) level or the sequence of nucleotides defining a protein-binding site. As it turns out, many ways of encoding functional information into a transcript are mutually compatible. For example, any given transcript may have RNA structure while simultaneously interacting with other molecules such as other transcripts, DNA or proteins. Or, one and the same stretch of RNA may encode a functional RNA structure as well as codon information on protein synthesis. Cases like these are not only found in viral genomes where space constraints force different layers of information to overlap (Pedersen et al. 2004b; Watts et al. 2009) but can also occur in otherwise perfectly ordinary coding transcripts of model organisms such as human, mouse and fruit fly. Luckily, overlapping layers of information can be detected in silico provided dedicated computational methods are employed that are capable of explicitly dis-entangling them (Pedersen et al. 2004a,b; Meyer and Miklos 2005). It is already known that RNA structure features can act as exquisite sensors of the complex in vivo environment and change according to sometimes subtle changes of intrinsic and extrinsic factors. Examples of these factors range from single-nucleotide modifications of the primary RNA transcript (e.g. tRNAs and rRNAs require a range of well-defined chemical modifications at distinct sequence positions in order to become functionally active in vivo) and other changes of the primary transcript sequence (cleavage, splicing, tail-adding, A-to-I RNA editing, etc.) to changes of the surrounding temperature, changing trans interacting partners (ligands, other RNAs, proteins, DNA) and changes of the transcription speed. A wealth of recent evidence supports the notion of alternative RNA structure expression (Meyer 2017), i.e. that a single transcript can encode and express not just one, but several distinct RNA structures which are differentially expressed depending on the specific in vivo environment. Known cases do include examples not only from bacteria, but also from model organisms such as the fruit fly (Steif and Meyer 2012; Zhu et al. 2013; Zhu and Meyer 2015; Mazloomian and Meyer 2015). There is, for example, strong statistical evidence for differentially expressed, local RNA structure features near splice sites that define tissue-specific splice isoforms (Mazloomian and Meyer 2015). The corresponding RNA structure changes are mediated by tissue-specific A-to-I RNA editing of these structural features (Mazloomian and Meyer 2015). Alternative RNA structure expression allows one and the same (coding or non-coding) transcript to wear a series of distinct functional hats throughout its cellular life depending on its directly surrounding in vivo environment (extrinsic factors) and any modifications it undergoes itself (intrinsic changes). Taken together, the transcriptome thus offers exceptional potential to functionally link all layers of the central dogma of biology in a well-regulated manner that depends on the specific in vivo environment, thereby influencing gene expression and determining the organism’s overall complexity. For better or worse, the days where we may silently assume the validity of the one-sequence-one-structure dogma are over. This has far-reaching implications on how we should experimentally probe RNA structures and trans RNA–RNA interactions in vivo and how we should model these features computationally.
Whereas protein–protein, DNA–protein and protein–RNA interactions have been the subject of intense experimental and computational research for a while, transcriptome-wide investigations of RNA structures and general methods for detecting trans RNA–RNA interactions in vivo have only emerged fairly recently. On the experimental side, one major step forward was made very recently (2016) via the publication of three experimental protocols that can directly probe both RNA structure features and trans RNA–RNA interactions in a transcriptome-wide fashion in vivo. On the computational side, ab initio methods for predicting truly novel trans RNA–RNA interactions based on primary sequence data are only just emerging (Lai and Meyer 2016). Even these most recent experimental methods rely heavily on the computational analysis of their raw data to infer any actual RNA structures or trans RNA–RNA interactions. Any biological insight gained from experimental in vivo studies is thus a complex function of the combined experimental and computational strategies employed. The purpose of this review is therefore to describe, highlight and discuss key features of these experimental and computational pipelines that contribute critically to the overall results. The focus here is thus almost exclusively on method development. We therefore refer the reader to the respective original papers and recent reviews, e.g. (Bevilacqua et al. 2016), regarding the biological insights gained.
2 Transcriptome-Wide Experimental Methods for Determining RNA Structures In Vivo in a Nucleotide-Specific Way
In vivo, RNAs are surrounded by aqueous solution. Any experimental investigation of RNA secondary structures and trans RNA–RNA interactions (RNA structures and RNA–RNA interactions in the following) with potential relevance to biological in vivo systems thus has to happen in solution (Ehresmann et al. 1987).
2.1 Brief Survey of Experimental In Vitro Methods
2.1.1 Physical Methods
Early experimental methods for RNA structure probing comprise physical methods such as X-ray crystallography and nuclear magnetic resonance spectroscopy (NMR) (Lengyel et al. 2014). Both methods take the RNA out of its cellular context, especially so X-ray crystallography, where the ability to crystallise implies the almost complete removal of the solvent. Even then, not all RNAs crystallise equally well (some not at all), so that database of RNA structures derived by X-ray crystallography has inherent biases. NMR imposes a considerable limitation on the length of the RNAs that can be investigated. Both in vitro methods are low-throughput in the sense that they typically investigate a single RNA at a time. Especially NMR requires considerable human expertise to design and interpret all experiments required to determine a RNA structure. Experiments for different RNAs are considered on a case by case basis. These general limitations none-withstanding, NMR and X-ray crystallography have generated a wealth of important insights on RNA structure properties in vitro. Discrepancies between the RNA structures derived from NMR and from X-ray crystallography experiments give an early indication that RNA structure features are fairly context-sensitive (Higgs 2000). Based on these early observations, differences between RNA structures in vitro and in vivo could thus be expected.
2.1.2 Enzymatic Methods
RNA structure features can also be probed using RNases. These ribonucleases correspond to naturally occurring proteins that cleave at specific paired (i.e. double-stranded (ds)) or unpaired (i.e. single-stranded (ss)) nucleotides. Each type of RNase comes with distinct specificities (e.g. RNAse T1 (ssG), RNAse A (ssC/U), RNAse S1 (ssRNA) and RNase V1 (dsRNA)). Probing the same RNA with different RNases in separate experiments is a good way to independently assess complementary RNA structure features (and to also estimate the corresponding false positive rates via consistency checks). The size of these proteins (> 10, 000 Da) (Ehresmann et al. 1987), however, prevents them from easily crossing cellular membranes and from resolving smaller RNA structure details, e.g. small bulges. Their use has thus been limited to in vitro studies so far (Ehresmann et al. 1987; Weeks 2010; Knapp 1989; Woese et al. 1980; Aultman and Chang 1982; Guerrier-Takada et al. 1983; Kertesz et al. 2010).
In vitro experiments have the advantage of allowing to examine select aspects of the complex in vivo environment in isolation, e.g. changes in the ion concentrations, temperature or interaction partners. In vivo, however, many such effects including those that cannot be easily replicated in vitro conspire to create a complex environment that cannot be readily replicated in vitro. This is mostly due to the fact in vivo, intrinsic and extrinsic changes to the transcript happen in a space-wise and time-wise carefully orchestrated way which is often impossible to replicate in vitro. Several experimental and theoretical studies have, for example, confirmed that RNA structure formation in vivo can happen co-transcriptionally and that this yields functional RNA structures that can differ significantly from the so-called minimum-free-energy (MFE) RNA structures predicted for already synthesised transcripts assuming thermodynamic equilibrium (Morgan and Higgs 1996; Meyer and Miklos 2004; Wiebe and Meyer 2010; Lai et al. 2013; Proctor and Meyer 2013). This effect is particularly pronounced for transcripts longer than around 200 nt (Morgan and Higgs 1996), i.e. a significant portion of any transcriptome.
Overall, it should not come as a surprise that RNA structures in vitro have been found to differ from those in vivo (Kwok et al. 2013; Tyrrell et al. 2013; Lai et al. 2013). This has major implications for how we should computationally model RNA structures and RNA–RNA interactions that are functionally relevant in vivo. As we will see in the following, many well-known and commonly-used computational methods for predicting these features are based on the assumption that the RNA in question is in thermodynamic equilibrium (and already fully synthesised).
2.2 Experimental Methods for Determining RNA Structures In Vivo
Many existing experimental methods for RNA structure determination in vivo rely on small structure probing molecules (< 500 Da) that (a) can either be readily introduced into living cells via the cellular membrane (Kwok et al. 2013; Zaug and Cech 1995; Wells et al. 2000; Moazed et al. 1986; Harris et al. 1995; Merino et al. 2005; Wilkinson et al. 2006; Mortimer and Weeks 2007; Watts et al. 2009; Steen et al. 2012; Rice et al. 2014; Spitale et al. 2015) or that (b) be generated directly inside the cell (e.g. hydroxyl radicals generated by the high-flux photon beam of a synchrotron source (Latham and Cech 1989; Sclavi et al. 1997)). One exception is RNA structure probing via cryo-electron-microscopy (cryo-EM) (Lengyel et al. 2014) which shall not be discussed here as it is a low-throughput. Similar to RNases, both strategies ((a) and (b)) can be used to probe many RNAs simultaneously, i.e. in a massively parallel fashion. Unlike RNases which act by cutting the transcripts into shorter sub-sequences, these strategies only modify individual nucleotides of the underlying transcripts chemically. Compared to RNases, these chemical RNA structure probing methods thus have the significant, strategic advantage of respecting the linear identity of the underlying transcript. One significant disadvantage of these chemical RNA structure probing methods, however, is that higher-dimensional information on secondary and tertiary RNA structure features is converted into position-specific information along the linear sequence of the transcript. This linearisation implies, in particular, that any direct information on actual base pairs is entirely lost.
The main task of the computational interpretation is thus to convert the experimental probing information for individual nucleotides back into RNA structures involving actual base pairs. It is important to note here that all of these experimental methods chemically modify single, individual nucleotides, but that the reason for each such modification typically extends well beyond the confines of the modified nucleotide itself. That is, the modified nucleotide captures its wider secondary and tertiary RNA structure context. It is thus not entirely appropriate to say that these chemical RNA structure probing methods have single-nucleotide resolution. We will see later on that this has important implications for the computational interpretation of the experimental structure probing data.
Depending on the chemical used for chemical RNA structure probing, these methods can be sub-divided into those that target unpaired nucleotides in a nucleotide-specific way and those that act in a ribose-specific way, see Table 1 for an overview. The first group comprising DMS and CMCT modifies distinct positions in a nucleotide-specific way, but unpaired nucleotides only, whereas reagents of the second group (so-called SHAPE reagents) alkylate the C2’-hydroxyl group of the ribose and thereby the group acts in a way which is neither nucleotide-specific nor completely pairing-status-specific. SHAPE stands for selective 2’-hydroxyl alkylation analysed by primer extension (McGinnis et al. 2012; Merino et al. 2005; Weeks 2010). SHAPE reagents assess the flexibility of the RNA backbone and thereby probe the local RNA structure environment of each type of nucleotide. Raw SHAPE reactivity values thus have the advantage of covering both paired and unpaired nucleotides in any given RNA. The downside, however, is that the distributions of SHAPE values for paired and unpaired nucleotides typically have a non-negligible overlap which requires carefully computational dis-entangling. An additional complication arises due to the fact that all SHAPE reagents also react with water. Different SHAPE reagents have different half-lives in water (t 1∕2 hydrolysis at a specific temperature) spanning several orders of magnitude. These details have to be carefully considered for making the correct choice for each specific research question, e.g. when trying to investigate RNA structure features as function of time.
In principle, it is also possible to probe RNA structure features with molecules that occur naturally to some extent in living cells, e.g. hydroxyl radicals (Latham and Cech 1989; Sclavi et al. 1997). Similar to SHAPE reagents, this chemical acts in a ribose-specific manner and acts both on paired and unpaired nucleotides. Unlike all SHAPE reagents, however, it modifies the C4’-H group (rather than the C2’-hydroxyl group) of the ribose and thereby tends to probe the tertiary RNA structure environment of individual sequence positions. In normal circumstances in vivo, the concentration of hydroxyl radicals is too low for RNA structure probing. In order to artificially increase the concentration for successful RNA structure probing in vivo, X-ray radiation can be used, e.g. generated by a synchrotron source which can generate photon beams of sufficiently high flux. This has already allowed RNA structure probing with high, time-wise resolution in vitro (Sclavi et al. 1997) and in vivo (Soper et al. 2013).
2.3 Experimental Methods for Transcriptome-Wide Probing RNA Structures In Vivo
The above methods for the chemical probing of RNA structures in vivo can naturally probe many RNAs simultaneously. The key achievement of the last few years was to realise that these methods can be combined with high-throughput transcriptome-wide next-generation sequencing (NGS). For this, RNA structure information is first converted into a linearised sequence signal. This is done for many transcripts in parallel. In a second step, these linearised sequence signals are efficiently read out using high-throughput sequencing (typically, NGS).
The corresponding experimental methods can be classified according to (a) the chemical used for RNA structure probing and (b) the protocol employed for converting structure probing information into sequence-based information that can be read in a parallelised fashion using NGS techniques. The second step can comprise a variety of different extraction, depletion and enrichment steps whose features are also key determinants of the overall sensitivity and specificity of the combined experimental protocol.
As the focus here is on in vivo methods, we review in vitro methods for transcriptome-wide RNA structure only briefly. Historically, PARS (parallel analysis of RNA structures) was the first to assess RNA structures in a massively parallel fashion using RNases for enzymatic RNA structure probing (Kertesz et al. 2010; Wan et al. 2012; Righetti et al. 2016; Wan et al. 2014, 2013; Del Campo et al. 2015). Other in vitro approaches have since included those based on enzymatic structure probing (ds/ssRNA-seq (Zheng et al. 2010; Li et al. 2012a,b) and Frag-seq (Underwood et al. 2010) as well as approaches based on chemical probing (DMS-seq (Rouskin et al. 2014) and RING-MaP (Homan et al. 2014) using DMS, HRF-seq (Kielpinski and Vinther 2014) and MOHCA-seq (Cheng et al. 2015) using hydroxyl radicals, SHAPE-seq and SHAPE-seq 2.0 (Lucks et al. 2011; Loughrey et al. 2014; Watters et al. 2016b) (using SHAPE reagent 1M7) and SHAPES (Poulsen et al. 2015) (using SHAPE reagent NPIA)). Some in vitro methods employ two or more chemical reagents, e.g. ChemMod-seq (Hector et al. 2014) (DMS and SHAPE reagent 1M7), MAP-seq (Seetin et al. 2014) (DMS, CMCT and SHAPE reagent 1M7) and CIRS-seq (Incarnato et al. 2014) (DMS and CMCT). It is especially advantageous to combine nucleotide-specific with ribose-specific chemical modifications as these complement each other and enable valuable cross-checks. These in vitro methods are appropriate for RNA structure probing, if the artificial setting can be justified for addressing specific scientific questions. Care has to be taken, however, not to simply generalise these in vitro results to various in vivo settings.
All of the existing in vivo methods employ chemical probes for RNA structure probing. In all cases, the raw structure probing data consists of probing values for individual sequence positions, not base pairs. Most of the currently existing in vivo methods employ DMS as structure probing reagent, such as Structure-seq (Ding et al. 2014, 2015), DMS-seq (Rouskin et al. 2014), Mod-seq (Talkish et al. 2014; Lucks et al. 2011) and targeted Structure-seq (Fang et al. 2015). In addition, SHAPE-based approaches such as SHAPE-MaP (Smola et al. 2015a,b; Siegfried et al. 2014; Lavender et al. 2015; Mauger et al. 2015) (SHAPE reagents: 1M7, 1M6 and NMIA) and icSHAPE (Spitale et al. 2015; Flynn et al. 2016) (SHAPE reagent: NAI-N3) now exist, as well as earlier in vitro approaches such as SHAPE-seq (Lucks et al. 2011; Mortimer et al. 2012) (SHAPE reagent: 1M7) that were extended to combine the earlier SHAPE-reagent with DMS-based probing in cell SHAPE-seq (Watters et al. 2016a,b), see Table 2 for an overview. The major steps of all currently existing in vivo RNA structure probing methods are
2.3.1 Step 1: RNA Structure Probing
The goal of this step is to probe RNA structures using a reagent that induces chemical modifications into individual nucleotides.
The key aspect to consider is: Could any step of the protocol for RNA structure probing actually interfere with the in vivo RNA structures in a way which would alter them before they are probed?
This is perhaps the most important aspect to optimise. If this fails, no subsequent step in the experimental or computational analysis can fix it. (1) For this, the chemical properties of the probing reagents need to considered and their potential direct or indirect impact on RNA structure features be examined, e.g. in dedicated in vitro experiments prior to the in vivo ones. These experiments have to be conducted in a way that can distinguish reactions on different time-scales. (2) It is also important to consider the possibility that the chemical modifications induced during RNA structure probing alter the RNA structure while it is being probed. (3) Lastly, if RNA structure probing is done by more than a single probing reagent, this should happen in separate experiments keeping everything, but the probing reagent, unchanged.
In terms of future developments, it would be beneficial to have fast and efficient ways to stop RNA structure probing in vivo. This would help to conserve the RNA structure probing signal and allow detailed investigations of RNA structures as function of time.
2.3.2 Step 2: RNA Extraction, rRNA Depletion and RNA Enrichment
In this step, the pool of chemically modified transcripts of interest is extracted and enriched and unwanted transcripts are removed to prevent them from being sequenced (e.g. rRNAs which account for the majority of transcripts, yet are typically not the focus of the investigation).
The key challenge here is to ensure that extraction and enrichment are done with maximum specificity. Any true signal lost cannot be recovered later on.
For enrichment, a polyA RNA enrichment step is often applied. This implies, however, that non-polyA transcripts (e.g. non-coding RNAs, circular RNAs) are omitted from all subsequent steps of the analysis. The user needs to decide whether this is actually wanted and otherwise adapt the original protocol.
2.3.3 Step 3: Library Preparation for High-Throughput Sequencing
Different in vivo RNA structure probing protocols differ substantially in how the enriched pool of chemically modified transcripts is converted into a library for NGS sequencing. As soon as the library has been sequenced, the corresponding reads have to be mapped back to the underlying genome/transcriptome before the computational analysis of RNA structure features can start. As this mapping comes with its own significant challenges, it is imperative to optimise the experimental library preparation w.r.t. the subsequent computational analysis.
2.3.4 Key Aspects to Consider for Optimisation
-
(A)
What is the expected average length of the final reads (excluding the length of any primers and/or adapters that are removed in silico prior to mapping the reads back to the genome/transcriptome)?
For those methods that detect RNA structure probing signals via chemical-induced reverse transcriptase halting, e.g. Structure-Seq and icSHAPE, this length is primarily determined by the average distance between the initiation site of reverse transcription (RT) and the first chemically modified nucleotide upstream. It thus depends both on the specificity of the chemical used for RNA structure probing as well as the mechanism used for RT initiation (example: DMS (which only probes unpaired nucleotides) and random hexamer primers for RT initiation in case of Structure-Seq). Note that the mechanism used for RT initiation (e.g. random primers of different lengths may preferentially bind to single-stranded regions of the transcript) may introduce its own biases that may be relevant to the subsequent, computational RNA structure interpretation. The effective average read length may also be influenced by additional RNA fragmentation steps, e.g. random fragmentation by Mg2+-mediated hydrolysis in icSHAPE. For these methods, a well-chosen combination of probing reagent and RT initiation can thus optimise the expected average read length.
For those methods that detect RNA structure probing signals via chemical-induced reverse transcriptase read-through, e.g. SHAPE-Map (Siegfried et al. 2014; Smola et al. 2015a,b; Lavender et al. 2015; Mauger et al. 2015), the natural average length of reads is primarily determined by the default fragmentation step of the corresponding library preparation protocol (Nextera in case of SHAPE-Map) and not by the average distance between RT initiation and any nucleotides modified via chemical RNA structure probing. This is a significant conceptual advantage over methods that detect RNA structure probing signals via reverse transcriptase halting.
-
(B)
How much RNA structure probing information is retained in a single read?
Ideally, we would like to retain structure probing information for entire, individual transcripts. If we loose this information, e.g. during library preparation, we cannot detect RNA structure diversity, i.e. the possibility that different copies of the same transcript assume different RNA structures in vivo. Also, in order to maximise the RNA structure information for each individual transcript, chemical RNA structure probing should happen in a way that saturates each transcript with structure probing signals (in a way which does not risk altering the underlying RNA structure itself).
For most of the existing protocols for RNA structure probing in vivo, however, the requirements for optimising the library preparation are not in line with the above requirements for optimising the RNA structure probing information. The library preparation of Structure-Seq and icSHAPE, for example, is set up to generate reads that correspond to one chemically modified nucleotide only, namely the chemically modified sequence position that is first encountered upstream of the RT initiation site (chosen by a hexamer primer in case of Structure-Seq and chosen by Mg2+-induced random fragmentation in case of icSHAPE). Any correlations between RNA structure probing information from the same transcript are thereby lost. In addition, saturated RNA structure probing would have the tendency to further lower the average read length, making the subsequent mapping even harder.
The best way to circumvent this problem is to choose a library preparation protocol that does not rely on RT transcriptase halting for detecting the RNA structure probing signal. This can, for example, be done using Mn 2+ mediated reverse transcriptase read-through of the modified nucleotide positions as in SHAPE-MaP. This strategy, however, has the undesired side effect of introducing a generally higher error rate for reverse transcription.
-
(C)
What is the overall efficiency of all steps in the protocol?
Some protocols, e.g. icSHAPE, incorporate a second enrichment step by chemically treating the RNA-structure-probed transcripts in a second step in vivo to prepare their subsequent biotinylation using click-chemistry (this happens after RNA extraction, rRNA depletion and RNA enrichment). This second biotin-based enrichment step has the advantage of further increasing the specificity.
Overall, protocols for in vivo RNA structure probing differ substantially in the number of steps required for library preparation. Any additional steps in the overall protocol, however, have the tendency of reducing the overall sensitivity and efficiency as the inefficiencies and biases of each step add up. Generally, it is thus advisable to minimise the total number of steps and to optimise each step in terms of specificity and sensitivity.
3 Interpreting the Experimental RNA Structure Probing Data In Silico
The above in vivo methods for transcriptome-wide RNA structure probing generate raw transcriptome sequencing data (reads) which must be computationally processed and interpreted for any actual RNA structures to be inferred.
Basically, any computational analysis has to achieve the reversal of the experimental protocol, namely to convert a purely sequence-based signal back into RNA structures involving base pairs. This is challenging due to a number of reasons:
-
(a)
The sequence signals induced by chemically encoded RNA structure probing can be noisy, biased and/or incomplete. For example, any particular SHAPE values cannot be unambiguously interpreted as being derived from a paired or unpaired nucleotide.
-
(b)
RNA structure probing information from any transcript is fragmented in the existing experimental protocols, i.e. the full sequence identity of the RNA structure probing signal is lost and cannot be retrieved later computationally. Correlated structure probing information is currently only retained within individual reads.
-
(c)
Next-generation sequencing itself introduces errors and biases, e.g. sequencing errors whose rate depends on the position within each read.
-
(d)
The mapping of sequenced reads to a reference genome/transcriptome is not straightforward and can induce different kinds of errors, biases and missing data. This is a particular concern for experimental protocols that encode RNA structure probing information in terms of nucleotide changes, e.g. SHAPE-MaP. There, sequenced reads cannot be readily mapped back to their original transcripts without carefully considering SNP-like discrepancies. This requires dedicated, probabilistic mapping methods such as those used in transcriptome-wide RNA editing studies, see e.g. (Mazloomian and Meyer 2015).
-
(e)
Only once the sequenced reads have been mapped to a reference transcriptome, can the actual inference of RNA structures begin. This can be done using a range of conceptually different computational strategies. These are introduced in the following.
Most existing computational methods focus on utilising SHAPE reactivity values as input information to infer RNA structure information. The following describes different underlying conceptual strategies for converting raw SHAPE reactivity values along one linear transcript into distinct RNA structure(s). These approaches not only employ different strategies for RNA structure prediction, but also differ in the (implicit or explicit) assumptions they make in interpreting the raw structure probing data. Roughly, all existing computational approaches can be classified according to how they address three main aspects:
-
(a)
How the raw, sequence-position-specific RNA structure probing is processed. Examples include re-scaling and normalisation procedures.
-
(b)
How the raw, sequence-position-specific RNA structure probing is interpreted and integrated into RNA structure prediction.
-
(c)
How RNA structures are captured in a predictive model that utilises experimental RNA structure probing data. All of these methods model RNA structures at secondary-structure level. These methods differ substantially in their implicit and explicit assumptions. Examples include thermodynamic methods that derive the thermodynamically most stable RNA secondary structure (so-called minimum-free energy (MFE) methods), methods that consider Boltzmann ensembles of RNA secondary structures in thermodynamic equilibrium and, most recently, probabilistic methods for RNA secondary structure prediction that predict the maximum likelihood RNA secondary structure, see Table 5.
Table 5 Characteristic features of the computer programs that predict RNA secondary structure by combining sequence data and chemical RNA structure probing data Table 3 Prediction performance of several computer programs that utilise individual sequences and corresponding SHAPE data as input to make RNA secondary structure predictions (optimal values highlighted in bold) Table 4 Changes in prediction performance of ProbFold and RNAstructure as different types of RNA structure probing are provided as combined input
As we will see in the following, early methods incorporate experimentally derived RNA structure probing information into thermodynamic methods for RNA secondary structure prediction (MFE approach). More recently, RNA structure probing information has been integrated in a fully probabilistic manner into probabilistic methods for RNA secondary structure prediction. These new methods offer conceptually convincing ways of seamlessly combining experimental RNA structure probing data with RNA structure prediction.
3.1 Interpreting SHAPE Reactivity Values as Pseudo-Energies for Paired Sequence Positions
Many commonly used computational methods for RNA secondary structure prediction, e.g. Mfold (Zuker 2003) and RNAfold (Zuker and Stiegler 1981), utilise a so-called thermodynamic model of RNA secondary structures. These methods decompose any (pseudo-knot-free) RNA secondary structure into a sum of Lego-like, structural RNA secondary structure building blocks and express the total free energy of the RNA structure as sum of the free-energy contributions of these structural building blocks. The underlying thermodynamic models, e.g. the well-known Turner model (Mathews et al. 1999) on which Mfold and RNAfold are based, rely on many parameters that correspond to physical entities that have been determined experimentally. For a given input RNA sequence, these models employ efficient dynamic programming algorithms such as the Zuker–Stiegler algorithm (Zuker and Stiegler 1981) to derive the RNA secondary structure with the minimum overall free energy. The corresponding minimum-free-energy (MFE) structure is reported as output. For any given input sequence, these methods predict a single MFE RNA secondary structure. Thermodynamic methods for RNA secondary structure prediction such as Mfold and RNAfold make the implicit assumptions that any given input sequence (a) is already fully synthesised and (b) that it will assume an MFE RNA secondary structure. In particular, these methods assume any input RNA to be in thermodynamic equilibrium and to be naked, i.e. without any trans interaction partners such as ligands, proteins or other RNAs. As we know, this assumption is generally not justified in in vivo settings.
Early efforts to integrate chemical RNA structure probing data into RNA structure prediction try to interpret these data as modifications to the default thermodynamic model used for RNA secondary structure prediction. For this, experimentally determined RNA structure probing values are somehow converted into free energy contributions assigned to individual sequence positions.
Deigan et al. (2009) were the first to interpret the position-specific SHAPE reactivity values α i as position-specific free-energy corrections \(\Delta G_i^D\) to the nominal free energy terms in the thermodynamic model for RNA structure prediction:
Here, α i denotes the experimentally determined SHAPE reactivity value for sequence position i in the transcript (i.e. i ∈{1, …, L} for a transcript of L nucleotides length) and m and b are free parameters with default values m = 2.6 and b = −0.8 kcal mol−1, see Low et al. (2014), Qi et al. (2012) for other parametrisations. In the dynamic programming recursion which derives the most stable RNA secondary structures, these \(\Delta G_i^D\) values are added to the nominal energy contribution for each base-paired sequence position i. Any contributions from SHAPE reactivity values from un-paired sequence positions are completely ignored.
This approach by Deigan was later extended to work on DMS input data (Cordero et al. 2012a); pseudo-energies are derived from a log-likelihood ratio of a nucleotide being unpaired versus paired. Eddy (2014) pointed out that base-pairing probabilities for individual sequence positions, p i, can be linked to position-specific pseudo-energies if one may assume that a naked, fully synthesised RNA is in thermodynamic equilibrium. This can be achieved because \(p_i(\pi _i = 1) \propto e^{-\Delta G_i/R T}\). That is, the probability that sequence position i is base-paired, i.e. p i(π i = 1), is proportional to \(e^{- \Delta G_i/R T}\), where ΔG i is the pseudo-energy assigned to position i (here, R denotes the universal Gas constant and T the absolute temperature in degrees Kelvin).
3.2 Interpreting SHAPE Reactivity Values as Pseudo-Energies for Paired and Unpaired Sequence Positions
The above approach by Deigan introduces an unnatural bias into the interpretation of SHAPE reactivity values. Even though experimentally determined SHAPE reactivity values have a continuous spectrum, covering both paired and unpaired nucleotides, SHAPE-derived pseudo-energies are effectively only assigned to paired sequence positions.
Zarringhalam et al. (2012) propose a strategy which is symmetric w.r.t. paired and unpaired sequence positions. Similar to Deigan, they interpret SHAPE reactivity values α i along the transcript as position-specific corrections \(\Delta G_i^Z\) to the free energy terms of the underlying transcript position i:
Here, \(\alpha _i^r\) denotes the (rescaled version of the) experimentally determined SHAPE reactivity value and π i is the corresponding pairing status of sequence position i, i.e. π i = 0 for an un-paired and π i = 1 for a paired sequence position. The rescaling of the original SHAPE reactivity values α i is achieved via a piecewise-linear function which re-scales the values so that the resulting values satisfy \(\alpha _i^r \in [0,1]\). The shape of this function was chosen to fit to the empirical likelihood ratio distribution, i.e. the paired-unpaired likelihood ratios as function of the SHAPE reactivity values. The scaling parameter β affects all sequence positions equally and can be interpreted as a universal knob to decrease or increase the contribution of SHAPE values in the thermodynamic model for RNA structure prediction.
The goal of the Zarringhalam approach is to minimise the overall difference between the experimentally derived SHAPE data and the predicted RNA structure as measured by the so-called Manhattan distance, i.e. to minimise \(\sum _i | \pi _i - \alpha _i^r|\). Unlike the above approach by Deigan, this strategy can be mathematically shown to yield a better fit of the predicted RNA structures to the SHAPE reactivities in terms of Manhattan distance (Zarringhalam et al. 2012).
3.3 Introducing Pseudo-Energy-Like Free Parameters in a Fit to a Thermodynamic Ensemble of RNA Secondary Structures
Both above approaches implicitly assume that all SHAPE reactivity values correspond to a single RNA secondary structure. Washietl et al. (2012) stick to the assumption of a naked, already synthesised RNA sequence in thermodynamic equilibrium but interpret the SHAPE reactivity values as ensemble-weighted average values over many identical RNAs with different RNA secondary structures. Many properties of this so-called Boltzmann distribution of RNA secondary structure in thermodynamic equilibrium can be calculated analytically (McCaskill 1990; Miklos et al. 2005).
Their method works as follows. In a first step, SHAPE values for each sequence position i, α i, are translated into so-called pairing probabilities p i(α i) with p i(α i) = 0 if α i > 0.25 and p i(α i) = 1 if α i ≤ 0.25. Using this simple thresholding procedure, SHAPE reactivity values are thus effectively interpreted as either being paired or unpaired (with 100% probability, i.e. certainty). These position-specific p i(α i) values should thus be viewed as pairing status indicators, e.g. denoted by s i := p i(α i), rather than pairing probabilities.
Any discrepancies between the position-specific pairing probabilities \(z_i(\theta , \vec {e})\) as they can be explicitly calculated from the Boltzmann ensemble of RNA structures in thermodynamic equilibrium (where θ denotes the set of default parameters of the underlying thermodynamic model and \(\vec {e}\) a vector of so-called pseudo-energy corrections e i introduced for each individual sequence position i) and the position-specific SHAPE-derived pairing status values s i are assumed to be normally distributed with a position-independent variance σ 2. Every sequence position i in the transcript of L nucleotides length, i.e. i ∈{1, …L}, is assigned a so-called pseudo-energy term e i. In contrast to the above approaches by Deigan and Zarringhalam, however, these e i values do not have a link to SHAPE reactivities. Rather, they correspond to position-specific free parameters in a global optimisation problem and have been artificially introduced. Also these e i terms are assumed to come with a position-independent, overall variance of τ 2. Using a gradient descent method, the method by Washietl et al. then tries to identify the vector of e i values that minimises the expression:
This optimisation can be expected to be mathematically challenging as the optimisation procedure is not guaranteed to find the global minimum and can get stuck in local minima. A priori, it is also not clear what the correct interpretation of the resulting e i values should be. They have no obvious link to SHAPE reactivity values nor to the free parameters of the underlying thermodynamic model (θ). Also, it should be noted that the number of free parameters e i increases linearly with the length of the input sequence and that the optimisation is done for each input sequence independently.
The current implementation of the Washietl approach into the ViennaPackage (Lorenz et al. 2016) allows users to explore different ways of converting structure probing data into p i values and provides several optimisation techniques.
3.4 Using SHAPE Reactivity Values in a Sample and Select Approach Using an Unperturbed Thermodynamic Ensemble of RNA Secondary Structures
All of the above approaches hinge on the validity of the assumption that experimental structure probing data can be interpreted as position-specific pseudo-energy corrections to an underlying thermodynamic model. As the detailed discussion of the above methods shows, even incorporating this assumption into a corresponding strategy for RNA structure prediction is technically and conceptually not entirely straightforward.
Some groups (Ouyang et al. 2013; Quarrier et al. 2010) have decided not to interpret structure probing data as position-specific pseudo-energy corrections at all. Instead, they assume that the in vivo environment introduces unknown changes to the nominal RNA structure of the underlying thermodynamic model (i.e. the MFE-structure as defined earlier) which cannot be modelled by tweaking the underlying parameters of the thermodynamic model. This makes sense as some effects of the in vivo environment, e.g. trans interaction partners, can conceptually not be captured by tweaking the free energy parameters of the thermodynamic model for RNA secondary structure prediction. Instead, they propose to address this challenge by sampling RNA secondary structures from the (unperturbed) thermodynamic ensemble of RNA secondary structure (Ding and Lawrence 2003; McCaskill 1990) and re-ranking the sampled RNA structures according to how well they fit the experimentally determined RNA structure probing data. This involves a distance metric such as the Manhattan distance introduced above. For calculating the fit, SHAPE reactivity values are first mapped to discrete paired/unpaired values for each sequence position using a simple thresholding approach before calculating the Manhattan distance to the sampled RNA.
These methods effectively allow for more than a single RNA secondary structure to correspond to one set of experimentally determined, position-specific RNA structure probing data, even though these RNA secondary structures conceptually derive from the same Boltzmann ensemble of many identical RNA sequences in thermodynamic equilibrium. By ranking the sampled RNA structures based on fit to the probing data only (rather than the respective probability of the sampled RNA structure in the Boltzmann ensemble), all sampled RNA secondary structures are effectively assumed to have equal prior probability (provided they are sampled at all). The obvious downside of this pragmatic approach is that RNA secondary structure with low probability in the Boltzmann ensemble may never be sampled at all, even if they could provide the best overall fit. Also, this approach only provides limited feedback in terms of insight gained.
3.5 Probabilistic Integration of Experimental RNA Structure Probing Data into Probabilistic Methods for RNA Secondary Structure Prediction
RNA secondary structure prediction does not necessarily need to involve the assumption that any input RNA folds into the minimum-free-energy structure and is in thermodynamic equilibrium. Using probabilistic methods such as stochastic context-free grammars (SCFGs) (Durbin et al. 1998) (or Markov Chain Monte Carlo (MCMC) methods), it is possible to explicitly capture different hypotheses on how RNA secondary structure may arise. This has given rise to a number of RNA secondary structure prediction methods, e.g. Pfold (Knudsen and Hein 2003), RNA-Decoder (Pedersen et al. 2004a,b), SimulFold (Meyer and Miklos 2007), that yield a high prediction performance for evolutionarily conserved RNA secondary structures. These methods combine a probabilistic model of RNA secondary structures with computationally efficient algorithms to derive the maximum likelihood RNA structure given the underlying RNA structure model. In terms of time-and-memory efficiency, they have the same complexity as thermodynamic methods, e.g. Mfold (Zuker 2003) and RNAfold (Zuker and Stiegler 1981), but offer several conceptual advantages. First, the user can decide the parametrisation of the model. Free parameters can thus be chosen to have a straightforward biological interpretation. Second, given a training set of sufficient size and complexity, the free parameters of the model can be explicitly trained. Third, alternative parametrisations of the same model can be explicitly evaluated and ranked based on likelihood fits to the data. Fourth, the predictive model for RNA secondary structures can be readily extended to take into account additional sources of input information, e.g. evolutionary information in terms of a multiple-sequence alignment (MSA) or experimental RNA structure probing data.
Technically, this can be achieved by replacing the so-called emission probabilities of SCFGs by probabilistic emission models that, for example, read entire alignment columns from an input MSA rather than individual nucleotides from an input sequence. These emission models are probabilistic models that can, for example, explicitly capture how we expect paired and unpaired nucleotides to evolve as function of evolutionary time.
Most importantly, fully probabilistic models allow information of different types (e.g. primary sequence features, RNA structure features, evolution) to be seamlessly merged as the corresponding probabilities for different sources of information can be readily combined in a single predictive framework. This elegantly avoids the need for converting conceptually different sources of information (e.g. chemical RNA structure probing data) into units with a physical interpretation (free energy terms). More importantly, probabilistic models allow us to move beyond the assumption of thermodynamic equilibrium.
3.5.1 Integration into Comparative Methods for RNA Secondary Structure Prediction
PPfold 3.0 (Sükösd et al. 2012) (PPfold in the following) were the first to integrate external RNA structure probing information into a fully probabilistic model of RNA secondary structure prediction.
The model for RNA structure prediction is identical to Pfold (Knudsen and Hein 2003), a comparative RNA secondary structure prediction method. It takes as input a multiple-sequence alignment (MSA) and a corresponding evolutionary tree linking the sequences in the MSA and returns as output the maximum-likelihood RNA secondary structure for the input alignment and input tree. Pfold captures the assumption that RNA secondary structures that have been conserved during evolution are likely to be functional. As far as we know, this is overall a decent assumption to make. In practice, the success of the comparative approach depends on a decent choice of the appropriate evolutionary distances of the sequences in the input alignment. The RNA structure predicted by Pfold corresponds to the maximum-likelihood RNA secondary structure given the input information and the predictive model and its parameters. The evolutionary relationships of the sequences in the input multiple-sequence alignment are explicitly modelled using two probabilistic models of evolution that capture how unpaired and base-paired nucleotides evolve as function of time, respectively.
The novelty of PPfold consists of combining comparative RNA secondary structure prediction with experimental RNA structure probing information. In order to do this, the user needs to specify a probability distribution P(H|σ) for a set of experimental probing data H and secondary structures σ. PPfold generally assumes that P(H|D, σ) = P(H|σ), i.e. that there is no dependence on the actual observed nucleotides sequences of the input alignment D. As the discussion of the more recently published method ProbFold (Sahoo et al. 2016) below shows, this is probably too simplistic: It can actually be shown that SHAPE-values typically do depend on nucleotide identity. As an alternative to P(H|σ), the user can also specify values P(H i|i unpaired) and P(H i|i paired), i.e. likelihood values that sequence position i in the input alignment is unpaired or paired given the experimental probing value of H i for that sequence position. Internally, PPfold uses these likelihood values as follows to bias the nominal likelihood values of Pfold for each paired (i, j) (subscript d for double) and unpaired i (subscript s for single) alignment column, P d(i, j) and P s(i):
This assumes that the experimental probing values for the two sequence positions involved in a base-pair are assumed to be independent. The validity of this assumption has since been confirmed by the more recent investigations of ProbFold, see below for details.
Similar to Pfold, PPfold naturally reduces to a non-comparative RNA secondary structure prediction method if the input alignment consists of only a single input sequence (although it should be stressed that this is not how Pfold nor PPfold are meant to be used). The authors of PPfold deliberately use it with single input sequences in order to make it directly comparable to the non-comparative RNA secondary structure program RNAstructure (Deigan et al. 2009; Mathews et al. 2004) which also utilises external RNA structure probing data as additional input information. RNAstructure and PPfold (using single sequences) have a similar performance in terms of F-value. The F-value is defined as the harmonic mean of sensitivity and specificity. This is an impressive result given that the RNA secondary structure model of PPfold is lightweight compared to the full- fledged thermodynamic model underlying RNAstructure. PPfold thus makes better use of the external RNA structure probing information than RNAstructure. The performance of PPfold w.r.t. RNAstructure can be further improved in terms of F-value when using PPfold with multiple sequence input alignments. As with many comparative RNA secondary structure prediction methods, however, the resulting performance in terms of F-value critically depends on the quality of the input alignment. A poor input alignment (with or without additional probing data) can lower the performance of PPfold below the corresponding single-sequence performance with experimental probing data. That is, a poor input alignment can provide more confusion than can be remedied by additional RNA structure probing data.
Note that due to the scarcity of training and testing data, the authors of PPfold could not avoid an overlap between their training set (16S and 23S rRNA structures and SHAPE data for Escherichia coli) and their test data set (16S rRNA of E. coli).
3.5.2 Integration into Non-comparative Methods for RNA Secondary Structure Prediction
Most recently, Sahoo et al. (2016) proposed ProbFold, a probabilistic method for non-comparative RNA secondary structure that can integrate information from one or more chemical RNA structure probing experiments. ProbFold employs a fully probabilistic stochastic context-free grammar (SCFG) for RNA secondary structure predictions and combines this with probabilistic graphical models (PGMs) (Koller and Friedman 2009) to capture experimental probing data. Compared to PPfold ProbFold offers a more general modelling approach that is also more readily extendible and more parameter-sparse. The SCFG employed by ProbFold is based on the original grammar underlying Pfold (Knudsen and Hein 2003) with extensions that capture stacking interaction, i.e. correlations between pairs of adjacent base pairs. Overall, the ProbFold grammar consists of six production rules in total, three of which emit terminals, i.e. read information from the input sequence. These three production rules require three corresponding emission models called single, pair and stack that model single, pairs and two adjacent pairs of sequence positions, respectively, see Fig. 2 in Sahoo et al. (2016) for a visualisation. The integration of experimental probing data into the RNA secondary structure prediction method happens via three corresponding PGMs that each specify a joint distribution over the RNA primary sequence data and the experimental probing data. Technically, each PGM corresponds to an undirected bipartite graph between so-called factor nodes and so-called variable nodes. The variable nodes represent random variables, whereas the factor nodes correspond to probability distributions between neighbouring random variables. ProbFold uses discrete random variables for the efficiency of the calculations. This is technically achieved by discretising the two distributions P single and P paired which model the corresponding distributions of experimental probing data. For this, probing data is first discretised into k bins using normalised histogram models (i.e. multinomials). This implies k − 1 free parameters specifying the boundaries of these bins. These are chosen to maximise the difference between the probing data distributions of paired and unpaired sequence positions using Kullback Leibler (KL) divergence.
During the development of ProbFold, a hierarchy of increasingly complex, fully probabilistic models with an increasing number of free parameters (ranging from 18 to 408, for the final model) was investigated. The final model of ProbFold has only a single user-specified meta-parameter, corresponding to the number of bins used for discretising the two distributions P single and P paired of the experimental probing data (default is six bins). All other free parameters can be explicitly derived using a dedicated set of training set of known RNA secondary structure with corresponding structure probing data. The final model captures not only stacking interactions between neighbouring base pairs (so-called stack-part of the model), but also correlations between the structure probing values of neighbouring positions along the linear sequence (so-called cor-part of the model). Due to the scarcity of the training data, the primary sequence and structure probing values are modelled independently in order to keep the number of free parameters low. The trade-off between the sensitivity and the specificity of performance can be explicitly adjusted via a parameter γ. Sahoo et al. carefully evaluate the performance of ProbFold, using a dedicated test set which has no overlap with the training set (they are actually the first to do this properly using a cross-evaluation procedure). Reassuringly, they can conclude that over-fitting is not an issue, implying that their method is sensibly parametrised and the number of free parameters in line with the information content provided by their training set.
In terms of performance, they compare ProbFold to PPfold 3.0 (Sükösd et al. 2012), RNAstructure v5.6 (Deigan et al. 2009; Mathews et al. 2004), GTFold-3.0 (Swenson et al. 2012) and RNAfold.zar (Lorenz et al. 2011, 2016) (this is how they RNAfold in combination with the approach by Zarringhalam for converting the raw SHAPE values) on an independent test data set of 11 RNA structures on which neither of these methods were initially trained. The resulting performance comparison thus allows a fair assessment of the prediction accuracy of several key predictive programs, see Table 3.
The overall performance is measured in terms of F-value, i.e. the harmonic mean of sensitivity and specificity with values of F ∈ [0, 1] with 1 corresponding to perfect predictions. For ProbFold, this is done for a fixed value of γ. ProbFold comes second in terms of overall F-value and accuracy across all structures after RNAfold.zar (F-values 0.77 and 0.71, respectively), but first in terms of performance gain w.r.t. purely sequence-based predictions without any SHAPE input (ΔF = 0.29 (ProbFold) compared to ΔF = 0.12 (RNAfold.zar)). This is impressive given that RNAfold employs the state-of-the-art thermodynamic model for predicting RNA secondary structures, whereas ProbFold uses a fairly light-weight SCFG with a significantly smaller number of parameters. (In that regard, it is also instructive to compare the baseline performance for single-sequence-only input between ProbFold and RNAstructure, see Table 4.) Of all methods assessed, ProbFold is found to be the most robust w.r.t. increasing levels of noise. This is quantitatively assessed using different levels of simulated noise. Based on these results, one can conclude that ProbFold makes best use of the external RNA structure probing information. Using a slightly more complex SCFG for modelling RNA secondary structures or employing a comparative approach such as Pfold should allow ProbFold’s baseline performance to be further improved in the future.
Apart from the benchmark performance evaluation, the ProbFold study offers several important biological insights. First, they find that the SHAPE reactivities for paired and unpaired regions depend significantly on the primary nucleotide sequence. Furthermore, they find that the SHAPE reactivities for neighbouring sequence positions are significantly correlated, both for base-paired and especially for unpaired nucleotides. This is to be expected given that the SHAPE reactivities measure the backbone flexibility of the RNA transcript which is a notion that extends beyond the confines of the single sequence position that ends up being chemically modified. Based on these observations, Sahoo et al. decided to explicitly capture these correlations within the probabilistic models of ProbFold. Somewhat surprisingly, they find no evidence that the SHAPE reactivities between two base-pairing nucleotides are correlated. They attribute this to the comparatively high level of noise for low SHAPE reactivities. In ProbFold, this finding is captured by modelling the emission models of the left- and right-pairing partner independently using separate distributions.
One of the key advantages of ProbFold is that it can seamlessly integrate more than one kind of experimental structure probing data, e.g. DMS and CMCT probing data in addition to SHAPE reactivities. Initial performance results with a model which assumes independence of the different kinds of experimental evidence show that the performance can indeed be significantly improved as more types of experimental evidence are added, see the results in Table 4. Technically, ProbFold can also be set up to work with SHAPE-seq data (Lucks et al. 2011).
Conceptually, the theoretical framework underlying ProbFold offers a mathematically and conceptually convincing way of integrating experimental RNA structure probing data into models for RNA secondary structure prediction. Unlike most existing methods that are based on thermodynamic models for RNA secondary structure prediction, the number of free parameters in ProbFold that are used to integrate experimental RNA structure probing information does not increase with the length of the RNA. Instead, it only depends on the complexity (i.e. parametrisation) of the underlying predictive model. Moreover, these free parameters have a straightforward interpretation in terms of the experimental RNA structure probing data. By employing purely probabilistic concepts, different assumptions about the dependence or independence between probing data and/or between sequence positions can be made explicit and quantitatively assessed, so we can quantitatively test different hypotheses and also learn something about our data from the model. In addition, its free parameters can be readily retrained as more training data or novel types of experimental RNA structure probing data become available. This is a pre-requisite for cross-evaluating the performance and for examining if over-fitting is an issue (Table 5).
4 Transcriptome-Wide Experimental Methods for Directly Determining RNA Structures and trans RNA–RNA Interactions In Vivo
The structural building blocks of RNA secondary structures and of trans RNA–RNA interactions are base pairs. Yet, none of the transcriptome-wide methods for chemically probing RNA structures in vivo described above retain direct information on base pairs. Rather, information on RNA structure probing is linearised and encoded in individual sequence positions. Any direct information on corresponding pairing partners is lost. This is the main reason why major computational efforts are required to covert the raw position-specific experimental data back into actual RNA structures involving base pairs.
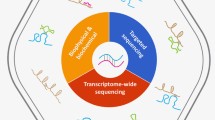
This recently changed as three groups simultaneously proposed experimental protocols for directly determining RNA secondary structure features in vivo in a transcriptome-wide fashion: PARIS (Lu et al. 2016), SPLASH (Aw et al. 2016) and LIGR-seq (Sharma et al. 2016). PARIS stands for psoralen analysis of RNA interactions and structures, SPLASH for sequencing of psoralen cross-linked, ligated and selected hybrids and LIGR-seq for ligation of interacting RNA followed by high-throughput sequencing. In constrast to earlier experimental protocols for probing transcriptome-wide in vivo probing, these three new methods allow to probe RNA structure features in a way which is not specific to any particular RNA-binding protein, see Fig. 1 for an overview.
Overview of the experimental protocols of PARIS, SPLASH and LIGR-seq. Lines in yellow and dark blue denote different transcripts. The black dots represent hydrogen bonds between transcripts. A red ellipse denotes the cross-linked psoralen derivative AMT. The complex between psoralen and biotin is shown in pink and light green, see the SPLASH pipeline. In the library preparation step, the red and green regions denote primers and adapters added, during the corresponding preparation protocols. The main difference between the protocols lies in the enrichment strategies for cross-linked duplexes. SPLASH focuses on biotin- dependent enrichment after fragmentation. PARIS utilises 2D-electrophoresis. LIGR-seq relies on the fact that AMT-cross-linked duplexes are more resistant to RNase R treatment. LIGR-seq requires additional samples to be made, see the text for details. In this figure, we only outline the protocol for making the +AMT+ligase sample
4.1 Experimental Protocols of PARIS, SPLASH and LIGR-seq
All three new methods, i.e. PARIS (Lu et al. 2016), SPLASH (Aw et al. 2016) and LIGR-seq (Sharma et al. 2016), directly probe so-called duplexes, i.e. stretches of more or less consecutive base pairs. Each duplex can either involve the same or two different RNAs and thus either correspond to an RNA structure feature or a trans RNA–RNA interaction. It is important to note that all three experimental protocols process both types of duplexes in an identical manner (and that it is up to their respective, subsequent computational analysis pipelines to detect and distinguish both cases). All three methods are thus methods for both direct RNA structure probing as well as direct probing of trans RNA–RNA interactions. Conceptually, all three protocols have common steps but differ in important details. Their overall logical flow is as follows, see also Fig. 1.
4.1.1 Experimental Protocol of PARIS
In the first step of PARIS, duplexes corresponding to RNA structure features or to trans RNA–RNA interactions are covalently cross-linked using the psoralen derivative 4’-aminomethyltrioxsalen (AMT) and UV-light at 365 nm. For this, AMT intercalates between base pairs and covalently cross-links preferentially juxtaposed pyrimidines (Calvet and Pederson 1979; Cimino et al. 1985). This effectively staples the two base-pairing arms involved in each duplex together. In the second step, RNase S1 digestion is utilised to remove single-stranded regions of RNA. Subsequently, ShortCut RNase III is used to make duplexes smaller and complete proteinase digestion and RNA purification yield short, directly base-pairing duplexes. In the third step, 2D-electrophoresis is employed for purification and enrichment as cross-linked duplexes appear off-diagonal, corresponding to 0.2%–0.5% of the RNA used as input to the 2D electrophoresis. This step is likely to reduce the overall sensitivity. In the fourth step, the ends of these selected duplexes are proximity-ligated before the cross-linking of the duplexes is reversed using UV-light at 254 nm. The efficiency of the cross-ligation is key for ensuring that information on the base-pairing arms involved in one duplex is not lost. The ligation step concatenates the two arms involved in one duplex into an artificial RNA in which the linear ordering of the two arms is a priori not clear. Finally, pre-adenylated adapters are added to the 3’ ends, the resulting RNAs are reverse-transcribed in an adapter-specific way, circularised cDNA are generated and PCR amplification is performed to generate the cDNA libraries for NGS.
PARIS was originally performed in HeLa, HEK293T and mouse embryonic stem (mES) cells. Lu et al. conduct −AMT control experiments and observe no detectable off-diagonal elements in the corresponding 2D electrophoresis.
4.1.2 Experimental Protocol of SPLASH
The overall logical flow of SPLASH is similar to PARIS. Unlike for PARIS, cross-linking of duplexes in the first step is done using a biotinylated version of psoralen (so-called biopsoralen) also using UV-light at 365 nm. The biotin group is key for the subsequent enrichment step. Similar to AMT, biopsoralen also has a preference for cross-linking pyrimidines (Garrett-Wheeler et al. 1984; Hearst 1981). In contrast to AMT, however, biopsoralen typically requires the addition of a mild detergent (e.g. digitonin) to sufficiently increase the cellular uptake. The details of this (i.e. concentrations and duration of treatments with biopsoralen and digitonin) have to be carefully adjusted for each cell type separately. In the second step, cross-linked duplexes are extracted, randomly fragmented using Mg2+-mediated hydrolysis and biotin enriched using streptavidin magnetic beads. Note that due to the random nature of fragmentation procedure a nick can occur in the hybridised region. Therefore, there is a chance that the detected length of the duplex does not correspond to the full length of the original duplex. The enrichment step of SPLASH is thus experimentally more efficient and conceptually more straightforward than the enrichment step of PARIS involving the more loosely-defined off-diagonal in a 2D-electrophoresis. In the third step, the ends of the resulting duplexes are ligated before UV-light at 254 nm is used as in PARIS to reverse the cross-linking. Similarly to PARIS, the fourth step involves the addition of pre-adenylated adapters the 3’ ends, the reverse-transcription of the resulting RNAs in an adapter-specific way and the generation of circularised cDNAs. Again, PCR amplification is performed to obtain the final cDNA library for NGS.
The SPLASH protocol was used to examine HeLa cells, human lymphoblastoid cells, human embryonic stem (hES) cells, cells differentiated using retinoic acid and two types of cells from S. cerevisiae, namely wild type cells and Prp43 helicase mutant cells. Using between two to four biological replicates for each type of cell, they measure a high correlation (R = 0.75–0.9). Aw et al. (2016) generate several control libraries without cross-linking and without ligation in order to confirm that the duplexes identified by SPLASH are indeed enriched for ligated, cross-linked cases and not due to random background events. Furthermore, they explicitly confirm that cross-linking using biopsoralen is largely independent of solvent accessibility and show that SPLASH can detect RNA structure features with similar precision as the proximity ligation-based approach by Ramani et al. (2015) and has even higher sensitivity regarding trans RNA–RNA interactions.
4.1.3 Experimental Protocol of LIGR-seq
Conceptually, LIGR-seq has the same aims as PARIS and SPLASH, namely the direct detection of duplexes formed via RNA structure features or via trans RNA–RNA interactions. Unlike these two protocols, it uses a few features that set it distinctly apart and that have a significant impact on the subsequent computational interpretation of the raw reads.
Similar to PARIS, the first step of LIGR-seq consists of in vivo cross-linking of duplexes using AMT and UV-light at 365 nm. In terms of the specificity of the resulting, cross-linked duplexes, LIGR-seq is therefore comparable to PARIS (AMT) and SPLASH (biopsoralen). In the second step, RNA is extracted from cells and a limited digest with single-strand S1 endonuclease applied. The third step employs a circRNA ligase to link RNA ends in proximity. The fourth step is an enrichment step which utilises RNase R (a 3’-to’-5’ exoribonuclease) to digest linear and structured RNAs whose duplexes have not been cross-linked (Vincent and Deutscher 2006). The pool of surviving RNAs consists of fully circularised RNAs and linear RNAs with cross-linked duplexes (as well as linear RNAs with uncross-linked duplexes whose 3’ ends are too short for RNase R to latch on). Some false positives may very well survive the RNase R treatment. The fifth step reverses the cross-linking of duplexes using UV-light at 254 nm. Finally, the resulting RNAs (so-called chimeras in the LIGR-seq paper) are used to prepare stranded libraries for NGS. Unlike PARIS and SPLASH, the experimental protocol of LIGR-seq includes as default the preparation of an −AMT sample without any AMT-induced cross-linking. All samples are conceptually key for the subsequent computational interpretation of the raw LIGR-seq data. Without these, it would be conceptually impossible to define a dedicated probabilistic model which can assign estimated p-values to the experimentally detected interactions. Out of the three methods, LIGR-seq is currently the only method that is trying to experimentally estimate significance values for its detected interactions. As we will see in the following discussion of the computational analysis pipelines, it is also possible to assign significance values or p-values to proposed RNA structure features based on purely theoretical considerations, but these are conceptually different from the p-values derived by LIGR-seq.
4.1.4 Summary of All Three Experimental Protocols
After NGS, the raw data from PARIS, SPLASH and LIGR-seq corresponds to reads that each encode the sequence of the two arms involved in a formerly cross-linked duplex. One key difference with respect to chemical RNA structure probing methods is that any duplex can only be probed once as the molecules of the duplex itself end up being examined by the protocol. In contrast to this, methods for chemical RNA structure can probe any individual transcript multiple times and at different time points as they do not consume the investigated molecule itself.
For any given duplex derived by PARIS, SPLASH or LIGR-seq, it is unclear if the corresponding duplex derives from an inter- or from an intramolecular duplex, i.e. from a trans RNA–RNA interaction or from RNA structure features. It is also unclear in which linear order the two arms involved in the corresponding duplex appear in the resulting RNA and where their boundary is. These are key challenges to be addressed in the subsequent computational analysis of the raw data.
All three experimental protocols involve a stapler (i.e. AMT (PARIS and LIGR-seq) or biopsoralen (SPLASH)) that has a significant bias towards intercalating and cross-linking pyrimidines (Calvet and Pederson 1979; Cimino et al. 1985). Perfectly ordinary duplexes such as those involving G–C base pairs only may thus not be detectable at all using PARIS, SPLASH and LIGR-seq. Any absence of detectable duplexes can therefore not necessarily be taken as experimental evidence that the corresponding RNA structure feature of trans RNA–RNA interactions does not exist.
In addition, all three experimental protocols involve many steps that each introduce specific errors and biases that add up. As we will see in the following, the overall sensitivity and specificity of the combined step of each experimental protocol is further influenced by the errors and biases introduced by the computational analysis of the raw experimental data. It thus makes sense to consider and, ideally, optimise both in parallel.
4.2 Computational Protocols of PARIS, SPLASH and LIGR-seq
The main tasks of the computational analysis of the raw data from PARIS, SPLASH and LIGR-seq are (1) to map the sequenced reads back to the corresponding genome/transcriptome and (2) to figure out, for each read, if it corresponds to an inter- or an intramolecular duplex. Conceptually, both tasks have to be addressed simultaneously which amounts to the key challenge of the in silico analysis of these experimental data. In contrast to the sequenced reads derived from chemical RNA structure probing experiments, the raw data generated by PARIS, SPLASH and LIGR-seq do not correspond to a consecutive sub-sequence of any single transcript. Rather, each read either encodes the two separate of a duplex within the same transcript (if the duplex corresponds to an RNA structure feature) or a duplex involving two transcripts (if the duplex corresponds to a trans RNA–RNA interaction).
In case of an RNA structure duplex, mapping the corresponding read requires a gapped alignment to a single transcript (with a gap inserted between the two base-paired arms of the duplex encoded in the read) or a chimeric alignment in case of the two parts being non-canonical due to circle formation. This is complicated by the fact that the linear order of the arms in the read need not correspond to the natural linear order of the two arms within the underlying transcript (so-called chiastic reads). In case of a trans RNA–RNA duplex, mapping the read involves the identification of a pair of transcripts to which either of the two base-paired arms in the read map. This is conceptually and computationally challenging as the search space of all pairs of transcripts is huge compared to the search space of individual transcripts. Also here, the linear order in which the two arms appear in the read need not correspond to the order in which the respective two transcripts appear (chiastic reads). Furthermore, for both kinds of duplexes, the boundary between the two arms, i.e. where the gap has to be inserted for mapping, is a priori not known. To complicate matters further, it is up to the computational analysis to figure out for each read whether it corresponds to an RNA structure duplex or a trans RNA–RNA duplex.
The computational data analyses published in conjunction with the experimental protocols of PARIS, SPLASH and LIGR-seq have some main features in common, but differ in key details. As these differences are not exclusively due to the differences in experimental protocols, but partly due to different underlying strategies for interpreting the raw data, we will discuss them here.
4.2.1 Computational Analysis of Raw PARIS Data
Raw PARIS reads are first pre-processed by removing adapters from the 3’ ends and PCR duplicates. The latter is possible due to the insertion of a bar-code (random hexamer) in the middle of the adapter. These reads are then mapped to the corresponding genome using the computer program STAR (Dobin et al. 2013) with a set of input parameters that explicitly allow gapped-reads as well as so-called chiastic reads.
In a chiastic read, the linear order of the mappable parts (in our case, the two arms of a duplex) needs to be inverted. So, a read encoding a 5’-R-L-3’ duplex with a right (R) and left (L) arm of an RNA structure duplex needs to be mapped as 5’-L-3’-gap-5’-R-3’ to the underlying transcript. These chiastic reads naturally arise in all protocols whenever the ligation of a cross-linked, RNA structure-derived duplex happens to fuse the two base-pairing arms of the duplex in the wrong linear order, i.e. 5’-R-L-3’ rather than 5’-L-R-3’. Chiastic reads can also arise in duplexes corresponding to trans RNA–RNA interactions whenever the mapping of the 5’-R-L-3’ read to the (linearly ordered) transcripts of the transcriptome requires the reversal of the linear ordering of the two arms involved in the duplex. The correct mapping of chiastic reads thus always implies the insertion of a gap.
Before the mapping with STAR can actually be performed, a corresponding STAR index needs to be generated. This needs to be done with a carefully adjusted parameter for genomeSAindexNbases whenever the index is generated for a so-called mini-genome. The authors of PARIS utilise these mini-genomes in order to artificially reduce the search space for mapping, in particular when searching for specific trans RNA–RNA interactions, but also when investigating select genes in terms of RNA structure features (e.g. Xist gene or sub-set of snRNAs only). The parameters of STAR have to be explicitly adjusted whenever mini-genomes are used.
Of all the resulting STAR-mapped PARIS reads, only gapped and chiastic ones are retained. Of the gapped reads, only those are retained whose gap is not due to splicing.
In the next step, the retained mapped reads are grouped into so-called duplex groups (DGs). This is done using a greedy algorithm involving two steps. In the first step, the mapped reads are clustered into initial DGs such that all reads in a DG share at least 5 nt common overlap in both duplex arms (these two regions of overlap define the so-called core regions of the DG). Any mapped read is thereby either merged with an already existing DG or used to start a new DG. In the second step, DGs are merged into single DGs if they are close to each other and “well-defined” for both arms, see supplementary information of PARIS (Lu et al. 2016) for details.
Once the DGs have been established, each duplex group DG is assigned a so-called connection score which is defined as \(cs(DG) = N_{\mathrm{span}}(DG) / \sqrt {N_{\mathrm{left}}(DG) \cdot N_{\mathrm{right}}(DG)}\), where N span(DG) is the number of reads spanning the two duplex arms of DG and N left(DG) and N right(DG) are the number of unique reads overlapping the left and the right arm of DG, respectively. Note that N left(DG) can be different from N right(DG) as the reads covering each arm of the DG can also be assigned to other duplex groups overlapping DG only in one arm. Any duplex group DG with a connection score cs(DG) < 0.01 is then discarded to focus the subsequent analysis on duplexes that are supported by a significant portion of overlapping transcript reads.
The resulting duplexes typically involve two arms of 20–30 nt. The specific base pairs involved in a duplex between these two arms can, however, not be directly inferred from any DG. Rather, they have to be predicted based on the arms of the DG.
Lu et al. (2016) find that known miRNA-mRNA interactions cannot be detected, either because the duplex involved in the seed region is fairly short (around 5 nt length) and/or because binding of the duplex by the Argonaute protein shields the duplex from cross-linking.
Lu et al. try to assign a statistical significance to each detected duplex (whether corresponding to an RNA structure feature or a trans RNA–RNA interaction). For this, they compare the free energy of the MFE structure predicted for a multiple-sequence alignment underlying this DG to the corresponding, predicted free energies for 100 randomised versions of this multiple-sequence alignment. They thereby obtain a Z-score (Gesell and von Haeseler 2006). By utilising a procedure which focuses on the multiple-sequence alignment underlying the DG only, however, the Z-score cannot assess the statistical significance of seeing this DG by chance within the same transcript, let alone within the entire transcriptome which is what one would ideally like to know. Lu et al. evaluate the overall performance of PARIS by examining select RNAs (rRNA, snRNA, microRNA, telomerase RNA). This is done by visually comparing corresponding DGs to known features.
4.2.2 Computational Analysis of Raw SPLASH Data
Conceptually, the overall logical flow of the computational analysis of SPLASH is similar to the above for PARIS. Key details, however, differ and these turn out to be important.
To start with, transcriptomes for mapping purposes are generated by downloading the corresponding reference transcriptomes (taking the longest known isoform for each coding or non-coding gene as representative transcript) and by manually adding in select classes of non-coding genes. Any sequence duplicates from the joint set are then removed.
In the first step, the raw SPLASH paired-end reads are pre-processed by removing adapters and merging overlapping paired-end reads into corresponding single reads. In the next step, only these single merged reads are retained and mapped to the respective reference transcriptome using BWA MEM (version 0.7.12) (Li and Durbin 2010) using parameter -T 20 to lower the minimum length of mapped regions to 20 nt. These mapped reads are then post-processed by sorting them and converting them to BAM-format using samtools. Reads are then filtered for potential PCR duplicates by examining sets of reads with identical start coordinates and identical CIGAR strings and by retaining only the first read in each such set (Ramani et al. 2015).
In the original SPLASH analysis, the authors decide to deliberately focus their entire subsequent analysis on long-range features, i.e. RNA structure features and trans RNA–RNA interactions where the two arms involved in the corresponding duplex are far apart in terms of the underlying search space. Technically, this is achieved by retaining only split alignments more than 50 nt apart from the BAM-file of mapped reads. The authors of SPLASH then apply several measures to increase the quality of the retained, mapped reads. Reads with a mapping quality below 20 are discarded. In addition, ambiguously mapped reads and mapped reads with similarly scored second best hits are discarded (e.g. pseudo-genes). To lower the number of false positives, any read spanning a known splice-junction is removed using STAR (Dobin et al. 2013) to map splits reads from the transcriptome back to the corresponding genome. For this, reference sets of known splice junctions are assumed to be correct and complete.
The quoted overall sensitivity of SPLASH of 78% is based on its performance for the known RNA structure features of the 80S ribosome. The overall precision is reported to be 75%. In order to estimate the false discovery rate, independently cross-linked total RNAs from human yeast were pooled to prepare and analyse SPLASH libraries for any human-yeast interactions. Based on this strategy, SPLASH is reported to have a false discovery rate < 3.7%.
In order to assign a statistical significance or p-value to the interactions detected by SPLASH, the free energy of the pairwise interaction in the detected duplex is compared to the free energy of many shuffled randomised versions of the sequences underlying the same pairwise interaction. The randomisation procedure keeps the di-nucleotide content preserved. SPLASH thus employs the same strategy as PARIS for estimating p-values to its detected interactions (in PARIS, this is done by shuffling multiple-sequence alignments; in SPLASH this is done by randomising only the sequences involved in the duplex). Both procedures are based on the validity of the assumption that true interactions in vivo have a lower minimum-free energy than interactions between corresponding randomised version of the same sequences. This assumption, however, is generally not justified (Rivas and Eddy 2000). In any case, the resulting p-value could not be interpreted as the probability of observing a corresponding RNA structure duplex or trans RNA–RNA interaction feature by chance. For this, entire transcripts (in case of RNA structure features) or pairs of transcripts (in case of trans RNA–RNA interactions) would need to be examined.
This could, for example, be achieved using Transat (Wiebe and Meyer 2010), a fully probabilistic method that takes a multiple-sequence alignment and a corresponding evolutionary tree as input and detects evolutionarily conserved duplexes (so-called helices) in the input alignment. Any predicted helices are assigned a log-likelihood score as well as a p-value. This p-value corresponds to the chance of observing the duplex in the same transcript by chance.
4.2.3 Computational Analysis of Raw LIGR-seq Data
Raw LIGR-seq data consists of stranded, single-end reads. Similar to the above procedures for PARIS and SPLASH, these raw reads first need to be computationally post-processed before their actual interpretation in terms of biological contents can begin.
For this, LIGR-seq proposes a dedicated computational analysis pipeline called aligater consisting of several steps. Unlike PARIS and SPLASH, the pipeline comprises a dedicated probabilistic model which is used to estimate p-values for the detected interactions. The first step removes the random bar-codes from the 5’ ends. In the second step, these trimmed reads are mapped to the corresponding transcriptome using bowtie2 with a set of especially adjusted input parameters that aim to maximise sensitivity while keeping the computational run-time of the analysis reasonable. In the third step, these initial bowtie2 alignments in BAM-format are re-analysed such that blocks for each read are recursively chained into longer alignments in order to detect chimeras. This procedure can also handle circular ligation products and identifies the best path through the read. This step assigns a score to each chained alignment and is conceptually key for all of the subsequent analysis. The key corresponding input parameter for this procedure (the so-called chaining penalty) has to be carefully adjusted depending on the library quality as well as the specs of the specific class of transcripts being investigated. Reads with best-scoring chained alignments are then assigned an individual LIGQ score which retains detailed information on the corresponding alignments.
These LIGQ scores are subsequently used to carefully address several potential problems by either discarding or re-classifying chimeras. For example, artifacts due the mis-mapping of spliced transcript and of near-identical sequence duplicates (due to repeats, pseudo-genes or paralogues) are identified via near-identical matches to contiguous stretches of the underlying genome overlapping the ligation site and discarded. Other artifacts that incorrectly identify intra-molecular interactions as inter-molecular ones are re-classified based on corresponding supporting evidence. Overall, five different post-processing steps are executed, resulting in a strategy that re-classifies events rather than simply discard them and that aims for high sensitivity.
Another significant, conceptual difference of LIGR-seq with respect to the two other protocols, i.e. SPLASH and PARIS, is that it proposes an experimental strategy for estimating the statistical significance of the detected duplexes. This is achieved via a dedicated probabilistic model that judges the observed versus the expected ratios of chimeric reads. Each observed to expected ratio (i.e. OE+AMT or OE−AMT) corresponds to the corresponding experiments (i.e. + AMT or −AMT) with and without ligase. For this, separate + AMT and −AMT control experiments are performed without the ligation step in order to assess the expected background levels of spurious ligation events. The resulting LIGR-seq reads are then computationally processed as described above to detect interaction events (chimeras). Any pair of genes g x and g y is assigned a probability for spurious trans interactions P B(g x, g y) (using subscript B for background) which is assumed to only be a function of the respective relative whole gene abundance P(g x) of gene g x and P(g y) of gene g y, respectively. Mathematically, it corresponds to the probability of two independent draws from a multinomial distribution that is proportional to the relative abundance of each gene in the transcriptome. This defines their so-called null model.
The relative whole gene abundance for each gene g is measured in terms of reads per million without length adjustment (the RNase R treatment prevents this normalisation) and denoted RPM(g). So, P B(g x, g y) ∝ P(g x)P(g y) if x≠y and if g x and g y have experimentally confirmed interactions events. In contrast, P B(g x, g y) = 0 if x = y or if x≠y and no interactions between these two genes are detected. The normalised probability for spurious interactions between gene g x and g y, p B(g x, g y) is then written as (using P(g j) = RPM(g j)∕∑iRPM(g i)):
This null model assumes that the probability of a direct, spurious trans RNA–RNA interaction between two genes g x and g y in the transcriptome is only a function of the abundance of the relative whole gene abundance for each gene in the transcriptome. This model does not capture the primary sequence identity of each gene which is likely to also influence the probability of spurious trans RNA–RNA interactions. Assuming the validity of their null model, each experimentally detected interaction between genes g x and g y can then be assigned a p-value based on the number of observed reads k that are supporting it. This allows to explicitly filter for significant, AMT-induced interactions. Technically, this is achieved by first defining an enrichment score r AMT which is defined as the ratio between OE+AMT and OE−AMT, i.e. r AMT = OE+AMT∕OE−AMT. For real, AMT-induced interactions, we expect OE+AMT > OE−AMT and require r AMT > 1.1, more than 2 reads (k > 2), a p-value < α and an RPM of more than 10 in support. Similarly, interactions with r AMT < 0.9 (and more than 2 reads (k > 2), a p-value < α and an RPM of more than 10) are considered false positives and allow to explicitly estimate the false positive rate of the overall protocol. In addition, LIGR-seq utilises two biological replicates. These allow to assess the overall technical reproducibility of the protocol (Spearman Rho = 0.38, p < 8 ⋅ 10−6).
Overall, the false discovery rate of LIGR-seq is estimated to range between 4.4% for highly expressed transcripts (> 250 RPM) and 25% for sparsely expressed transcripts (> 10 RPM). These numbers can be viewed as worst-case estimates as some known, stable interactions can be detected in both +AMT and -AMT samples. The high sensitivity of LIGR-seq can be explicitly confirmed based on known interactions in select groups of genes, e.g. known RNA structure features in the 80S ribosome (Anger et al. 2013) and trans RNA–RNA interactions between the 28S and 5S rRNA.
Overall, LIGR-seq is the only of the three protocols for measuring RNA structure features and trans RNA–RNA interactions in vivo that tries to assign experimentally estimated significance values to the detected features. This is done by proposing an explicit null model and by utilising dedicated, experimentally determined control samples. As mentioned above, Transat (Wiebe and Meyer 2010) could be readily used to assign p-values to any experimentally determined duplexes in order to estimate their statistical significance in terms of the probability of seeing each duplex in the underlying transcript by chance.
5 Outlook
The last few years have seen an explosion of novel experimental and computational methods for determining RNA structures and trans RNA–RNA interactions in vivo. All experimental protocols require substantial computational strategies for analysing and for converting the raw experimental data into actual RNA structures or trans RNA–RNA interactions. Experimental and computational approaches are closely intertwined and therefore require simultaneous optimisation in order to optimise the overall performance.
Significant future improvements could be made in various ways.
First, we need to fully acknowledge the complexities of transcriptomes in vivo, in particular on the computational side of things. Any transcript in vivo may be long (long in this case meaning longer than 200 nt), may have various, unknown trans interaction partners (which may introduce RNA structure changes, e.g. Mazloomian and Meyer (2015)), may assume more than a single functional RNA structure or trans RNA–RNA interaction throughput its cellular life (e.g. Zhu and Meyer 2015; Lai et al. 2013) and, in particular, is unlikely to ever experience true thermodynamic equilibrium as a naked RNA. In particular for long RNAs such as coding transcripts, there is no reason to assume that they fold into a minimum-free energy structure spanning the entire transcript.
As advances in the field of ab initio RNA structure prediction showed, we may tackle this challenge best by employing a comparative strategy, i.e. by simply trying to identify RNA structure features or trans RNA–RNA interactions that have been conserved during well-chosen evolutionary times. Conceptually, this is currently the only way to detect the overall effects of various complexities in vivo without having to explicitly model them. Probabilistic methods are particularly well suited to seamlessly integrating experimental probing data into RNA structure predictions. In order for this line of research to flourish, we require gold-standard data sets of experimental probing data from different experimental probing protocols that examine the same in vivo situation using different methods. This needs, in particular, to include transcripts longer than 200 nt (see the captions of Tables 3 and 4 for the specs of the current data sets) from diverse biological classes of transcripts, not only short and non-coding RNAs that are known to contain global RNA structures spanning the entire transcript. There is, for example, by now ample evidence that short- and long-range RNA structure features are involved in regulating key cellular processes such as alternative splicing (Meyer and Miklos 2005; Raker et al. 2009; Pervouchine et al. 2012; Mazloomian and Meyer 2015). These gold-standard data sets thus have to be large and diverse enough to allow for parameter training as well as cross-evaluation procedures to avoid and evaluate potential issues due to over-fitting. The same applies to methods for predicting trans RNA–RNA interaction, where the currently assembled benchmark set (Lai and Meyer 2016) could be significantly increased, diversified and complemented by different kinds of experimental probing data.
On the experimental side of things, it would be beneficial to further reduce the inherent biases and limitation that the current methods have. PARIS, SPLASH and LIGR-seq are currently all based on psoralen-derivatives for cross-linking. This makes them blind to duplexes without juxtaposed pyrimidines. It would thus be great to remedy this by identifying intercalators that have complementary chemical specificities. The mapping of raw duplexes could be significantly facilitated by introducing artificial, known linker-sequences during the ligation of duplex-ends. Conceptually, another major step forward could be made by devising experimental protocols that are capable of detecting RNA structure diversity, i.e. cases where different copies of the same transcript engage in different RNA structures or trans RNA–RNA interactions in vivo. Right now, any RNA structure variation is mis-interpreted as noise when interpreting chemical RNA structure probing data. Using specific variants of SHAPE-MaP (Smola et al. 2015b) may be able to change this conceptually by allowing structure probing information from individual transcripts to be retained throughout the entire protocol. Overall, Smola et al. propose three strategies. The standard Randomer workflow which uses random primers and default fragmentation and library preparation for creating a map of SHAPE-induced mutations, see Fig. 2. Due the fragmentation procedure, probing information on entire transcripts is typically lost. They propose two other strategies for addressing this problem. One is to perform size selection on RNAs with short lengths (< 500 nt) in order to retain full probing information on their entire sequences. This will, however, ignore a large proportion of typical transcriptomes (the average length for human mRNAs is 2.7 kb). To specifically address transcripts longer than 500 nt, i.e. particular isoforms of one gene, the so-called Amplicon workflow can be applied. In that strategy, specific primers, unique to one isoform, can be used to amplify only a region of the transcript. Then, multiple non-overlapping regions can be sequenced similar to the Randomer strategy to produce isoform specific information. This experimental strategy should in particular allow us to gain conceptually novel biological insight into how long coding or non-coding transcripts in eukaryotic genes use RNA structure features as mechanisms of gene regulation at RNA level. In the long run, the most elegant way of retaining RNA structure information on entire individual transcripts would be to combine chemical RNA structure probing with single-molecule sequencing techniques. This, however, will require significant changes of the currently existing protocols.
Overview of the strategies recently proposed by SHAPE-MaP (Smola et al. 2015b). Shown here are two isoforms A and B of the same gene with partially overlapping sequences, where only one isoform assumes an RNA structure. Black ellipses correspond to the adducts produced by the SHAPE reagent. Black stars indicate mutations indicated during reverse transcription. The primer used in the Randomer workflow is shown in dark green. Region-specific primers of the Amplicon workflow are shown in orange and blue. The unpaired region that is paired in isoform A and unpaired in isoform B is highlighted by a red circle. The addition of SHAPE reagents to isoform B in combination with the Randomer workflow will produce a signal confirming that the region is unpaired. To confirm the presence of the RNA structure feature in isoform A, an alternative approach is required. This can be achieved with the Amplicon workflow using primers that are specific for a region in isoform A. This ensures that the adduct that is specific to isoform B is not amplified and thereby ignored
These are truly exciting times for in vivo transcriptome research, with many significant recent contributions both on the experimental and the computational side. Only by simultaneously optimising both experimental and computational procedures, however, will we be able to combine the best of both worlds. Both aspects currently come with a range of in-built assumptions and limitations. Questioning and, ideally, further reducing those will be key to discovering truly novel features
References
Anger A, Armache J, Berninghausen O, Habeck M, Subklewe M, Wilson D, Beckmann R (2013) Structures of the human and drosophila 80S ribosome. Nature 497(7447):80–85
Aultman K, Chang S (1982) Partial P1 nuclease digestion as a probe of tRNA structure. Eur J Biochem 124(3):471–476
Aw J, Shen Y, Wilm A, Sun M, Lim X, Boon K, Tapsin S, Chan Y, Tan C, Sim A, Zhang T, Susanto T, Fu Z, Nagarajan N, Wan Y (2016) In vivo mapping of eukaryotic RNA interactomes reveals principles of Higher-Order organization and regulation. Mol Cell 62(4):603–617
Bevilacqua P, Ritchey L, Su Z, Assmann S (2016) Genome-Wide analysis of RNA secondary structure. Annu Rev Genet 50:235–266
Calvet J, Pederson T (1979) Heterogeneous nuclear RNA double-stranded regions probed in living HeLa cells by crosslinking with the psoralen derivative aminomethyltrioxsalen. Proc Natl Acad Sci USA 76(2):755–759
Cheng C, Chou F, Kladwang W, Tian S, Cordero P, Das R (2015) Consistent global structures of complex RNA states through multidimensional chemical mapping. Elife 4:e07600
Cimino G, Gamper H, Isaacs S, Hearst J (1985) Psoralens as photoactive probes of nucleic acid structure and function: organic chemistry, photochemistry, and biochemistry. Annu Rev Biochem 54:1151–1193
Cordero P, Kladwang W, VanLang C, Das R (2012a) Quantitative dimethyl sulfate mapping for automated RNA secondary structure inference. Biochemistry 51(36):7037–7039
Cordero P, Lucks J, Das R (2012b) An RNA mapping DataBase for curating RNA structure mapping experiments. Bioinformatics 28(22):3006–3008
Deigan K, Li T, Mathews D, Weeks K (2009) Accurate SHAPE-directed RNA structure determination. Proc Natl Acad Sci USA 106(1):97–102
Del Campo C, Bartholomäus A, Fedyunin I, Ignatova Z (2015) Secondary structure across the bacterial transcriptome reveals versatile roles in mRNA regulation and function. PLoS Genet 11(10):e1005613
Ding Y, Lawrence C (2003) A statistical sampling algorithm for RNA secondary structure prediction. Nucleic Acids Res 31(24):7280–7301
Ding Y, Tang Y, Kwok C, Zhang Y, Bevilacqua P, Assmann S (2014) In vivo genome-wide profiling of RNA secondary structure reveals novel regulatory features. Nature 505(7485):696–700
Ding Y, Kwok C, Tang Y, Bevilacqua P, Assmann S (2015) Genome-wide profiling of in vivo RNA structure at single-nucleotide resolution using structure-seq. Nat Protoc 10(7):1050–1066
Dobin A, Davis C, Schlesinger F, Drenkow J, Zaleski C, Jha S, Batut P, Chaisson M, Gingeras T (2013) STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29(1):15–21
Durbin R, Eddy S, Krogh A, Mitchison G (1998) Biological sequence analysis: probabilistic models of proteins and nucleic acids. Cambridge University Press, Cambridge
Eddy S (2014) Computational analysis of conserved RNA secondary structure in transcriptomes and genomes. Annu Rev Biophys 43:433–456
Ehresmann C, Baudin F, Mougel M, Romby P, Ebel J, Ehresmann B (1987) Probing the structure of RNAs in solution. Nucleic Acids Res 15(22):9109–9128
Fang R, Moss W, Rutenberg-Schoenberg M, Simon M (2015) Probing Xist RNA structure in cells using targeted Structure-Seq. PLoS Genet 11(12):e1005668
Flynn R, Zhang Q, Spitale R, Lee B, Mumbach M, Chang H (2016) Transcriptome-wide interrogation of RNA secondary structure in living cells with icSHAPE. Nat Protoc 11(2): 273–290
Garrett-Wheeler E, Lockard R, Kumar A (1984) Mapping of psoralen cross-linked nucleotides in RNA. Nucleic Acids Res 12(7):3405–3423
Gesell T, von Haeseler A (2006) In silico sequence evolution with site-specific interactions along phylogenetic trees. Bioinformatics 22(6):716–722
Guerrier-Takada C, Gardiner K, Marsh T, Pace N, Altman S (1983) The RNA moiety of ribonuclease P is the catalytic subunit of the enzyme. Cell 35(3):849–857
Harris K Jr, Crothers D, Ullu E (1995) In vivo structural analysis of spliced leader RNAs in trypanosoma brucei and leptomonas collosoma: a flexible structure that is independent of cap4 methylations. RNA 1(4):351–362
Hearst J (1981) Psoralen photochemistry and nucleic acid structure. J Invest Dermatol 77(1):39–44
Hector R, Burlacu E, Aitken S, Le Bihan T, Tuijtel M, Zaplatina A, Cook A, Granneman S (2014) Snapshots of pre-rRNA structural flexibility reveal eukaryotic 40S assembly dynamics at nucleotide resolution. Nucleic Acids Res 42(19):12138–12154
Higgs PG (2000) RNA secondary structure: physical and computational aspects. Q Rev Biophys 33(3):199–253
Homan P, Favorov O, Lavender C, Kursun O, Ge X, Busan S, Dokholyan N, Weeks K (2014) Single-molecule correlated chemical probing of RNA. Proc Natl Acad Sci USA 111(38):13858–13863
Incarnato D, Neri F, Anselmi F, Oliviero S (2014) Genome-wide profiling of mouse RNA secondary structures reveals key features of the mammalian transcriptome. Genome Biol 15(10):491
Kertesz M, Wan Y, Mazor E, Rinn J, Nutter R, Chang H, Segal E (2010) Genome-wide measurement of RNA secondary structure in yeast. Nature 467(7311):103–107
Kielpinski L, Vinther J (2014) Massive parallel-sequencing-based hydroxyl radical probing of RNA accessibility. Nucleic Acids Res 42(8):e70
Knapp G (1989) Enzymatic approaches to probing of RNA secondary and tertiary structure. Methods Enzymol 180:192–212
Knudsen B, Hein J (2003) Pfold: RNA secondary structure prediction using stochastic context-free grammars. Nucleic Acids Res 31(13):3423–3428
Koller D, Friedman N (2009) Probabilstic graphical models. MIT, Cambridge
Kwok C, Ding Y, Tang Y, Assmann S, Bevilacqua P (2013) Determination of in vivo RNA structure in low-abundance transcripts. Nat Commun 4:2971
Lai D, Meyer IM (2016) A comprehensive comparison of general RNA-RNA interaction prediction methods. Nucleic Acids Res 44(7):e61
Lai D, Proctor JR, Meyer IM (2013) On the importance of cotranscriptional RNA structure formation. RNA 19(11):1461–1473
Latham J, Cech T (1989) Defining the inside and outside of a catalytic RNA molecule. Science 245(4915):276–282
Lavender C, Gorelick R, Weeks K (2015) Structure-based alignment and consensus secondary structures for three HIV-related RNA genomes. PLoS Comput Biol 11(5):e1004230
Lengyel J, Hnath E, Storms M, Wohlfarth T (2014) Towards an integrative structural biology approach: combining Cryo-TEM, X-ray crystallography, and NMR. J Struct Funct Genom 15(3):117–124
Li H, Durbin R (2010) Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26(5):589–595
Li F, Zheng Q, Ryvkin P, Dragomir I, Desai Y, Aiyer S, Valladares O, Yang J, Bambina S, Sabin L, Murray J, Lamitina T, Raj A, Cherry S, Wang L, Gregory B (2012a) Global analysis of RNA secondary structure in two metazoans. Cell Rep 1(1):69–82
Li F, Zheng Q, Vandivier L, Willmann M, Chen Y, Gregory B (2012b) Regulatory impact of RNA secondary structure across the arabidopsis transcriptome. Plant Cell 24(11):4346–4359
Lorenz R, Bernhart S, Höner Zu Siederdissen C, Tafer H, Flamm C, Stadler P, Hofacker I (2011) ViennaRNA package 2.0. Algorithms Mol Biol 6:26
Lorenz R, Luntzer D, Hofacker I, Stadler P, Wolfinger M (2016) SHAPE directed RNA folding. Bioinformatics 32(1):145–147
Loughrey D, Watters K, Settle A, Lucks J (2014) SHAPE-Seq 2.0: systematic optimization and extension of high-throughput chemical probing of RNA secondary structure with next generation sequencing. Nucleic Acids Res 42(21):e165
Low J, Garcia-Miranda P, Mouzakis K, Gorelick R, Butcher S, Weeks K (2014) Structure and dynamics of the HIV-1 frameshift element RNA. Biochemistry 53(26):4282–4291
Lu Z, Zhang Q, Lee B, Flynn R, Smith M, Robinson J, Davidovich C, Gooding A, Goodrich K, Mattick J, Mesirov J, Cech T, Chang H (2016) RNA duplex map in living cells reveals Higher-Order transcriptome structure. Cell 165(5):1267–1279
Lucks J, Mortimer S, Trapnell C, Luo S, Aviran S, Schroth G, Pachter L, Doudna J, Arkin A (2011) Multiplexed RNA structure characterization with selective 2’-hydroxyl acylation analyzed by primer extension sequencing (shape-seq). Proc Natl Acad Sci USA 108(27):11063–11068
Mathews DH, Sabina J, Zuker M, Turner DH (1999) Expanded sequence dependence of thermodynamic parameters improves prediction of RNA secondary structure. J Mol Biol 288(5):911–940
Mathews D, Disney M, Childs J, Schroeder S, Zuker M, Turner D (2004) Incorporating chemical modification constraints into a dynamic programming algorithm for prediction of RNA secondary structure. Proc Natl Acad Sci USA 101(19):7287–7292
Mauger D, Golden M, Yamane D, Williford S, Lemon S, Martin D, Weeks K (2015) Functionally conserved architecture of hepatitis C virus RNA genomes. Proc Natl Acad Sci USA 112(12):3692–3697
Mazloomian A, Meyer IM (2015) Genome-wide identification and characterization of tissue-specific RNA editing events in D. melanogaster and their potential role in regulating alternative splicing. RNA Biol 12(12):1391–1401
McCaskill J (1990) The equilibrium partition function and base pair binding probabilities for RNA secondary structure. Biopolymers 29(6–7):1105–1119
McGinnis J, Dunkle J, Cate J, Weeks K (2012) The mechanisms of RNA SHAPE chemistry. J Am Chem Soc 134(15):6617–6624
Merino E, Wilkinson K, Coughlan J, Weeks K (2005) RNA structure analysis at single nucleotide resolution by selective 2’-hydroxyl acylation and primer extension (shape). J Am Chem Soc 127(12):4223–4231
Meyer I (2017) In silico methods for co-transcriptional RNA secondary structure prediction and for investigating alternative RNA structure expression. Methods 120:3–16
Meyer I, Miklos I (2004) Co-transcriptional folding is encoded within RNA genes. BMC Mol Biol 5:10
Meyer I, Miklos I (2005) Statistical evidence for conserved, local secondary structure in the coding regions of eukaryotic mRNAs and pre-mRNAs. Nucleic Acids Res 33(19):6338–6348
Meyer IM, Miklos I (2007) SimulFold: simultaneously inferring RNA structures including pseudoknots, alignments, and trees using a Bayesian MCMC framework. PLOS Comput Biol 3(8):1441–1454
Miklos I, Meyer I, Nagy B (2005) Moments of the Boltzmann distribution for RNA secondary structures. Bull Math Biol 67(5):1031–1047
Moazed D, Stern S, Noller H (1986) Rapid chemical probing of conformation in 16 S ribosomal RNA and 30 S ribosomal subunits using primer extension. J Mol Biol 187(3):399–416
Morgan S, Higgs PG (1996) Evidence for kinetic effects in the folding of large RNA molecules. Annu Rev Biophys 105:7152–7157
Mortimer S, Weeks K (2007) A fast-acting reagent for accurate analysis of RNA secondary and tertiary structure by SHAPE chemistry. J Am Chem Soc 129(14):4144–4145
Mortimer S, Trapnell C, Aviran S, Pachter L, Lucks J (2012) Shape-seq: high-throughput RNA structure analysis. Curr Protoc Chem Biol 4(4):275–297
Ouyang Z, Snyder M, Chang H (2013) SeqFold: genome-scale reconstruction of RNA secondary structure integrating high-throughput sequencing data. Genome Res 23(2):377–387
Pedersen J, Forsberg R, Meyer I, Hein J (2004a) An evolutionary model for protein-coding regions with conserved RNA structure. Mol Biol Evol 21(10):1913–1922
Pedersen J, Meyer I, Forsberg R, Simmonds P, Hein J (2004b) A comparative method for finding and folding RNA secondary structures within protein-coding regions. Nucleic Acids Res 32(16):4925–4936
Pervouchine DD, Khrameeva EE, Pichugina MY, Nikolaienko OV, Gelfand MS, Rubtsov PM, Mironov AA (2012) Evidence for widespread association of mammalian splicing and conserved long-range RNA structures. RNA 18(1):1–15
Poulsen L, Kielpinski L, Salama S, Krogh A, Vinther J (2015) SHAPE selection (shapes) enrich for RNA structure signal in SHAPE sequencing-based probing data. RNA 21(5):1042–1052
Proctor JR, Meyer IM (2013) CoFold: an RNA secondary structure prediction method that takes co-transcriptional folding into account. Nucleic Acids Res 41(9):e102
Qi L, Lucks J, Liu C, Mutalik V, Arkin A (2012) Engineering naturally occurring trans-acting non-coding RNAs to sense molecular signals. Nucleic Acids Res 40(12):5775–5786
Quarrier S, Martin J, Davis-Neulander L, Beauregard A, Laederach A (2010) Evaluation of the information content of RNA structure mapping data for secondary structure prediction. RNA 16(6):1108–1117
Raker VA, Mironov AA, Gelfand MS, Pervouchine DD (2009) Modulation of alternative splicing by long-range RNA structures in Drosophila. Nucleic Acids Res 37(14):4533–4544
Ramani V, Qiu R, Shendure J (2015) High-throughput determination of RNA structure by proximity ligation. Nat Biotechnol 33(9):980–984
Rice G, Leonard C, Weeks K (2014) RNA secondary structure modeling at consistent high accuracy using differential SHAPE. RNA 20(6):846–854
Righetti F, Nuss A, Twittenhoff C, Beele S, Urban K, Will S, Bernhart S, Stadler P, Dersch P, Narberhaus F (2016) Temperature-responsive in vitro RNA structurome of yersinia pseudotuberculosis. Proc Natl Acad Sci USA 113(26):7237–7242
Rivas E, Eddy S (2000) Secondary structure alone is generally not statistically significant for the detection of noncoding RNAs. Bioinformatics 16(7):583–605
Rouskin S, Zubradt M, Washietl S, Kellis M, Weissman J (2014) Genome-wide probing of RNA structure reveals active unfolding of mRNA structures in vivo. Nature 505(7485):701–705
Sahoo S, Świtnicki M, Pedersen J (2016) ProbFold: a probabilistic method for integration of probing data in RNA secondary structure prediction. Bioinformatics 32(17):2626–2635
Sclavi B, Woodson S, Sullivan M, Chance M, Brenowitz M (1997) Time-resolved synchrotron x-ray “footprinting”, a new approach to the study of nucleic acid structure and function: application to protein-DNA interactions and RNA folding. J Mol Biol 266(1):144–159
Seetin M, Kladwang W, Bida J, Das R (2014) Massively parallel RNA chemical mapping with a reduced bias MAP-seq protocol. Methods Mol Biol 1086:95–117
Sharma E, Sterne-Weiler T, O’Hanlon D, Blencowe B (2016) Global mapping of human RNA-RNA interactions. Mol Cell 62(4):618–626
Siegfried N, Busan S, Rice G, Nelson J, Weeks K (2014) RNA motif discovery by SHAPE and mutational profiling (shape-map). Nat Methods 11(9):959–965
Smola M, Calabrese J, Weeks K (2015a) Detection of RNA-protein interactions in living cells with SHAPE. Biochemistry 54(46):6867–6875
Smola M, Rice G, Busan S, Siegfried N, Weeks K (2015b) Selective 2’-hydroxyl acylation analyzed by primer extension and mutational profiling (shape-map) for direct, versatile and accurate RNA structure analysis. Nat Protoc 10(11):1643–1669
Soper SFC, Dator RP, Limbach PA, Woodson SA (2013) In vivo x-ray footprinting of pre-30s ribosomes reveals chaperone-dependent remodeling of late assembly intermediates. Mol Cell 52(4):506–516
Spitale R, Flynn R, Zhang Q, Crisalli P, Lee B, Jung J, Kuchelmeister H, Batista P, Torre E, Kool E, Chang H (2015) Structural imprints in vivo decode RNA regulatory mechanisms. Nature 519(7544):486–490
Steen K, Rice G, Weeks K (2012) Fingerprinting noncanonical and tertiary RNA structures by differential SHAPE reactivity. J Am Chem Soc 134(32):13160–13163
Steif A, Meyer IM (2012) The hok mRNA family. RNA Biol 9(12):1399–1404
Sükösd Z, Knudsen B, Kjems J, Pedersen C (2012) PPfold 3.0: fast RNA secondary structure prediction using phylogeny and auxiliary data. Bioinformatics 28(20):2691–2692
Swenson MS, Anderson J, Ash A, Gaurav P, Sükösd Z, Bader DA, Harvey SC, Heitsch CE (2012) GTfold: enabling parallel RNA secondary structure prediction on multi-core desktops. BMC Res Notes 5:341
Talkish J, May G, Lin Y, Woolford J Jr, McManus C (2014) Mod-seq: high-throughput sequencing for chemical probing of RNA structure. RNA 20(5):713–20
Tyrrell J, McGinnis J, Weeks K, Pielak G (2013) The cellular environment stabilizes adenine riboswitch RNA structure. Biochemistry 52(48):8777–8785
Underwood J, Uzilov A, Katzman S, Onodera C, Mainzer J, Mathews D, Lowe T, Salama S, Haussler D (2010) FragSeq: transcriptome-wide RNA structure probing using high-throughput sequencing. Nat Methods 7(12):995–1001
Vincent H, Deutscher M (2006) Substrate recognition and catalysis by the exoribonuclease RNase R. J Biol Chem 281(40):29769–29775
Wan Y, Qu K, Ouyang Z, Kertesz M, Li J, Tibshirani R, Makino D, Nutter R, Segal E, Chang H (2012) Genome-wide measurement of RNA folding energies. Mol Cell 48(2):169–181
Wan Y, Qu K, Ouyang Z, Chang H (2013) Genome-wide mapping of RNA structure using nuclease digestion and high-throughput sequencing. Nat Protoc 8(5):849–869
Wan Y, Qu K, Zhang Q, Flynn R, Manor O, Ouyang Z, Zhang J, Spitale R, Snyder M, Segal E, Chang H (2014) Landscape and variation of RNA secondary structure across the human transcriptome. Nature 505(7485):706–709. https://doi.org/10.1038/nature12946
Washietl S, Hofacker I, Stadler P, Kellis M (2012) RNA folding with soft constraints: reconciliation of probing data and thermodynamic secondary structure prediction. Nucleic Acids Res 40(10):4261–4272
Watters K, Abbott T, Lucks J (2016a) Simultaneous characterization of cellular RNA structure and function with in-cell SHAPE-Seq. Nucleic Acids Res 44(2):e12
Watters K, Yu A, Strobel E, Settle A, Lucks J (2016b) Characterizing RNA structures in vitro and in vivo with selective 2’-hydroxyl acylation analyzed by primer extension sequencing (shape-seq). Methods 103:34–48
Watts J, Dang K, Gorelick R, Leonard C, Bess J Jr, Swanstrom R, Burch C, Weeks K (2009) Architecture and secondary structure of an entire HIV-1 RNA genome. Nature 460(7256): 711–716
Weeks K (2010) Advances in RNA structure analysis by chemical probing. Curr Opin Struct Biol 20(3):295–304
Wells S, Hughes J, Igel A, Ares M Jr (2000) Use of dimethyl sulfate to probe RNA structure in vivo. Methods Enzymol 318:479–493
Wiebe NJP, Meyer IM (2010) TRANSAT-a method for detecting the conserved helices of functional RNA structures, including transient, pseudo-knotted and alternative structures. PLOS Comput Biol 6(6):e1000823
Wilkinson K, Merino E, Weeks K (2006) Selective 2’-hydroxyl acylation analyzed by primer extension (shape): quantitative RNA structure analysis at single nucleotide resolution. Nat Protoc 1(3):1610–1616
Woese C, Magrum L, Gupta R, Siegel R, Stahl D, Kop J, Crawford N, Brosius J, Gutell R, Hogan J, Noller H (1980) Secondary structure model for bacterial 16S ribosomal RNA: phylogenetic, enzymatic and chemical evidence. Nucleic Acids Res 8(10):2275–2293
Zarringhalam K, Meyer M, Dotu I, Chuang J, Clote P (2012) Integrating chemical footprinting data into RNA secondary structure prediction. PLoS One 7(10):e45160
Zaug A, Cech T (1995) Analysis of the structure of tetrahymena nuclear RNAs in vivo: telomerase RNA, the self-splicing rRNA intron, and U2 snRNA. RNA 1(4):363–374
Zheng Q, Ryvkin P, Li F, Dragomir I, Valladares O, Yang J, Cao K, Wang L, Gregory B (2010) Genome-wide double-stranded RNA sequencing reveals the functional significance of base-paired RNAs in arabidopsis. PLoS Genet 6(9):e1001141
Zhu JYA, Meyer IM (2015) Four RNA families with functional transient structures. RNA Biol 12(1):5–20
Zhu JYA, Steif A, Proctor JR, Meyer IM (2013) Transient RNA structure features are evolutionarily conserved and can be computationally predicted. Nucleic Acids Res 41(12):6273–6285
Zuker M (2003) Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res 31(13):3406–3415
Zuker M, Stiegler P (1981) Optimal computer folding of large RNA sequences using thermodynamics and auxiliary information. Nucleic Acids Res 9(1):133–148
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made. The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2018 Springer International Publishing AG, part of Springer Nature
About this chapter
Cite this chapter
Stefanov, S.R., Meyer, I.M. (2018). Deciphering the Universe of RNA Structures and trans RNA–RNA Interactions of Transcriptomes In Vivo: From Experimental Protocols to Computational Analyses. In: Rajewsky, N., Jurga, S., Barciszewski, J. (eds) Systems Biology. RNA Technologies. Springer, Cham. https://doi.org/10.1007/978-3-319-92967-5_9
Download citation
DOI: https://doi.org/10.1007/978-3-319-92967-5_9
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-92966-8
Online ISBN: 978-3-319-92967-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)