Abstract
The three main linear phenotypic eigen selection index methods are the eigen selection index method (ESIM), the restricted ESIM (RESIM) and the predetermined proportional gain ESIM (PPG-ESIM). The ESIM is an unrestricted index, but the RESIM and PPG-ESIM allow null and predetermined restrictions respectively to be imposed on the expected genetic gains of some traits, whereas the rest remain without any restrictions. These indices are based on the canonical correlation, on the singular value decomposition, and on the linear phenotypic selection indices theory. We extended the ESIM theory to the molecular-assisted and genomic selection context to develop a molecular ESIM (MESIM), a genomic ESIM (GESIM), and a genome-wide ESIM (GW-ESIM). Also, we extend the RESIM and PPG-ESIM theory to the restricted genomic ESIM (RGESIM), and to the predetermined proportional gain genomic ESIM (PPG-GESIM) respectively. The latter five indices use marker and phenotypic information jointly to predict the net genetic merit of the candidates for selection, but although MESIM uses only statistically significant markers linked to quantitative trait loci, the GW-ESIM uses all genome markers and phenotypic information and the GESIM, RGESIM, and PPG-GESIM use the genomic estimated breeding values and the phenotypic values to predict the net genetic merit. Using real and simulated data, we validated the theoretical results of all five indices.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
8.1 The Molecular Eigen Selection Index Method
The molecular eigen selection index method (MESIM) is very similar to the linear molecular selection index (LMSI) described in Chap. 4; thus, it uses the same set of information to predict the net genetic merit of individual candidates for selection, and therefore needs the same set of conditions as those of the LMSI. The only difference between the two indices is how the vector of coefficients is obtained and the assumption associated with the vector of economic weights. Thus, although the LMSI obtains the vector of coefficients according to the linear phenotypic selection index (LPSI) described in Chap. 2 and assumes that the economic weights are known and fixed, the MESIM assumes that the economic weights are unknown and fixed and obtains the vector of coefficients according to the ESIM theory.
8.1.1 The MESIM Parameters
In the MESIM context, the net genetic merit can be written as
where \( {\mathbf{g}}^{\prime }=\left[{g}_1\kern0.5em \dots \kern0.5em {g}_t\right] \) is the vector of true breeding values, t is the number of traits, \( {\mathbf{w}}_1^{\prime }=\left[{w}_1\kern0.5em \cdots \kern0.5em {w}_t\right] \) is a vector of unknown economic weights associated with g, \( {\mathbf{w}}_2^{\prime }=\left[{0}_1\kern0.5em \cdots \kern0.5em {0}_t\right] \) is a null vector associated with the vector of marker score values \( {\mathbf{s}}^{\prime }=\left[{s}_1\kern0.5em {s}_2\kern0.5em \dots \kern0.5em {s}_t\right] \), \( {\mathbf{w}}^{\prime }=\left[{\mathbf{w}}_1^{\prime}\kern0.5em {\mathbf{w}}_2^{\prime}\right] \) and \( {\mathbf{a}}^{\prime }=\left[{\mathbf{g}}^{\prime}\kern0.5em {\mathbf{s}}^{\prime}\right] \) (Chap. 4 for details). The MESIM index can be written as
where \( {\mathbf{y}}^{\prime }=\left[{y}_1\kern0.5em \cdots \kern0.5em {y}_t\right] \) is the vector of phenotypic values; \( {\mathbf{s}}^{\prime }=\left[{s}_1\kern0.5em {s}_2\kern0.5em \dots \kern0.5em {s}_t\right] \) is the vector of marker scores; \( {\boldsymbol{\upbeta}}_{\mathbf{y}}^{\prime } \) and βs are vectors of phenotypic and marker score weight values respectively, \( {\boldsymbol{\upbeta}}^{\prime }=\left[{\boldsymbol{\upbeta}}_{\mathbf{y}}^{\prime}\kern0.5em {\boldsymbol{\upbeta}}_G^{\prime}\right] \) and \( {\mathbf{t}}^{\prime }=\left[{\mathbf{y}}^{\prime}\kern0.5em {\mathbf{s}}^{\prime}\right] \). The objectives of the MESIM are the same as those of the ESIM (see Chap. 7 for details).
Let \( Var(H)={\mathbf{w}}^{\prime }{\boldsymbol{\Psi}}_M\mathbf{w}={\sigma}_H^2 \) be the variance of H, \( Var(I)={\boldsymbol{\upbeta}}^{\prime }{\mathbf{T}}_M\boldsymbol{\upbeta} ={\sigma}_I^2 \) the variance of I, and Cov(H, I) = w′ΨMβ the covariance between H and I, where \( {\boldsymbol{\Psi}}_M= Var\left[\begin{array}{c}\mathbf{g}\\ {}\mathbf{s}\end{array}\right]=\left[\begin{array}{cc}\mathbf{C}& {\mathbf{S}}_M\\ {}{\mathbf{S}}_M& {\mathbf{S}}_M\end{array}\right] \) and \( {\mathbf{T}}_M= Var\left[\begin{array}{c}\mathbf{y}\\ {}\mathbf{s}\end{array}\right]=\left[\begin{array}{cc}\mathbf{P}& {\mathbf{S}}_M\\ {}{\mathbf{S}}_M& {\mathbf{S}}_M\end{array}\right] \) are block matrices of size 2t × 2t (t is the number of traits) of covariance matrices where P, SM, and C are covariance matrices t × t of phenotypic (y), marker score (s), and genetic breeding (g) values respectively. Let \( {\rho}_{HI}=\frac{{\mathbf{w}}^{\prime }{\boldsymbol{\Psi}}_M\boldsymbol{\upbeta}}{\sqrt{{\mathbf{w}}^{\prime }{\boldsymbol{\Psi}}_M\mathbf{w}}\sqrt{{\boldsymbol{\upbeta}}^{\prime }{\mathbf{T}}_M\boldsymbol{\upbeta}}} \) and \( {h}_I^2=\frac{{\boldsymbol{\upbeta}}^{\prime }{\boldsymbol{\Psi}}_M\boldsymbol{\upbeta}}{{\boldsymbol{\upbeta}}^{\prime }{\mathbf{T}}_M\boldsymbol{\upbeta}} \) be the correlation between H and I, and the heritability of I respectively; then, the MESIM selection response can be written as
and
where kI is the standardized selection differential (or selection intensity) associated with MESIM; \( {\sigma}_H=\sqrt{{\mathbf{w}}^{\prime }{\boldsymbol{\Psi}}_M\mathbf{w}} \) and \( {\sigma}_I=\sqrt{{\boldsymbol{\upbeta}}^{\prime }{\mathbf{T}}_M\boldsymbol{\upbeta}} \) are the standard deviations of the variance of H and I respectively. It is assumed that kI is fixed, and that matrices TM and ΨM are known; therefore, we can maximize R by maximizing ρHI (Eq. 8.3) with respect to vectors w and β, or by maximizing \( {h}_I^2 \) (Eq. 8.4) only with respect to vector β.
Maximizing \( {h}_I^2 \) only with respect to β is simpler than maximizing ρHI with respect to w and β; however, in the latter case the maximization process of ρHI gives more information associated with MESIM parameters than when \( {h}_I^2 \) is maximized only with respect to β (see Chap. 7, Eq. 7.13, for details). In this subsection, we maximize ρHI with respect to vectors w and β similar to the ESIM in Chap. 7, Sect. 7.1.1. Thus, we omit the steps and details of the maximization process of ρHI.
We maximize \( {\rho}_{HI}=\frac{{\mathbf{w}}^{\prime }{\boldsymbol{\Psi}}_M\boldsymbol{\upbeta}}{\sqrt{{\mathbf{w}}^{\prime }{\boldsymbol{\Psi}}_M\mathbf{w}}\sqrt{{\boldsymbol{\upbeta}}^{\prime }{\mathbf{T}}_M\boldsymbol{\upbeta}}} \) with respect to vectors w and β under the restrictions \( {\sigma}_H^2={\mathbf{w}}^{\prime}\boldsymbol{\Psi} \mathbf{w} \), \( {\sigma}_I^2={\boldsymbol{\upbeta}}^{\prime}\mathbf{T}\boldsymbol{\upbeta } \), and 0 < \( {\sigma}_H^2 \), \( {\sigma}_I^2 \) < ∞, where \( {\sigma}_H^2 \) is the variance of H = w′a and \( {\sigma}_I^2 \) is the variance of I = β′t. Thus, it is necessary to maximize the function
with respect to β, w, μ, and ϕ, where μ and ϕ are Lagrange multipliers. The derivatives of Eq. (8.5) with respect to β, w, μ, and ϕ are:
respectively, where Eq. (8.8) denotes the restrictions imposed for maximizing ρHI. It can be shown (see Chap. 7) that vector w can be obtained as
and the net genetic merit in the MESIM context can be written as \( {H}_M={\mathbf{w}}_M^{\prime}\mathbf{a} \); thus, the correlation between \( {H}_M={\mathbf{w}}_M^{\prime}\mathbf{a} \) and I is \( {\rho}_{H_MI}=\frac{\sqrt{{\boldsymbol{\upbeta}}^{\prime}\mathbf{T}\boldsymbol{\upbeta }}}{\sqrt{{\boldsymbol{\upbeta}}^{\prime }{\mathbf{T}\boldsymbol{\Psi}}^{-1}\mathbf{T}\boldsymbol{\upbeta }}} \) and the MESIM vector of coefficients (β) that maximizes \( {\rho}_{H_MI} \) can be obtained from equation
where I2t is an identity matrix of size 2t × 2t (t is the number of traits), and \( {\lambda}_M^2 \) and βM are the eigenvalue and eigenvector of matrix \( {\mathbf{T}}_M^{-1}{\boldsymbol{\Psi}}_M \). The words eigenvalue and eigenvector are derived from the German word eigen, which means owned by or peculiar to. Eigenvalues and eigenvectors are sometimes called characteristic values and characteristic vectors, proper values and proper vectors, or latent values and latent vectors (Meyer 2000). The square root of \( {\lambda}_M^2 \) (λM) is the canonical correlation between \( {H}_M={\mathbf{w}}_M^{\prime}\mathbf{a} \) and \( {I}_M={\boldsymbol{\upbeta}}_M^{\prime}\mathbf{t} \), and the optimized MESIM index can be written as \( {I}_M={\boldsymbol{\upbeta}}_M^{\prime}\mathbf{t} \). Using a similar procedure to that described in Chap. 7 (Eq. 7.17), it can be show that vector βM can be transformed into βC = FβM, where F is a diagonal matrix with values equal to any real number, except zero values.
The maximized correlation between \( {H}_M={\mathbf{w}}_M^{\prime}\mathbf{a} \) and \( {I}_M={\boldsymbol{\upbeta}}_M^{\prime}\mathbf{t} \), or MESIM accuracy, is
where \( {\sigma}_{I_M}=\sqrt{{\boldsymbol{\upbeta}}_M^{\prime }{\mathbf{T}}_M{\boldsymbol{\upbeta}}_M} \) is the standard deviation of \( {I}_M={\boldsymbol{\upbeta}}_M^{\prime}\mathbf{t} \), and \( {\sigma}_{H_M}=\sqrt{{\boldsymbol{\upbeta}}_M^{\prime }{\mathbf{T}}_M{\boldsymbol{\Psi}}_M^{-1}{\mathbf{T}}_M{\boldsymbol{\upbeta}}_M} \) is the standard deviation of \( {H}_M={\mathbf{w}}_M^{\prime}\mathbf{a} \).
The maximized selection response and expected genetic gain per trait of MESIM are
and
respectively, where \( {\boldsymbol{\upbeta}}_{M_1} \) is the first eigenvector of matrix \( {\mathbf{T}}_M^{-1}{\boldsymbol{\Psi}}_M \). If vector \( {\boldsymbol{\upbeta}}_{M_1} \) is multiplied by matrix F, we obtain \( {\boldsymbol{\upbeta}}_{C_1}={\mathbf{F}\boldsymbol{\upbeta}}_{M_1} \); in this case, we can replace \( {\boldsymbol{\upbeta}}_{M_1} \) with \( {\boldsymbol{\upbeta}}_{C_1}={\mathbf{F}\boldsymbol{\upbeta}}_{M_1} \) in Eqs. (8.12) and (8.13), and the optimized MESIM index should be written as \( {I}_M={\boldsymbol{\upbeta}}_{C_1}^{\prime}\mathbf{y} \).
8.1.2 Estimating MESIM Parameters
We estimate the MESIM parameters using the same procedure described in Chap. 7 (Sect. 7.1.4) to estimate the ESIM parameters. Let \( \widehat{\mathbf{C}} \), \( \widehat{\mathbf{P}} \), and \( {\widehat{\mathbf{S}}}_M \) be the estimates of the genotypic, phenotypic, and marker scores covariance matrices, \( {\widehat{\mathbf{T}}}_M=\left[\begin{array}{cc}\widehat{\mathbf{P}}& {\widehat{\mathbf{S}}}_M\\ {}{\widehat{\mathbf{S}}}_M& {\widehat{\mathbf{S}}}_M\end{array}\right] \) and \( {\widehat{\boldsymbol{\Psi}}}_M=\left[\begin{array}{cc}\widehat{\mathbf{C}}& {\widehat{\mathbf{S}}}_M\\ {}{\widehat{\mathbf{S}}}_M& {\widehat{\mathbf{S}}}_M\end{array}\right] \) the estimated block matrices (Chap. 4) and \( \widehat{\mathbf{W}}={\widehat{\mathbf{T}}}_M^{-1}{\widehat{\boldsymbol{\Psi}}}_M \); then, to find the estimators \( {\widehat{\boldsymbol{\upbeta}}}_{M_1} \) and \( {\widehat{\lambda}}_{M_1}^2 \) of the first eigenvector (\( {\boldsymbol{\upbeta}}_{M_1} \)) and the first eigenvalue (\( {\lambda}_{M_1}^2 \)) respectively, we need to solve the equation
where \( {\widehat{\mu}}_j={\widehat{\lambda}}_{M_j}^4 \), j= 1, 2, …, 2t. For additional details, see Eqs. (7.22) and (7.23), and Sect. 7.1.5 of Chap. 7. The result of Equation (8.14) allow the MESIM index (\( {I}_M={\boldsymbol{\upbeta}}_{M_1}^{\prime}\mathbf{t} \)) to be estimated as \( {\widehat{I}}_M={\widehat{{\boldsymbol{\upbeta}}^{\prime}}}_{M_1}\mathbf{t} \), whereas the estimator of the maximized ESIM selection response and its expected genetic gain per trait can be denoted by
respectively.
8.1.3 Numerical Examples
To validate the MESIM theoretical results, we use a real maize (Zea mays) F2 population with 247 genotypes (each with two repetitions), 195 molecular markers, and two traits—plant height (PHT, cm) and ear height (EHT, cm)—evaluated in one environment. We coded the marker homozygous loci for the allele from the first parental line by 1, whereas the marker homozygous loci for the allele from the second parental line was coded by −1 and the marker heterozygous loci by 0. The estimated phenotypic, genetic, and marker scores covariance matrices were \( \widehat{\mathbf{P}}=\left[\begin{array}{cc}191.81& 106.89\\ {}106.89& 167.93\end{array}\right] \), \( \widehat{\mathbf{C}}=\left[\begin{array}{cc}83.00& 57.44\\ {}57.44& 59.80\end{array}\right] \), and \( {\widehat{\mathbf{S}}}_M=\left[\begin{array}{cc}15.750& 0.983\\ {}0.983& 28.083\end{array}\right] \) respectively, and the vector of economic weights was \( {\mathbf{a}}^{\prime }=\left[{\mathbf{w}}^{\prime}\kern0.5em {\mathbf{0}}^{\prime}\right] \), where \( {\mathbf{w}}^{\prime }=\left[-1\kern0.5em -1\right] \) and \( {\mathbf{0}}^{\prime }=\left[0\kern0.5em 0\right] \). Details of how to estimate the marker scores and their variance were given in Chap. 4.
We compare LMSI versus MESIM efficiency. The estimated LMSI vector of coefficients was \( {\widehat{\boldsymbol{\upbeta}}}^{\prime }={\mathbf{a}}^{\prime }{\widehat{\boldsymbol{\Psi}}}_M{\widehat{\mathbf{T}}}_M^{-1}=\left[-0.59\kern0.5em -0.18\kern0.5em -0.41\kern0.5em -0.82\right] \). Using a 10% selection intensity (kI = 1.755), the estimated LMSI selection response and the expected genetic gain per trait were \( \widehat{R}={k}_I\sqrt{{\widehat{{\boldsymbol{\upbeta}}^{\prime }}\widehat{\mathbf{T}}}_M\widehat{\boldsymbol{\upbeta}}}=20.41 \) and \( {\widehat{\mathbf{E}}}^{\prime }={k}_I\frac{{\widehat{{\boldsymbol{\upbeta}}^{\prime }}\widehat{\boldsymbol{\Psi}}}_M}{\sqrt{{\widehat{{\boldsymbol{\upbeta}}^{\prime }}\widehat{\mathbf{T}}}_M\widehat{\boldsymbol{\upbeta}}}}=\left[-10.09\kern0.5em -10.31\kern0.5em -2.53\kern0.5em -4.39\right] \) respectively, whereas the estimated LMSI accuracy was \( {\widehat{\rho}}_{H\widehat{I}}=\frac{{\widehat{\sigma}}_I}{{\widehat{\sigma}}_H}=0.72 \).
Vector \( {\widehat{\boldsymbol{\upbeta}}}_{M_1}^{\prime }=\left[0.089\kern0.5em -0.061\kern0.5em -0.536\kern0.5em 0.837\right] \) was the original estimated MESIM vector of coefficients. Using matrix \( \mathbf{F}=\left[\begin{array}{cccc}-0.1& 0& 0& 0\\ {}0& -0.1& 0& 0\\ {}0& 0& 0.75& 0\\ {}0& 0& 0& -0.75\end{array}\right] \), vector \( {\widehat{\boldsymbol{\upbeta}}}_{M_1}^{\prime } \) was transformed as \( {\widehat{\boldsymbol{\upbeta}}}_{C_1}^{\prime }={\widehat{\boldsymbol{\upbeta}}}_{M_1}^{\prime}\mathbf{F}=\left[-0.009\kern0.5em 0.006\kern0.5em -0.402\kern0.5em 0.628\right] \) and then the estimated MESIM index was \( {\widehat{I}}_M=-0.009\;\mathrm{PHT}+0.006\;\mathrm{EHT}-0.402\;{\mathrm{S}}_{\mathrm{PHT}}+0.628\;{\mathrm{S}}_{\mathrm{EHT}} \), where SPHT and SEHT denote the marker scores associated with PHT and EHT respectively. The estimated MESIM expected genetic gain, selection response, and accuracy were \( {\widehat{\mathbf{E}}}_M^{\prime }={k}_I\frac{{\widehat{{\boldsymbol{\upbeta}}^{\prime}}}_{C_1}{\widehat{\boldsymbol{\Psi}}}_M}{\sqrt{{\widehat{{\boldsymbol{\upbeta}}^{\prime}}}_{C_1}{\widehat{\mathbf{T}}}_M{\widehat{\boldsymbol{\upbeta}}}_{C_1}}}=\left[-3.438\kern0.5em -8.516\kern0.5em -3.319\kern0.5em -8.372\right] \), \( {\widehat{R}}_M={k}_I\sqrt{{\widehat{{\boldsymbol{\upbeta}}^{\prime}}}_{C_1}{\widehat{\mathbf{T}}}_M{\widehat{\boldsymbol{\upbeta}}}_{C_1}}=6.573 \) and \( {\widehat{\rho}}_{H_M{\widehat{I}}_M}=\frac{{\widehat{\sigma}}_{I_M}}{{\widehat{\sigma}}_{H_M}}=0.99 \) respectively.
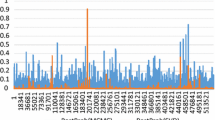
The inner product of the estimated LMSI and MESIM vector of coefficients were 1.221 and 0.556 respectively, whence the estimated LMSI selection response (20.41) divided by 1.221 was 16.716, and the estimated MESIM selection response (6.573) divided by 0.556 was 11.821. That is, the estimated LMSI selection response was higher than the estimated MESIM selection response for this data set. Similar results were found when we compared the estimated LMSI expected genetic gain per trait with the estimated MESIM expected genetic gain per trait. Finally, Fig. 8.1 presents the frequency distribution of the 247 estimated MESIM values for the real data set described earlier, which approaches normal distribution, as we would expect.
Frequency distribution of 247 estimated molecular eigen selection index method (MESIM) values for one selection cycle in an environment for a real maize (Zea mays) F2 population with 195 molecular markers and two traits, plant height (PHT, cm) and ear height (EHT, cm), and their associated marker scores SPHT and SEHT respectively
Now with a selection intensity of 10% (kI = 1.755), we compare the LMSI and MESIM efficiency using the simulated data set described in Sect. 2.8.1 of Chap. 2 for four phenotypic selection cycles, each with four traits (T1, T2, T3 and T4), 500 genotypes, and four replicates of each genotype. The economic weights for T1, T2, T3, and T4 were 1, −1, 1, and 1 respectively. For this data set, we did not use the linear transformation \( {\widehat{\boldsymbol{\upbeta}}}_{C_1}=\mathbf{F}{\widehat{\boldsymbol{\upbeta}}}_{M_1} \).
The estimated selection responses of the linear marker, combined genomic and genome-wide selection indices (LMSI, CLGSI, and GW-LMSI respectively; see Chaps. 4 and 5 for details) for four simulated selection cycles when their vectors of coefficients were normalized, are presented in Table 8.1. Also, in this table the selection responses of the estimated linear molecular, genomic, and genome-wide eigen selection index methods (MESIM, GESIM, and GW-ESIM respectively; details in Sect. 8.2) are shown for four simulated selection cycles. The average of the estimated LMSI selection response was 2.22, whereas the average of the estimated MESIM selection response was 1.69. The estimated LMSI selection response was higher than that of the MESIM.
Table 8.2 presents the estimated LMSI and MESIM expected genetic gains for four traits (T1, T2, T3, and T4) and their associated marker scores (S1, S2, S3, and S4) for four simulated selection cycles. The averages of the estimated LMSI expected genetic gains for the four traits and their associated marker scores were 12.74, −2.10, 1.60, 0.94, 5.70, −2.19, 0.71, and 0.64 respectively, whereas the averages of the estimated MESIM expected genetic gains for the four traits and their associated marker scores were 14.40, −0.38, −0.39, 0.34, 8.65, 0.47, −0.21, and −0.70 respectively. Except for trait T1 and its associated molecular scores, the estimated LMSI expected genetic gains per trait were higher than the estimated MESIM expected genetic gains. Thus, for this data set, LMSI efficiency was greater than MESIM efficiency.
Chapter 11 presents RIndSel, a user-friendly graphical unit interface in JAVA that is useful for estimating the LMSI and ESIM parameters and selecting parents for the next selection cycle.
8.2 The Linear Genomic Eigen Selection Index Method
The linear genomic eigen selection index method (GESIM) is based on the standard CLGSI described in Chap. 5, and uses genomic estimated breeding values (GEBVs) and phenotypic values jointly to predict the net genetic merit. Thus, conditions for constructing a valid GESIM are the same as those for constructing the CLGSI. Also, the MESIM theory described in Sect. 8.1 is directly applied to the GESIM and only minor changes are necessary in GESIM theory. For example, instead of marker scores, the GESIM uses GEBVs to predict the net genetic merit; thus, the details of the estimation process are the same as for the MESIM.
8.2.1 The GESIM Parameters
In the GESIM context, the net genetic merit can be written as
where \( {\mathbf{g}}^{\prime }=\left[{g}_1\kern0.5em \dots \kern0.5em {g}_t\right] \) is the vector of true breeding values, t is the number of traits, \( {\mathbf{w}}_1^{\prime }=\left[{w}_1\kern0.5em \cdots \kern0.5em {w}_t\right] \) is a vector of unknown economic weights associated with g, \( {\mathbf{w}}_2^{\prime }=\left[{0}_1\kern0.5em \cdots \kern0.5em {0}_t\right] \) is a null vector associated with the vector of genomic breeding values \( {\boldsymbol{\upgamma}}^{\prime }=\left[{\gamma}_1\kern0.5em {\gamma}_2\kern0.5em \dots \kern0.5em {\gamma}_t\right] \), \( {\mathbf{w}}^{\prime }=\left[{\mathbf{w}}_1^{\prime}\kern0.5em {\mathbf{w}}_2^{\prime}\right] \), and \( {\boldsymbol{\upalpha}}^{\prime }=\left[{\mathbf{g}}^{\prime}\kern0.5em {\boldsymbol{\upgamma}}^{\prime}\right] \). The estimator of γ is the GEBV (see Chap. 5 for additional details). The GESIM index can be written as
where \( {\mathbf{y}}^{\prime }=\left[{y}_1\kern0.5em \cdots \kern0.5em {y}_t\right] \) is the vector of phenotypic values; \( {\boldsymbol{\upbeta}}_{\mathbf{y}}^{\prime } \) and βγ are vectors of weights of phenotypic and genomic breeding values weights respectively; \( {\boldsymbol{\upbeta}}^{\prime }=\left[{\boldsymbol{\upbeta}}_{\mathbf{y}}^{\prime}\kern0.5em {\boldsymbol{\upbeta}}_{\boldsymbol{\upgamma}}^{\prime}\right] \) and \( {\mathbf{f}}^{\prime }=\left[{\mathbf{y}}^{\prime}\kern0.5em {\boldsymbol{\upgamma}}^{\prime}\right] \).
Let \( Var(H)={\mathbf{w}}^{\prime}\mathbf{Aw}={\sigma}_H^2 \) be the variance of H = w′α, \( Var(I)={\boldsymbol{\upbeta}}^{\prime}\boldsymbol{\Phi} \boldsymbol{\upbeta} ={\sigma}_I^2 \) the variance of I = β′f, and Cov(H, I) = w′Aβ = σHI the covariance between H and I, where \( \mathbf{A}= Var\left[\begin{array}{c}\mathbf{g}\\ {}\boldsymbol{\upgamma} \end{array}\right]=\left[\begin{array}{cc}\mathbf{C}& \boldsymbol{\Gamma} \\ {}\boldsymbol{\Gamma} & \boldsymbol{\Gamma} \end{array}\right] \) and \( \boldsymbol{\Phi} = Var\left[\begin{array}{c}\mathbf{y}\\ {}\boldsymbol{\upgamma} \end{array}\right]=\left[\begin{array}{cc}\mathbf{P}& \boldsymbol{\Gamma} \\ {}\boldsymbol{\Gamma} & \boldsymbol{\Gamma} \end{array}\right] \) are block matrices 2t × 2t (t is the number of traits) of covariance matrices and P, Γ, and C are covariance matrices of phenotypic (y), genomic (γ), and genetic (g) values respectively. Then, \( {\rho}_{HI}=\frac{{\mathbf{w}}^{\prime}\mathbf{A}\boldsymbol{\upbeta }}{\sqrt{{\mathbf{w}}^{\prime}\mathbf{A}\mathbf{w}}\sqrt{{\boldsymbol{\upbeta}}^{\prime}\boldsymbol{\Phi} \boldsymbol{\upbeta}}} \) is the correlation between H = w′α and I = β′f and the GESIM selection response can be written as
where kI is the standardized selection differential (or selection intensity) associated with the GESIM and \( {\sigma}_H=\sqrt{{\mathbf{w}}^{\prime}\mathbf{Aw}} \) is the standard deviation of the variance of H. It is assumed that kI is fixed, and that matrices Φ and A are known; then, we can maximize R by maximizing ρHI with respect to vectors w and β under the restrictions \( {\sigma}_H^2={\mathbf{w}}^{\prime}\mathbf{Aw} \), \( {\sigma}_I^2={\boldsymbol{\upbeta}}^{\prime}\boldsymbol{\Phi} \boldsymbol{\upbeta} \), and 0 < \( {\sigma}_H^2 \), \( {\sigma}_I^2 \) < ∞; similar to the MESIM.
It can be shown that the vector w in the GESIM context is
and that the net genetic merit can be written as \( {H}_G={\mathbf{w}}_G^{\prime}\boldsymbol{\upalpha} \). The correlation between \( {H}_G={\mathbf{w}}_G^{\prime}\boldsymbol{\upalpha} \) and I = β′f is \( {\rho}_{H_GI}=\frac{\sqrt{{\boldsymbol{\upbeta}}^{\prime}\boldsymbol{\Phi} \boldsymbol{\upbeta}}}{\sqrt{{\boldsymbol{\upbeta}}^{\prime }{\boldsymbol{\Phi} \mathbf{A}}^{-1}\boldsymbol{\Phi} \boldsymbol{\upbeta}}} \) and the GESIM index vector of coefficients that maximizes \( {\rho}_{H_GI} \) can be obtained from the equation
where I2t is an identity matrix of size 2t × 2t (t is the number of traits); the optimized GESIM index can be written as \( {I}_G={\boldsymbol{\upbeta}}_G^{\prime}\mathbf{f} \). By Eqs. (8.19) and (8.20), GESIM accuracy can be written as
where \( {\sigma}_{I_G}=\sqrt{{\boldsymbol{\upbeta}}_G^{\prime }{\boldsymbol{\Phi} \boldsymbol{\upbeta}}_G} \) is the standard deviation of \( {I}_G={\boldsymbol{\upbeta}}_G^{\prime}\mathbf{f} \), and \( {\sigma}_{H_G}=\sqrt{{\boldsymbol{\upbeta}}_G^{\prime }{\boldsymbol{\Phi} \mathbf{A}}^{-1}{\boldsymbol{\Phi} \boldsymbol{\upbeta}}_G} \) is the standard deviation of \( {H}_G={\mathbf{w}}_G^{\prime}\boldsymbol{\upalpha} \). In Eq. (8.20), \( {\lambda}_G^2={\rho}_{H_G{I}_G}^2 \) is the square of the canonical correlation between HG and IG, and βG is the canonical vector associated with \( {\lambda}_G^2={\rho}_{H_G{I}_G}^2 \).
The maximized GESIM selection response and expected genetic gain per trait are
and
respectively, where βG is the first eigenvector of matrix Φ−1A. Vector βG can be transformed as βCG = FβG, where F is a diagonal matrix defined earlier.
8.2.2 Numerical Examples
To compare the CLGSI versus GESIM theoretical results, we use a real maize (Zea mays) F2 population with 244 genotypes (each with two repetitions), 233 molecular markers, and three traits—grain yield (GY, ton ha−1), ear height (EHT, cm), and plant height (PHT, cm). We estimated matrices P and C using Eqs. (2.22) to (2.24) described in Chap. 2, whence the estimated matrices were \( \widehat{\mathbf{P}}=\left[\begin{array}{lll}0.45& 1.33& 2.33\\ {}1.33& 65.07& 83.71\\ {}2.33& 83.71& 165.99\end{array}\right] \) and \( \widehat{\mathbf{C}}=\left[\begin{array}{lll}0.07& 0.61& 1.06\\ {}0.61& 17.93& 22.75\\ {}1.06& 22.75& 44.53\end{array}\right] \). In a similar manner, we estimated matrix Γ by applying Eqs. (5.21) to (5.23) described in Chap. 5 using phenotypic and marker information jointly; the estimated matrix was \( \widehat{\boldsymbol{\Gamma}}=\left[\begin{array}{lll}0.07& 0.65& 1.05\\ {}0.65& 10.62& 14.25\\ {}1.05& 14.25& 26.37\end{array}\right] \). The selection intensity for making a selection cycle was 10% (kI = 1.755) and the vector of economic weights was \( {\mathbf{w}}^{\prime }=\left[5\kern0.5em -0.1\kern0.5em -0.1\kern0.5em 0\kern0.5em 0\kern0.5em 0\right] \). To obtain the estimated vector of coefficient of CLGSI (\( \widehat{\boldsymbol{\upbeta}}={\widehat{\boldsymbol{\Phi}}}^{-1}\widehat{\mathbf{A}}\mathbf{w} \)) and GESIM (Eq. 8.20), it is necessary to construct matrices \( \widehat{\mathbf{A}}=\left[\begin{array}{ll}\widehat{\mathbf{C}}& \widehat{\boldsymbol{\Gamma}}\\ {}\widehat{\boldsymbol{\Gamma}}& \widehat{\boldsymbol{\Gamma}}\end{array}\right] \) and \( \widehat{\boldsymbol{\Phi}}=\left[\begin{array}{ll}\widehat{\mathbf{P}}& \widehat{\boldsymbol{\Gamma}}\\ {}\widehat{\boldsymbol{\Gamma}}& \widehat{\boldsymbol{\Gamma}}\end{array}\right] \).
The estimated CLGSI vector of coefficients for the traits GY, EHT, and PHT and their associated GEBVs (GEBVGY, GEBVEHT, and GEBVPHT respectively) was \( {\widehat{\boldsymbol{\upbeta}}}^{\prime }=\left[0.08\kern0.5em -0.02\kern0.5em -0.01\kern0.5em 4.92\kern0.5em -0.08\kern0.5em -0.09\right] \), whereas the estimated CLGSI selection response, accuracy, and expected genetic gain per trait were \( \widehat{R}={k}_I\sqrt{{\widehat{\boldsymbol{\upbeta}}}^{\prime}\widehat{\boldsymbol{\Phi}}\widehat{\boldsymbol{\upbeta}}}=1.54 \), \( {\widehat{\rho}}_{HI}=\frac{{\widehat{\sigma}}_I}{{\widehat{\sigma}}_H}=0.814 \), and \( {\widehat{\mathbf{E}}}^{\prime }={k}_I\frac{{\widehat{\boldsymbol{\upbeta}}}^{\prime}\widehat{\mathbf{A}}}{\sqrt{{\widehat{\boldsymbol{\upbeta}}}^{\prime}\widehat{\boldsymbol{\Phi}}\widehat{\boldsymbol{\upbeta}}}}=\left[0.36\kern0.5em 1.04\kern0.5em 1.70\kern0.5em 0.36\kern0.5em 1.53\kern0.5em 2.38\right] \) respectively. Finally, \( \widehat{I}=0.08\mathrm{GY}-0.02\mathrm{EHT}-0.01\mathrm{PHT}+4.92{\mathrm{GEBV}}_{\mathrm{GY}}-0.08{\mathrm{GEBV}}_{\mathrm{EHT}}-0.09{\mathrm{GEBV}}_{\mathrm{PHT}} \) was the estimated CLGSI.
The estimated GESIM vector of coefficients, selection response, accuracy, and expected genetic gain per trait were \( {\widehat{\boldsymbol{\upbeta}}}_{G_1}^{\prime }=\left[-0.207\kern0.5em 0.029\kern0.5em 0.041\kern0.5em 0.820\kern0.5em 0.337\kern0.5em 0.411\right] \), \( {\widehat{R}}_G={k}_I\sqrt{{\widehat{\boldsymbol{\upbeta}}}_{G_1}^{\prime }{\widehat{\boldsymbol{\Phi}}\widehat{\boldsymbol{\upbeta}}}_{G_1}}=6.288 \), \( {\widehat{\rho}}_{{\widehat{H}}_G{\widehat{I}}_G}=\frac{\sqrt{{\widehat{\boldsymbol{\upbeta}}}_{G_1}^{\prime }{\widehat{\boldsymbol{\Phi}}\widehat{\boldsymbol{\upbeta}}}_{G_1}}}{\sqrt{{\widehat{\boldsymbol{\upbeta}}}_{G_1}^{\prime }{\widehat{\boldsymbol{\Phi}}\widehat{\mathbf{A}}}^{-1}{\widehat{\boldsymbol{\Phi}}\widehat{\boldsymbol{\upbeta}}}_{G_1}}}=0.9056 \), and \( {\widehat{\mathbf{E}}}_G^{\prime }={k}_1\frac{{\widehat{\boldsymbol{\upbeta}}}_{G_1}^{\prime}\widehat{\mathbf{A}}}{\sqrt{{\widehat{\boldsymbol{\upbeta}}}_{G_1}^{\prime }{\widehat{\boldsymbol{\Phi}}\widehat{\boldsymbol{\upbeta}}}_{G_1}}}=\left[0.369\kern0.5em 5.528\kern0.5em 9.186\kern0.5em 0.370\kern0.5em 5.250\kern0.5em 8.702\right] \) respectively.
Fig. 8.2 presents the frequency distribution of the 244 estimated GESIM index values for one (Fig. 8.2a) and three traits (Fig. 8.2b) using the real data set described earlier. The frequency distribution of the estimated GESIM index values approaches the normal distribution for both indices.
Frequency distribution of the 244 estimated genomic eigen selection index method (GESIM) values for the one-trait case (a) and for the three-trait case (b) for one selection cycle in an environment for a real maize (Zea mays) F2 population with 233 molecular markers. Note that the frequency distribution of the estimated GESIM index values approaches normal distribution for both indices
Now, we compare the estimated CLGSI and GESIM selection response and expected genetic gain per trait using the simulated data set described in Sect. 2.8.1 of Chap. 2 for four phenotypic selection cycles, each with four traits (T1, T2, T3 and T4), 500 genotypes, and four replicates per genotype. The economic weights of T1, T2, T3, and T4 were 1, −1, 1, and 1 respectively and the selection intensity for both indices was 10% (kI = 1.755). For this data set, matrix F was an identity matrix of size 8 × 8 in all four selection cycles.
For this data set, the averages of the estimated CLGSI and GESIM selection responses were 0.68 and 2.74 (Table 8.1) respectively. The estimated CLGSI selection response was lower than the estimated GESIM selection response. Table 8.3 presents the estimated CLGSI and GESIM expected genetic gain for four traits (T1, T2, T3, and T4) and their associated genomic estimated breeding values (GEBV1, GEBV2, GEBV3, and GEBV4) for four simulated selection cycles. The averages of the estimated CLGSI expected genetic gains for the four traits and their associated GEBVs were 7.45, −3.35, 2.68, 1.09, 7.13, −3.68, 3.13, and 2.69 respectively, whereas the averages of the estimated GESIM expected genetic gains for the four traits and their associated GEBVs were 8.18, −3.08, 2.27, 0.71, 7.46, −3.53, 2.86, and 2.39 respectively. The estimated CLGSI and GESIM expected genetic gains per trait were very similar.
8.3 The Genome-Wide Linear Eigen Selection Index Method
The MESIM requires regressing phenotypic values on marker coded values to predict the marker score values for each individual candidate for selection, and then combining the marker scores with phenotypic information using the MESIM to obtain a final prediction of the net genetic merit. In addition, the GESIM requires fitting of a statistical model to estimate all available marker effects in the training population; these estimates are then used to obtain GEBVs, which are predictors of breeding values. Crossa and Cerón-Rojas (2011) extended the ESIM theory to a genome-wide linear molecular ESIM (GW-ESIM) similar to the GW-LMSI described in Chap. 4. The GW-LMSI and GW-ESIM are very similar and only minor changes are necessary in GW-ESIM; for example, instead of estimating the GW-LMSI vector of coefficients according to the LPSI method (Chap. 2), the GW-ESIM vector of coefficients is estimated according to the singular value decomposition (SVD) described in Chap. 7.
8.3.1 The GW-ESIM Parameters
In the GW-ESIM context, the net genetic merit can be written as
where \( {\mathbf{g}}^{\prime }=\left[{g}_1\kern0.5em \dots \kern0.5em {g}_t\right] \) is the vector of true breeding values, t is the number of traits, \( {\mathbf{w}}_1^{\prime }=\left[{w}_1\kern0.5em \cdots \kern0.5em {w}_t\right] \) is the vector of unknown economic weights associated with the breeding values; \( {\mathbf{w}}_2^{\prime }=\left[{0}_1\kern0.5em \cdots \kern0.5em {0}_N\right] \) is a null vector associated with the vector of marker code values \( {\mathbf{m}}^{\prime }=\left[{m}_1\kern0.5em \cdots \kern0.5em {m}_N\right] \), where mj (j = 1, 2, …, N = number of markers) is the jth marker in the training population; \( {\mathbf{w}}^{\prime }=\left[{\mathbf{w}}_1^{\prime}\kern0.5em {\mathbf{w}}_2^{\prime}\right] \) and \( \mathbf{x}=\left[{\mathbf{g}}^{\prime}\kern0.5em {\mathbf{m}}^{\prime}\right] \). The GW-ESIM (I) index combines the phenotypic value and all the marker information of individuals to predict Eq. (8.24) values in each selection cycle and can be written as
where \( {\boldsymbol{\upbeta}}_y^{\prime } \) and βm are vectors of phenotypic and marker weights respectively; \( {\mathbf{y}}^{\prime }=\left[{y}_1\kern0.5em \cdots \kern0.5em {y}_t\right] \) is the vector of phenotypic values; m was defined in Eq. (8.24); \( {\boldsymbol{\upbeta}}^{\prime }=\left[{\boldsymbol{\upbeta}}_y^{\prime}\kern0.5em {\boldsymbol{\upbeta}}_m^{\prime}\right] \) and \( {\mathbf{q}}^{\prime }=\left[{\mathbf{y}}^{\prime}\kern0.5em {\mathbf{m}}^{\prime}\right] \).
Let \( {\sigma}_I^2={\boldsymbol{\upbeta}}^{\prime}\mathbf{Q}\boldsymbol{\upbeta } \) and \( {\sigma}_H^2={\mathbf{w}}^{\prime}\mathbf{Zw} \) be the variance of I = β′q and H = w′z respectively, and σHI = w′Zβ the covariance between I and H, where \( \mathbf{Q}= Var\left[\begin{array}{l}\mathbf{y}\\ {}\mathbf{m}\end{array}\right]=\left[\begin{array}{cc}\mathbf{P}& {\mathbf{G}}_M^{\prime}\\ {}{\mathbf{G}}_M& \mathbf{M}\end{array}\right] \) and \( \mathbf{X}= Var\left[\begin{array}{l}\mathbf{g}\\ {}\mathbf{m}\end{array}\right]=\left[\begin{array}{cc}\mathbf{C}& {\mathbf{G}}_M^{\prime}\\ {}{\mathbf{G}}_M& \mathbf{M}\end{array}\right] \) are block matrices of size (t + N) × (t + N) (t is the number of traits and N is the number of markers) where P = Var(y), M = Var(m), C = Var(g), and GM = cov (y, m) = cov (g, m) are covariance matrices of phenotypic (y), coded marker (m), and genetic (g) values respectively, whereas GM is the covariance matrix between y and m, and between g and m (for details see Chap. 4); w and β were defined earlier. Note that although the size of matrices P and C are t × t, the sizes of matrices M and GM are N × N and N × t respectively. Thus, if the number of markers is very high, the size of matrices M and GM could also be very high.
In Chap. 4 we described matrix M as
where (1 − 2θij) and θij (i, j= 1, 2, …, N= number of markers) are the covariance (or correlation) and the recombination frequency between the ith and jth marker respectively, whereas matrix GM can be written as
where (1 − 2rik)αqk (i= 1, 2, …, N, k= 1, 2, …, NQ = number of quantitative trait loci (QTL), q = 1, 2, …, t) is the covariance between the qth trait and the ith marker; rik is the recombination frequency between the ith and kth QTL, and αqk is the effect of the kth QTL over the qth trait.
Let \( {\rho}_{HI}=\frac{{\mathbf{w}}^{\prime}\mathbf{X}\boldsymbol{\upbeta }}{\sqrt{{\mathbf{w}}^{\prime}\mathbf{X}\mathbf{w}}\sqrt{{\boldsymbol{\upbeta}}^{\prime}\mathbf{Q}\boldsymbol{\upbeta }}} \) be the correlation between I = β′q and H = w′x; then, the GW-ESIM selection response can be written as
where kI is the standardized selection differential (or selection intensity) associated with GW-ESIM and \( {\sigma}_H=\sqrt{{\mathbf{w}}^{\prime}\mathbf{Xw}} \) is the standard deviation of the variance of H.
Assuming that kI is fixed, and that matrices Q and X are known, we can maximize R (Eq. 8.28) by maximizing ρHI with respect to vectors w′ and β under the restrictions \( {\sigma}_H^2={\mathbf{w}}^{\prime}\mathbf{Xw} \), \( {\sigma}_I^2={\boldsymbol{\upbeta}}^{\prime}\mathbf{Q}\boldsymbol{\upbeta } \), and 0 < \( {\sigma}_H^2 \),\( {\sigma}_I^2 \) < ∞, similar to the MESIM and GESIM. It can be shown that vector w can be written as
and that \( {H}_W={\mathbf{w}}_W^{\prime}\mathbf{x} \) is the net genetic merit in the GW-ESIM context. The correlation between \( {H}_W={\mathbf{w}}_W^{\prime}\mathbf{x} \) and I = β′q is \( {\rho}_{H_WI}=\frac{\sqrt{{\boldsymbol{\upbeta}}^{\prime}\mathbf{Q}\boldsymbol{\upbeta }}}{\sqrt{{\boldsymbol{\upbeta}}^{\prime }{\mathbf{QX}}^{-1}\mathbf{Q}\boldsymbol{\upbeta }}} \) and the GW-ESIM vector of coefficients (β) that maximizes \( {\rho}_{H_WI} \) can be obtained from equation
where I(t + N) is an identity matrix of size (t + N) × (t + N) and \( {I}_W={\boldsymbol{\upbeta}}_W^{\prime}\mathbf{q} \) is the optimized GW-ESIM. The accuracy of the GW-ESIM can be written as
where \( {\sigma}_{I_W}=\sqrt{{\boldsymbol{\upbeta}}_W^{\prime }{\mathbf{Q}\boldsymbol{\upbeta}}_W} \) is the standard deviation of \( {I}_W={\boldsymbol{\upbeta}}_W^{\prime}\mathbf{q} \), and \( {\sigma}_{H_W}=\sqrt{{\boldsymbol{\upbeta}}_W^{\prime }{\mathbf{Q}\mathbf{X}}^{-1}{\mathbf{Q}\boldsymbol{\upbeta}}_W} \) is the standard deviation of \( {H}_W={\mathbf{w}}_W^{\prime}\mathbf{x} \). In Eq. (8.30) \( {\lambda}_W^2={\rho}_{H_W{I}_W}^2 \) is the square of the canonical correlation between HW and IW.
The maximized GW-ESIM selection response and expected genetic gain per trait are
and
respectively, where βW is the first eigenvector of Eq. (8.30).
8.3.2 Estimating GW-ESIM Parameters
In Chap. 2, Eqs. (2.22) to (2.24), we described the restricted maximum likelihood methods to estimate matrices C and P, which can be denoted by \( \widehat{\mathbf{C}} \) and \( \widehat{\mathbf{P}} \). In Chap. 4, we described how to estimate matrices M and GM, which can be denoted by \( \widehat{\mathbf{M}} \) and \( {\widehat{\mathbf{G}}}_M \). With these estimates, we constructed the block estimated matrices as \( \widehat{\mathbf{Q}}=\left[\begin{array}{cc}\widehat{\mathbf{P}}& {\widehat{\mathbf{G}}}_M^{\prime}\\ {}{\widehat{\mathbf{G}}}_M& \widehat{\mathbf{M}}\end{array}\right] \) and \( \widehat{\mathbf{X}}=\left[\begin{array}{cc}\widehat{\mathbf{C}}& {\widehat{\mathbf{G}}}_M^{\prime}\\ {}{\widehat{\mathbf{G}}}_M& \widehat{\mathbf{M}}\end{array}\right] \), whence we obtained the equation
j = 1, 2, …, (t + N), where (t + N) is the number of traits and markers in the GW-ESIM index. Similar to the MESIM, we obtained estimators \( {\widehat{\boldsymbol{\upbeta}}}_{W_1} \) and \( {\widehat{\lambda}}_{W_1}^2 \) of the first eigenvector \( {\boldsymbol{\upbeta}}_{W_1} \) and the first eigenvalue \( {\widehat{\lambda}}_{W_1}^2 \) respectively, from equation
where \( \widehat{\mathbf{E}}={\widehat{\mathbf{Q}}}^{-}\widehat{\mathbf{X}} \) and \( {\widehat{\mu}}_j={\widehat{\lambda}}_{W_j}^4 \). These results allow the GW-ESIM index selection response and its expected genetic gain per trait to be estimated as \( {\widehat{I}}_W={\widehat{{\boldsymbol{\upbeta}}^{\prime}}}_{W_1}\widehat{\mathbf{q}} \), \( {\widehat{R}}_W={k}_I\sqrt{{\widehat{{\boldsymbol{\upbeta}}^{\prime}}}_{W_1}\widehat{\mathbf{Q}}{\boldsymbol{\upbeta}}_{W_1}^{\prime }} \) and \( {\widehat{\mathbf{E}}}_w={k}_I\frac{\widehat{\mathbf{X}}{\widehat{{\boldsymbol{\upbeta}}^{\prime}}}_{W_1}}{\sqrt{{\widehat{{\boldsymbol{\upbeta}}^{\prime}}}_{W_1}\widehat{\mathbf{Q}}{\boldsymbol{\upbeta}}_{W_1}^{\prime }}} \) respectively, whereas the estimator of GW-ESIM accuracy is \( {\widehat{\lambda}}_{W_1} \).
8.3.3 Numerical Examples
We compare the estimated GW-LMSI and GW-ESIM selection responses using the simulated data set described in Sect. 2.8.1 of Chap. 2, with a selection intensity of 10% (kI = 1.755). Table 8.1 presents the estimated GW-LMSI selection response for four simulated selection cycles when their vectors of coefficients are normalized, whence it can be seen that the average estimated GW-LMSI selection response was 0.87. Table 8.1 also presents the estimated GW-ESIM selection response for four simulated selection cycles; the average of the estimated GW-ESIM selection responses was 0.93. Thus, for this data set, the estimated GW-LMSI and selection responses were very similar.
8.4 The Restricted Linear Genomic Eigen Selection Index Method
The restricted linear genomic eigen selection index method (RGESIM) is based on the restricted linear phenotypic ESIM (RESIM) theory described in Chap. 7. In the RESIM, the breeder’s objective is to improve only (t − r) of t (r < t) traits, leaving r of them fixed. The same is true for RGESIM, but in this case, we should impose 2r restrictions, i.e., we need to fix r traits and their associated r GEBV to obtain results similar to those obtained with the RESIM (see Chap. 7 for details). This is the main difference between the RGESIM and the RESIM.
It can be shown that Cov(I, α) = Aβ is the covariance between the breeding value vector (α′ = [g′ γ′]) and the GESIM index (I = β′f). In the RGESIM, we want some covariances between the linear combinations of α (\( {\mathbf{U}}_{\mathbf{G}}^{\prime}\boldsymbol{\upalpha} \)) and I = β′f to be zero, i.e., \( Cov\left({\mathrm{I}}_{\mathrm{G}},{\mathbf{U}}_G^{\prime}\boldsymbol{\upalpha} \right)={\mathbf{U}}_G^{\prime}\mathbf{A}\boldsymbol{\upbeta } =\mathbf{0} \), where \( {\mathbf{U}}_{\mathbf{G}}^{\prime } \) is a matrix 2(t − 1) × 2t of 1s and 0s (1 indicates that the trait and its associated GEBV are restricted, and 0 indicates that the trait and its GEBV have no restrictions). We can solve this problem by maximizing \( \frac{{\boldsymbol{\upbeta}}^{\prime}\mathbf{A}\boldsymbol{\upbeta }}{\sqrt{{\boldsymbol{\upbeta}}^{\prime}\boldsymbol{\Phi} \boldsymbol{\upbeta}}} \) with respect to vector β under the restriction \( {\mathbf{U}}_G^{\prime}\mathbf{A}\boldsymbol{\upbeta } =\mathbf{0} \) and β′β = 1 similar to the RESIM, or by maximizing the correlation between H = w′α and I = β′f, \( {\rho}_{HI}=\frac{{\mathbf{w}}^{\prime}\mathbf{A}\boldsymbol{\upbeta }}{\sqrt{{\mathbf{w}}^{\prime}\mathbf{A}\mathbf{w}}\sqrt{{\boldsymbol{\upbeta}}^{\prime}\boldsymbol{\Phi} \boldsymbol{\upbeta}}} \), with respect to vectors w′ and β under the restrictions \( {\mathbf{U}}_G^{\prime}\mathbf{A}\boldsymbol{\upbeta } =\mathbf{0} \), \( {\sigma}_H^2={\mathbf{w}}^{\prime}\mathbf{Aw} \), \( {\sigma}_I^2={\boldsymbol{\upbeta}}^{\prime}\boldsymbol{\Phi} \boldsymbol{\upbeta} \) and 0 < \( {\sigma}_H^2 \), \( {\sigma}_I^2 \) < ∞, as we did for the GESIM.
8.4.1 The RGESIM Parameters
To obtain the RGESIM vector of coefficients, we maximize the function
with respect to β and v′, where v′ = [v1 v2 ⋯ v2(r − 1)] is a vector of Lagrange multipliers. The derivatives of function f(β, v′) with respect to β and v′ can be written as
respectively, where Eq. (8.38) denotes the restriction imposed for maximizing Eq. (8.36). Using algebraic methods on Eq. (8.37), we get
where \( {\lambda}_{RG}^2={h}_{I_{RG}}^2 \), \( {h}_{I_{RG}}^2 \) is the RGESIM heritability obtained under the restriction \( {\mathbf{U}}_G^{\prime}\mathbf{A}\boldsymbol{\upbeta } =\mathbf{0} \); KRG = [I2t − QRG], I2t is an identity matrix of size 2t × 2t, and \( {\mathbf{Q}}_{RG}={\boldsymbol{\Phi}}^{-1}{\mathbf{A}\mathbf{U}}_G{\left({\mathbf{U}}_G^{\prime }{\mathbf{A}\boldsymbol{\Phi}}^{-1}{\mathbf{A}\mathbf{U}}_G\right)}^{-1}{\mathbf{U}}_G^{\prime}\mathbf{A} \). When \( {\mathbf{U}}_G^{\prime } \) is a null matrix, \( {\boldsymbol{\upbeta}}_{RG}^{\prime }={\boldsymbol{\upbeta}}_G^{\prime } \) (the vector of the GESIM coefficients); thus, the RGESIM is more general than the GESIM and includes the GESIM as a particular case. The RGESIM index \( {I}_{GR}={\boldsymbol{\upbeta}}_{RG}^{\prime}\mathbf{y} \) and its selection response and expected genetic gain per trait use the first eigenvector of matrix KGΦ−1A. It can be shown that the vector of coefficients of \( H={\mathbf{w}}_{RG}^{\prime}\boldsymbol{\upalpha} \) in the RGESIM can be written as
where \( {\mathbf{Q}}_{RG}^{\prime }={\mathbf{A}\mathbf{U}}_G{\left({\mathbf{U}}_G^{\prime }{\mathbf{A}\boldsymbol{\Phi}}^{-1}{\mathbf{A}\mathbf{U}}_G\right)}^{-1}{\mathbf{U}}_G^{\prime }{\mathbf{A}\boldsymbol{\Phi}}^{-1} \).
Note that the restriction \( {\mathbf{U}}_G^{\prime}\mathbf{A}\boldsymbol{\upbeta } =\mathbf{0} \) can be written as β′AUG = 0; this means that β′Q′RG = 0 and that the covariance between \( {H}_{RG}={\mathbf{w}}_{RG}^{\prime}\boldsymbol{\upalpha} \) and \( {I}_{RG}={\boldsymbol{\upbeta}}_{RG}^{\prime}\mathbf{f} \) (\( {\sigma}_{H_{RG}{I}_{RG}} \)) can be written as
Equation (8.41) indicates that \( {\sigma}_{H_{RG}{I}_{RG}} \) is equal to the variance of \( {I}_{RG}={\boldsymbol{\upbeta}}_{RG}^{\prime}\mathbf{f} \) (\( {\sigma}_{I_{RG}}^2={\boldsymbol{\upbeta}}_{RG}^{\prime }{\boldsymbol{\Phi} \boldsymbol{\upbeta}}_{RG} \)); therefore, the maximized correlation between IRG and HRG or RGESIM accuracy can be written as
where \( {\mathbf{w}}_{RG}^{\prime }{\mathbf{Aw}}_{RG} \) is the variance of HRG. Hereafter, to simplify the notation, we write Eq. (8.42) as λRG.
The maximized selection response and the expected genetic gain per trait of the RGESIM are
and
respectively, where βRG is the first eigenvector of matrix KRGΦ−1A.
8.4.2 Estimating RGESIM Parameters
In Sect. 8.2, we indicated how to estimate matrices P, Γ, and C using phenotypic and genomic information, whence we can estimate matrices \( \mathbf{A}=\left[\begin{array}{ll}\mathbf{C}& \boldsymbol{\Gamma} \\ {}\boldsymbol{\Gamma} & \boldsymbol{\Gamma} \end{array}\right] \) and \( \Phi =\left[\begin{array}{ll}\mathbf{P}& \boldsymbol{\Gamma} \\ {}\boldsymbol{\Gamma} & \boldsymbol{\Gamma} \end{array}\right] \). Those methods are also valid for the RGESIM. This means that the SVD methods described for estimating MESIM parameters are also valid for estimating RGESIM parameters.
8.4.3 Numerical Examples
With a selection intensity of 10% (kI = 1.755), we compare the CRLGSI (for details see Chap. 6) versus the RGESIM theoretical results using a real maize (Zea mays) F2 population with 244 genotypes (each with two repetitions), 233 molecular markers, and three traits—GY (ton ha−1), EHT (cm), and PHT (cm)—described in Sect. 8.2.2, where \( \widehat{\mathbf{P}}=\left[\begin{array}{ccc}0.45& 1.33& 2.33\\ {}1.33& 65.07& 83.71\\ {}2.33& 83.71& 165.99\end{array}\right] \), \( \widehat{\mathbf{C}}=\left[\begin{array}{ccc}0.07& 0.61& 1.06\\ {}0.61& 17.93& 22.75\\ {}1.06& 22.75& 44.53\end{array}\right] \), and \( \widehat{\boldsymbol{\Gamma}}=\left[\begin{array}{ccc}0.07& 0.65& 1.05\\ {}0.65& 10.62& 14.25\\ {}1.05& 14.25& 26.37\end{array}\right] \) were the estimated matrices of P, C, and Γ respectively.
We have indicated that the main difference between the RLPSI and the CRLGSI is the matrix \( {\mathbf{U}}_C^{\prime } \), on which we now need to impose two restrictions: one for the trait and another for its associated GEBV. Consider the data set described earlier and suppose that we restrict the trait GY (ton ha−1) and its associated GEBVGY; then, matrix \( {\mathbf{U}}_C^{\prime } \) should be constructed as \( {\mathbf{U}}_{C1}^{\prime }=\left[\begin{array}{llllll}1& 0& 0& 0& 0& 0\\ {}0& 0& 0& 1& 0& 0\end{array}\right] \). If we restrict traits GY and EHT (cm) and their associated GEBVGY and GEBVEHT, matrix \( {\mathbf{U}}_C^{\prime } \) should be constructed as \( {\mathbf{U}}_{C2}^{\prime }=\left[\begin{array}{llllll}1& 0& 0& 0& 0& 0\\ {}0& 1& 0& 0& 0& 0\\ {}0& 0& 0& 1& 0& 0\\ {}0& 0& 0& 0& 1& 0\end{array}\right] \), etc. The procedure for obtaining matrices \( {\widehat{\mathbf{K}}}_{RG}=\left[{\mathbf{I}}_{2t}-{\widehat{\mathbf{Q}}}_{RG}\right] \) and \( {\widehat{\mathbf{Q}}}_{RG}={\widehat{\boldsymbol{\Phi}}}^{-1}\widehat{\mathbf{A}}{\mathbf{U}}_G{\left({\mathbf{U}}_G^{\prime }{\widehat{\mathbf{A}}\widehat{\boldsymbol{\Phi}}}^{-1}\widehat{\mathbf{A}}{\mathbf{U}}_{\mathrm{G}}\right)}^{-1}{\mathbf{U}}_G^{\prime}\widehat{\mathbf{A}} \) was described in Chap. 6, and is also valid for estimating RGESIM parameters.
The estimated CRLGSI vector of coefficients is \( {\widehat{\boldsymbol{\upbeta}}}_{CR}={\widehat{\mathbf{K}}}_{RG}\widehat{\boldsymbol{\upbeta}} \), where \( \widehat{\boldsymbol{\upbeta}}={\widehat{\boldsymbol{\Phi}}}^{-1}\widehat{\mathbf{A}}\mathbf{w} \) is the estimated CLGSI vector of coefficients (Chap. 6). Let w′ = [5 − 0.1 − 0.1 0 0 0] be the vector of economic weights and suppose that we restrict trait GY and its associated GEBVGY; in this case, \( {\mathbf{U}}_{C1}^{\prime }=\left[\begin{array}{llllll}1& 0& 0& 0& 0& 0\\ {}0& 0& 0& 1& 0& 0\end{array}\right] \), and according to matrices \( \widehat{\mathbf{P}} \), \( \widehat{\mathbf{C}} \), and \( \widehat{\boldsymbol{\Gamma}} \) described earlier, \( {\widehat{\boldsymbol{\upbeta}}}_{CR}^{\prime }=\left[0.076\kern0.5em -0.004\kern0.5em -0.018\kern0.5em 2.353\kern0.5em -0.096\kern0.5em -0.082\right] \) was the estimated CRLGSI vector of coefficients and the estimated CRLGSI was
where GEBVGY, GEBVEHT, and GEBVPHT are the GEBVs associated with the traits GY, EHT, and PHT respectively. The same procedure is valid for two or more restrictions.
The estimated CRLGSI selection response and expected genetic gain per trait were \( {\widehat{R}}_{CR}={k}_I\sqrt{{\widehat{\boldsymbol{\upbeta}}}_{CR}^{\prime }{\widehat{\boldsymbol{\Phi}}\widehat{\boldsymbol{\upbeta}}}_{CR}}=0.96 \) and \( {\widehat{\mathbf{E}}}_{CR}^{\prime }={k}_I=\frac{{\widehat{\boldsymbol{\upbeta}}}_{CR}^{\prime}\widehat{\mathbf{A}}}{\sqrt{{\widehat{\boldsymbol{\upbeta}}}_{CR}^{\prime }{\widehat{\boldsymbol{\Phi}}\widehat{\boldsymbol{\upbeta}}}_{CR}}}\left[0\kern0.5em -3.53\kern0.5em -6.03\kern0.5em 0\kern0.5em -2.93\kern0.5em -4.87\right] \) respectively, whereas the estimated CRLGSI accuracy was \( {\widehat{\rho}}_{Hl_{\mathrm{C}R}}=\frac{{\widehat{\sigma}}_{I_{\mathrm{C}R}}}{{\widehat{\sigma}}_H}=0.51 \). Note that in \( {\widehat{\mathbf{E}}}_{CR}^{\prime } \), the trait GY and its associated GEBVGY have null values, as we would expect.
The estimated RGESIM vector of coefficients was \( {\widehat{\boldsymbol{\upbeta}}}_{CR}^{\prime }=\left[0.015\kern0.5em -0.001\kern0.5em -0.004\kern0.5em 0.998\kern0.5em -0.029\kern0.5em -0.045\right] \), and the estimated RGESIM index was \( {\widehat{I}}_{RG}=0.015\mathrm{GY}-0.001\mathrm{EHT}-0.004\mathrm{PHT}+0.998{\mathrm{GEBV}}_{\mathrm{GY}}-0.029{\mathrm{GEBV}}_{\mathrm{EHT}}-0.045{\mathrm{GEBV}}_{\mathrm{PHT}} \) where GEBVGY, GEBVEHT, and GEBVPHT are the GEBVs associated with traits GY, EHT, and PHT respectively. The same procedure is valid for two or more restrictions.
The estimated RGESIM selection response and expected genetic gain per trait were \( {\widehat{R}}_{RG}={k}_I\sqrt{{\widehat{\boldsymbol{\upbeta}}}_{RG}^{\prime }{\widehat{\boldsymbol{\Phi}}\widehat{\boldsymbol{\upbeta}}}_{RG}}=0.37 \) and \( {\widehat{\mathbf{E}}}_{RG}^{\prime }={k}_I=\frac{{\widehat{\boldsymbol{\upbeta}}}_{RG}^{\prime}\widehat{\mathbf{A}}}{\sqrt{{\widehat{\boldsymbol{\upbeta}}}_{RG}^{\prime }{\widehat{\boldsymbol{\Phi}}\widehat{\boldsymbol{\upbeta}}}_{RG}}}\left[0\kern0.5em -3.28\kern0.5em -6.03\kern0.5em 0\kern0.5em -2.93\kern0.5em -5.40\right] \) respectively, whereas the estimated RGESIM accuracy was \( {\widehat{\rho}}_{{\widehat{H}}_{RG}{\widehat{I}}_{RG}}=\frac{{\widehat{\sigma}}_{{\widehat{I}}_{RG}}}{{\widehat{\sigma}}_{{\widehat{H}}_{RG}}}=0.86 \).
Fig. 8.3 presents the frequency distribution of the 244 estimated RGESIM index values for two null restrictions on traits GY and EHT and their associated GEBVGY and GEBVEHT, for one selection cycle in an environment for a real maize (Zea mays) F2 population with 233 molecular markers. Note that the frequency distribution of the estimated RGESIM index values approaches the normal distribution.
Frequency distribution of the 244 estimated restricted genomic eigen selection index method (RGESIM) values for two null restrictions on traits grain yield (GY) and EHT and their associated genomic estimated breeding values (GEBVs), GEBVGY and GEBVEHT respectively, for one selection cycle in an environment for a real maize (Zea mays) F2 population with 233 molecular markers. Note that the frequency distribution of the estimated RGESIM index values approaches normal distribution
Now we compare the estimated CRLGSI and RGESIM selection responses and expected genetic gains per trait using the simulated data set described in Sect. 2.8.1 of Chap. 2. We used that data set for four phenotypic selection cycles (C2, C3, C4, and C5), each with four traits (T1, T2, T3, and T4), 500 genotypes, and four replicates per genotype. The economic weights for T1, T2, T3, and T4 were 1, −1, 1, and 1 respectively. For this data set, matrix F was an identity matrix of size 8 × 8 for all four selection cycles.
Columns 2, 3, and 4 (from left to right) of Table 8.4 present the estimated CRLGSI selection responses when their vectors of coefficients are normalized and the estimated RGESIM and selection responses for one, two, and three restrictions for four simulated selection cycles. The averages of the estimated CRLGSI selection responses of the traits and their associated GEBVs for each of the three null restrictions were 3.24 for one restriction, 4.08 for two restrictions, and 5.06 for three restrictions, whereas the averages of the estimated RGESIM selection responses were 3.08 for one restriction, 2.79 for two restrictions, and 3.23 for three restrictions. Note that although for one restriction the selection response was similar for both indices, for two and three restrictions the CRLGSI selection responses were greater than the RGESIM selection responses.
Table 8.5 presents the estimated CRLGSI and RGESIM expected genetic gains per trait for four traits (T1, T2, T3, and T4) and their associated GEBVs (in this case denoted by G1, G2, G3, and G4 to simplify the notation) in four simulated selection cycles and for one, two, and three null restrictions in four simulated selection cycles. Note that the null values of the traits and their restricted GEBVs are not shown in Table 8.5 with the aim of simplifying the table. The averages of the estimated CRLGSI expected genetic gains for the three traits and their associated GEBVs were −2.60, 2.16, 2.84, −1.21, 0.67, and 1.02 for one restriction; 2.74, 3.23, 0.78, and 0.99 for two restrictions; and 4.02 and 1.33 for three restrictions. On the other hand, the averages of the estimated RGESIM expected genetic gains for the three traits and their associated GEBVs were 3.27, −1.67, −1.33, 2.16, −0.92, and −0.84 for one restriction; 3.29, 1.02, 1.76, and 0.46 for two restrictions; and 3.53 and 2.07 for three restrictions. These results indicate that in terms of absolute values, the estimated expected genetic gains for the traits and their associated GEBVs were similar for both indices.
8.5 The Predetermined Proportional Gain Linear Genomic Eigen Selection Index Method
The predetermined proportional gain linear genomic eigen selection index method (PPG-GESIM) theory is based on the predetermined proportional gain linear phenotypic ESIM (PPG-ESIM) described in Chap. 7. In the PPG-ESIM, the vector of PPG (predetermined proportional gain) imposed by the breeder was \( {\mathbf{d}}^{\prime }=\left[{d}_1\kern0.5em {d}_2\kern0.5em \cdots \kern0.5em {d}_r\right] \). However, because the PPG-GESIM uses phenotypic and GEBV information jointly to predict the net genetic merit, the vector of PPG imposed by the breeder (dPG) should be twice the standard vector d′, that is, \( {\mathbf{d}}_{PG}^{\prime }=\left[{d}_1\kern0.5em {d}_2\kern0.5em \cdots \kern0.5em \begin{array}{cccc}{d}_r& {d}_{r+1}& {d}_{r+2}& \begin{array}{cc}\cdots & {d}_{2r}\end{array}\end{array}\right] \), where we would expect that if d1 is the PPG imposed on trait 1, then dr + 1 should be the PPG imposed on the GEBV associated with trait 1, etc. Thus, in the PPG-GESIM we have three possible options for determining (for each trait and GEBV) the PPG: e.g., for trait 1, d1 = dr + 1, d1 > dr + 1 or d1 < dr + 1. This is the main difference between the standard PPG-ESIM described in Chap. 7 and the PPG-GESIM.
8.5.1 The PPG-GESIM Parameters
Using the same procedure described for RGESIM and PPG-ESIM, the PPG-GESIM vector of coefficients (βPG), which maximizes the PPG-GESIM selection response and the expected genetic gain per trait, is the first eigenvector of the following equation
where TPG = KRGΦ−1A + B, KPG = [I2t − QRG], I2t is an identity matrix of size 2t × 2t, \( {\mathbf{Q}}_{RG}={\boldsymbol{\Phi}}^{-1}{\mathbf{A}\mathbf{U}}_G{\left({\mathbf{U}}_G^{\prime }{\mathbf{A}\boldsymbol{\Phi}}^{-1}{\mathbf{A}\mathbf{U}}_G\right)}^{-1}{\mathbf{U}}_G^{\prime}\mathbf{A} \), B = δφ′, \( \boldsymbol{\updelta} ={\boldsymbol{\Phi}}^{-1}{\mathbf{A}\mathbf{U}}_G{\left({\mathbf{U}}_G^{\prime }{\mathbf{A}\boldsymbol{\Phi}}^{-1}{\mathbf{A}\mathbf{U}}_G\right)}^{-1}{\mathbf{d}}_{PG} \), and \( {\varphi}^{\prime }=\frac{{\mathbf{d}}_{PG}^{\prime }{\left({\mathbf{U}}_G^{\prime }{\mathbf{A}\boldsymbol{\Phi}}^{-1}{\mathbf{A}\mathbf{U}}_G\right)}^{-1}{\mathbf{U}}_G^{\prime }{\mathbf{A}\boldsymbol{\Phi}}^{-1}\mathbf{A}}{{\mathbf{d}}_{PG}^{\prime }{\left({\mathbf{U}}_G^{\prime }{\mathbf{A}\boldsymbol{\Phi}}^{-1}{\mathbf{A}\mathbf{U}}_G\right)}^{-1}{\mathbf{d}}_{PG}} \). When B is a null matrix, TPG = KRGΦ−1A (matrix of the RGESIM), and when \( {\mathbf{U}}_G^{\prime } \) is a null matrix, TPG = Φ−1A (matrix of the GESIM); this means that the PPG-GESIM includes the RGESIM and GESIM as particular cases. The optimized PPG-GESIM index can be written as \( {I}_{PG}={\boldsymbol{\upbeta}}_{PG}^{\prime}\mathbf{f} \).
The vector of coefficients of \( H={\mathbf{w}}_{PG}^{\prime}\boldsymbol{\upalpha} \) in the PPG-GESIM can be written as
where \( {\mathbf{Q}}_{PG}^{\prime }={\mathbf{A}\mathbf{U}}_G{\mathbf{D}}_G{\left({\mathbf{D}}_G^{\prime }{\mathbf{U}}_G^{\prime }{\mathbf{A}\boldsymbol{\Phi}}^{-1}{\mathbf{A}\mathbf{U}}_G{\mathbf{D}}_G\right)}^{-1}{\mathbf{D}}_G^{\prime }{\mathbf{U}}_G^{\prime }{\mathbf{A}\boldsymbol{\Phi}}^{-1} \), and \( {\mathbf{D}}_G^{\prime }=\left[\begin{array}{lllll}{d}_{2r}& 0& \cdots & 0& -{d}_1\\ {}0& {d}_{2r}& \cdots & 0& -{d}_2\\ {}\vdots & \vdots & \ddots & \vdots & \vdots \\ {}0& 0& \cdots & {d}_{2r}& -{d}_{2r-1}\end{array}\right] \). Similar to RGESIM, it can be shown that the covariance between \( {H}_{RG}={\mathbf{w}}_{PG}^{\prime}\boldsymbol{\upalpha} \) and \( {I}_{PG}={\boldsymbol{\upbeta}}_{PG}^{\prime}\mathbf{f} \) (\( {\sigma}_{H_{PG}{I}_{PG}} \)) is equal to the variance of \( {I}_{PG}={\boldsymbol{\upbeta}}_{PG}^{\prime}\mathbf{f} \) (\( {\sigma}_{I_{PG}}^2={\boldsymbol{\upbeta}}_{PG}^{\prime }{\boldsymbol{\Phi} \boldsymbol{\upbeta}}_{PG} \)), that is, \( {\sigma}_{H_{PG}{I}_{PG}}={\mathbf{w}}_{PG}^{\prime }{\mathbf{A}\boldsymbol{\upbeta}}_{PG}={\boldsymbol{\upbeta}}_{PG}^{\prime }{\boldsymbol{\Phi} \boldsymbol{\upbeta}}_{PG}={\sigma}_{I_{PG}}^2 \).
The maximized correlation between IPG and HPG, or PPG-GESIM accuracy, is
where \( {\mathbf{w}}_{PG}^{\prime }{\mathbf{Aw}}_{PG} \) is the variance of HPG. Hereafter, to simplify the notation, we write Eq. (8.47) as λPG.
The maximized selection response and the expected genetic gain per trait of the PPG-GESIM are
and
respectively, where βPG is the first eigenvector of Eq. (8.45).
8.5.2 Numerical Examples
The process for estimating PPG-ESIM parameters is similar to the method described for estimating RGESIM parameters. With a selection intensity of 10% (kI = 1.755), we compare the combined predetermined proportional gain linear genomic selection index (CPPG-LGSI) and PPG-GESIM results using the real maize (Zea mays) F2 population with 244 genotypes, 233 molecular markers, and three traits—GY (ton ha−1), EHT (cm), and PHT (cm)—where \( \widehat{\mathbf{P}}=\left[\begin{array}{lll}0.45& 1.33& 2.33\\ {}1.33& 65.07& 83.71\\ {}2.33& 83.71& 165.99\end{array}\right] \), \( \widehat{\mathbf{G}}=\left[\begin{array}{lll}0.07& 0.61& 1.06\\ {}0.61& 17.93& 22.75\\ {}1.06& 22.75& 44.53\end{array}\right] \) and \( \widehat{\boldsymbol{\Gamma}}=\left[\begin{array}{lll}0.07& 0.65& 1.05\\ {}0.65& 10.62& 14.25\\ {}1.05& 14.25& 26.37\end{array}\right] \) are the estimated matrices of P, G, and Γ respectively, whereas \( {\mathbf{w}}^{\prime }=\left[5\kern0.5em -0.1\kern0.5em -0.1\kern0.5em 0\kern0.5em 0\kern0.5em 0\right] \) was the vector of economic weights.
The estimated CPPG-LGSI vector of coefficients was \( {\widehat{\boldsymbol{\upbeta}}}_{CP}={\widehat{\boldsymbol{\upbeta}}}_{CG}+{\widehat{\theta}}_{CP}\widehat{\boldsymbol{\updelta}} \) (see Chap. 6 for additional details). Let \( \widehat{\mathbf{A}}=\left[\begin{array}{ll}\widehat{\mathbf{G}}& \widehat{\boldsymbol{\Gamma}}\\ {}\widehat{\boldsymbol{\Gamma}}& \widehat{\boldsymbol{\Gamma}}\end{array}\right] \) and \( \widehat{\boldsymbol{\Phi}}=\left[\begin{array}{ll}\widehat{\mathbf{P}}& \widehat{\boldsymbol{\Gamma}}\\ {}\widehat{\boldsymbol{\Gamma}}& \widehat{\boldsymbol{\Gamma}}\end{array}\right] \) be the estimated block matrices and \( {\mathbf{d}}_{PG}^{\prime }=\left[7\kern0.5em -3\kern0.5em 3.5\kern0.5em -1.5\right] \) the vector of PPG imposed by the breeder on the traits GY and EHT, and their associated genomic estimated breeding values (GEBVGY and GEBVEHT), and let \( {\mathbf{U}}_C^{\prime }=\left[\begin{array}{cccccc}1& 0& 0& 0& 0& 0\\ {}0& 1& 0& 0& 0& 0\\ {}0& 0& 0& 1& 0& 0\\ {}0& 0& 0& 0& 1& 0\end{array}\right] \) be the matrix of null restrictions on the CPPG-LGSI and \( {\mathbf{w}}^{\prime }=\left[5\kern0.5em -0.1\kern0.5em -0.1\kern0.5em 0\kern0.5em 0\kern0.5em 0\right] \) the vector of economic weights. It can be shown that \( {\widehat{\uptheta}}_{\mathrm{CP}}=-0.00009 \) is the estimated value of the proportionality constant, \( {\widehat{\boldsymbol{\updelta}}}^{\prime }=\left[-112.92\kern0.5em -72.16\kern0.5em 61.35\kern0.5em 231.79\kern0.5em 64.75\kern0.5em -61.35\right] \), \( {\widehat{\boldsymbol{\upbeta}}}_{CP}^{\prime }=\left[-0.01\kern0.5em 0.01\kern0.5em -0.01\kern0.5em 0.59\kern0.5em 0.09\kern0.5em -0.09\right] \) is the estimated CPPG-LGSI vector of coefficients, and the estimated CPPG-LGSI can be written as
where GEBVGY, GEBVEHT, and GEBVPHT are the GEBVs associated with traits GY, EHT, and PHT respectively. The same procedure is valid for more than two predetermined restrictions. The estimated CPPG-LGSI selection response and expected genetic gain per trait were \( {\widehat{R}}_{CP}={k}_I\sqrt{{\widehat{\boldsymbol{\upbeta}}}_{CP}^{\prime }{\widehat{\boldsymbol{\Phi}}\widehat{\boldsymbol{\upbeta}}}_{CP}}=0.443 \) and \( {\widehat{\mathbf{E}}}_{CP}^{\prime }={k}_I\frac{{\widehat{\boldsymbol{\upbeta}}}_{CP}^{\prime}\widehat{\mathbf{A}}}{\sqrt{{\widehat{\boldsymbol{\upbeta}}}_{CP}^{\prime }{\widehat{\boldsymbol{\Phi}}\widehat{\boldsymbol{\upbeta}}}_{CP}}}=\left[-0.004\kern0.5em 0.002\kern0.5em -4.639\kern0.5em -0.002\kern0.5em 0.001\kern0.5em -4.326\right] \) respectively, whereas the estimated CPPG-LGSI accuracy is \( {\widehat{\rho}}_{HI_{CP}}=\frac{{\widehat{\sigma}}_{I_{CP}}}{{\widehat{\sigma}}_H}=0.234 \).
Because the estimated value of the proportionality constant was negative (\( {\widehat{\uptheta}}_{\mathrm{CP}}=-0.00009 \)), the expected genetic gains of the traits GY and EHT, and their associated genomic estimated breeding values (GEBVGY and GEBVEHT), which appeared in the \( {\widehat{\mathbf{E}}}_{CP}^{\prime } \) values, were not in accordance with the values of the vector of PPG imposed by the breeder, \( {\mathbf{d}}_{PG}^{\prime }=\left[7\kern0.5em -3\kern0.5em 3.5\kern0.5em -1.5\right] \), as we would expect, and CPPG-LGSI accuracy (0.234) was low. These results indicate that in the CPPG-LGSI, it is very important for the estimated values of \( {\widehat{\uptheta}}_{\mathrm{CP}} \) to be positive (see Chaps. 3 and 6 for details).
In the PPG-GESIM, we need to find the solutions to equation \( \left({\widehat{\mathbf{T}}}_{PG}-{\widehat{\lambda}}_{PG_j}^2{\mathbf{I}}_{2t}\right){\widehat{\boldsymbol{\upbeta}}}_{PG_j}=\mathbf{0} \), for \( {\widehat{\lambda}}_{PG_j}^2 \) and \( {\widehat{\boldsymbol{\upbeta}}}_{PG_j} \) (see Eq. 8.45). The estimated PPG-GESIM vector of coefficients was \( {\widehat{\boldsymbol{\upbeta}}}_{PG}^{\prime }=\left[0.001\kern0.5em -0.050\kern0.5em 0.029\kern0.5em 0.975\kern0.5em 0.154\kern0.5em -0.157\right] \), which was transformed using matrix \( \mathbf{F}=\left[\begin{array}{cccccc}-0.1& 0& 0& 0& 0& 0\\ {}0& 3& 0& 0& 0& 0\\ {}0& 0& 2& 0& 0& 0\\ {}0& 0& 0& -1& 0& 0\\ {}0& 0& 0& 0& -1& 0\\ {}0& 0& 0& 0& 0& -1\end{array}\right] \), that is, we changed the direction of the original vector. With the \( {\widehat{\boldsymbol{\upbeta}}}_{PG}^{\prime } \) values, we can estimate the PPG-GESIM index as
where GEBVGY, GEBVEHT, and GEBVPHT are the GEBVs associated with the traits GY, EHT, and PHT respectively. The estimated PPG-GESIM selection response, accuracy, and expected genetic gain per trait were \( {\widehat{R}}_{PG}={k}_I\sqrt{{\widehat{\boldsymbol{\upbeta}}}_{PG}^{\prime }{\widehat{\boldsymbol{\Phi}}\widehat{\boldsymbol{\upbeta}}}_{PG}}=0.696 \), \( {\widehat{\rho}}_{{\widehat{H}}_{PG}{\widehat{I}}_{PG}}=\frac{{\widehat{\sigma}}_{{\widehat{I}}_{PG}}}{{\widehat{\sigma}}_{{\widehat{H}}_{PG}}}=0.843 \), and \( {\widehat{\mathbf{E}}}_{PG}^{\prime }={k}_I\frac{{\widehat{\boldsymbol{\upbeta}}}_{PG}^{\prime}\widehat{\mathbf{A}}}{\sqrt{{\widehat{\boldsymbol{\upbeta}}}_{PG}^{\prime }{\widehat{\boldsymbol{\Phi}}\widehat{\boldsymbol{\upbeta}}}_{PG}}}=\left[0.01\kern0.5em -1.00\kern0.5em -3.56\kern0.5em 0\kern0.5em -0.46\kern0.5em -3.98\right] \) respectively.
Fig. 8.4 presents the frequency distribution of the 244 estimated PPG-GESIM index values for two predetermined restrictions on the traits GY and EHT and their associated GEBVs (GEBVGY and GEBVEHT), for one selection cycle in an environment for a real maize (Zea mays) F2 population with 233 molecular markers. Note that the frequency distribution of the estimated PPG-GESIM index values approaches normal distribution.
Frequency distribution of the 244 estimated predetermined proportional gain genomic eigen selection index method (PPG-GESIM) values for two predetermined restrictions on the traits GY and EHT and their associated GEBVs, GEBVGY and GEBVEHT, for one selection cycle in an environment for a real maize (Zea mays) F2 population with 233 molecular markers
Now, with a selection intensity of 10% (kI = 1.755) and a vector of predetermined restrictions \( {\mathbf{d}}_{PG}^{\prime }=\left[7\kern0.5em -3\kern0.5em 5\kern0.5em 3.5\kern0.5em -1.5\kern0.5em 2.5\right] \), we compare the estimated CPPG-LGSI and PPG-GESIM selection responses and expected genetic gains per trait using the simulated data set described in Sect. 2.8.1 of Chap. 2. Traits T1, T2, and T3 and their associated GEBVs (GEBV1, GEBV2, and GEBV3 respectively) were restricted, but trait T4 and its associated GEBV4 were not restricted. For this data set, matrix F was an identity matrix of size 8 × 8 for all four selection cycles.
Table 8.6 presents the estimated CPPG-LGSI selection responses when their vectors of coefficients are normalized, and the estimated PPG-GESIM selection responses for one, two, and three predetermined restrictions for four simulated selection cycles. The averages of the estimated CPPG-LGSI selection responses were 5.08 for one restriction, 3.42 for two restrictions, and 1.60 for three restrictions, whereas the averages of the estimated PPG-GESIM selection responses were 1.96 for one restriction, 4.14 for two restrictions, and 5.46 for three restrictions. For this data set, when the number of restrictions increases, the estimated CPPG-LGSI selection response tends to decrease, whereas the estimated PPG-GESIM selection response increases.
Tables 8.7 presents the estimated CPPG-LGSI and PPG-GESIM expected genetic gains for one, two, and three predetermined restrictions respectively, for four simulated selection cycles. The averages of the estimated CPPG-LGSI expected genetic gains for the four traits and their four associated GEBVs were 8.28, −4.12, 3.23, 2.23, 4.14, −2.26, 1.71, and 1.01 for one restriction; 8.43, −3.61, 3.28, 2.13, 4.22, −1.81, 1.72, and 0.93 for two restrictions; and 5.81, −2.49, 4.15, 2.26, 2.90, −1.24, 2.07, and 0.89 for three restrictions. On the other hand, the averages of the estimated PPG-GESIM expected genetic gains for the four traits and their four associated GEBVs were 6.97, −1.31, 1.78, 0.52, 5.64, −1.74, 1.75, and 0.58 for one restriction; 6.93, −2.73, 1.29, 0.85, 5.75, −2.55, 1.49, and 0.79 for two restrictions, and 8.12, −3.27, 2.99, 1.13, 2.19, −1.15, 1.30, and 0.45 for three restrictions. These results indicate that the estimated CPPG-LGSI expected genetic gains for the four traits and their four associated GEBVs were generally higher than the estimated PPG-GESIM expected genetic gains for the four traits and their four associated GEBVs.
References
Crossa J, Cerón-Rojas JJ (2011) Multi-trait multi-environment genome-wide molecular marker selection indices. J Indian Soc Agric Stat 62(2):125–142
Meyer CD (2000) Matrix analysis and applied linear algebra. Society for Industrial and Applied Mathematics (SIAM), Philadelphia, PA
Author information
Authors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2018 The Author(s)
About this chapter
Cite this chapter
Céron-Rojas, J.J., Crossa, J. (2018). Linear Molecular and Genomic Eigen Selection Index Methods. In: Linear Selection Indices in Modern Plant Breeding. Springer, Cham. https://doi.org/10.1007/978-3-319-91223-3_8
Download citation
DOI: https://doi.org/10.1007/978-3-319-91223-3_8
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-91222-6
Online ISBN: 978-3-319-91223-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)