Abstract
Scholia is a tool to handle scientific bibliographic information through Wikidata. The Scholia Web service creates on-the-fly scholarly profiles for researchers, organizations, journals, publishers, individual scholarly works, and for research topics. To collect the data, it queries the SPARQL-based Wikidata Query Service. Among several display formats available in Scholia are lists of publications for individual researchers and organizations, plots of publications per year, employment timelines, as well as co-author and topic networks and citation graphs. The Python package implementing the Web service is also able to format Wikidata bibliographic entries for use in LaTeX/BIBTeX. Apart from detailing Scholia, we describe how Wikidata has been used for bibliographic information and we also provide some scientometric statistics on this information.
You have full access to this open access chapter, Download conference paper PDF
Similar content being viewed by others
Keywords
- Wikidata
- Wikipedias
- Functional Requirements For Bibliographic Records (FRBR)
- Public Library Of Science (PLOS)
- National Institute For Occupational Safety And Health (NIOSH)
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 Introduction
Wikipedia contains significant amounts of data relevant for scientometrics, and it has formed the basis for several scientometric studies [4, 14, 15, 17, 18, 20, 21, 28, 29, 34, 39]. Such studies can use the structured references found in Wikipedia articles or use the intrawiki hyperlinks, e.g., to compare citations from Wikipedia to scholarly journals with Thomson Reuters journal citation statistics as in [20] or to rank universities as in [39].
While many Wikipedia pages have numerous references to scientific articles, the current Wikipedias have very few entries about specific scientific articles. This is most evident when browsing the Academic journal articles category on the English Wikipedia.Footnote 1 Among the few items in that category are famed papers such as the 1948 physics paper The Origin of Chemical Elements [2] – described in the English Wikipedia article Alpher–Bethe–Gamow paperFootnote 2 – as well as the 1953 article Molecular Structure of Nucleic Acids: A Structure for Deoxyribose Nucleic Acid [37] on eight Wikipedias. Another scientific article is Hillary Putnam’s Is Semantics Possible? [31]Footnote 3 from 1970 on the Estonian Wikipedia.
References in Wikipedia are often formatted in templates, and it takes some effort to extract and match information in the template fields. For instance, in a study of journals cited on Wikipedia, a database was built containing journal name variations to match the many different variations that Wikipedia editors used when citing scientific articles [20]. The use of standard identifiers — such as the Digital Object Identifier (DOI) — in citations on Wikipedia can help to some extent to uniquely identify works and journals.
Several other wikis have been set up to describe scientific articles, such as WikiPapers, AcaWiki, Wikilit [25] and Brede Wiki [22].Footnote 4 They are all examples of MediaWiki-based wikis that primarily describe scientific articles. Three of them use the Semantic MediaWiki extension [16], while the fourth uses MediaWiki’s template functionalityFootnote 5 to structure bibliographic information.
Since the launch of WikidataFootnote 6 [36], the Wikimedia family includes a platform to better handle structured data such as bibliographic data and to enforce input validation to a greater degree than Wikipedia. Wikidata data can be reified to triples [5, 9], and RDF/graph-oriented databases, including SPARQL databases, can represent Wikidata data [10]. The Wikidata Query Service (WDQS)Footnote 7 is an extended SPARQL endpoint that exposes the Wikidata data. Apart from offering a SPARQL endpoint, it also features an editor and a variety of frontend result display options. It may render the SPARQL query result as, e.g., bubble charts, line charts, graphs, timelines, list of images, points on a geographical map, or just provide the result as a table. These results can also be embedded on other Web pages via an HTML iframe element. We note that Wikidata is open data published under the Public Domain Dedication and Waiver (CC0),Footnote 8 and that it is available not only through the SPARQL endpoint, but also as Linked Data FragmentsFootnote 9 [35] and—like any other project of the Wikimedia family—through an API and dump files.Footnote 10
In the following sections, we describe how Wikidata has been used for bibliographic information, some statistics on it and present Scholia, our website built to expose such information. We furthermore show how Scholia can be used for bibliography generation and discuss limitations and advantages with Wikidata and Scholia.
2 Bibliographic Information on Wikidata
Wikidata editors have begun to systematically add scientific bibliographic data to Wikidata across a broad range of scientific domains — see Table 1 for a summary of Wikidata as a digital library. Individual researchers and scientific articles not described by their own Wikipedia article in any language are routinely added to Wikidata, and we have so far experienced very few deletions of such data in reference to a notability criterion. The current interest in expanding bibliographic information on Wikidata has been boosted by the WikiCite project, which aims at collecting bibliographic information in Wikidata and held its first workshop in 2016 [33].
The bibliographic information collected on Wikidata is about books, articles (including preprints), authors, organizations, journals, publishers and more. These items (corresponding to subject in Semantic Web parlance) can be interlinked through Wikidata properties (corresponding to the predicate), such as author (P50),Footnote 11 published in (P1433), publisher (P123), series (P179), main theme (P921), educated at (P69), employer (P108), part of (P361), sponsor (P859, can be used for funding), cites (P2860) and several other properties.Footnote 12
Numerous properties exist on Wikidata for deep linking to external resources, e.g., for DOI, PMID, PMCID, arXiv, ORCID, Google Scholar, VIAF, Crossref funder ID, ZooBank and Twitter. With these many identifiers, Wikidata can act as a hub for scientometrics studies between resources. If no dedicated Wikidata property exists for a resource, one of the URL properties can work as a substitute for creating a deep link to a resource. For instance, P1325 (external data available at) can point to raw or supplementary data associated with a paper. We have used this scheme for scientific articles associated with datasets stored in OpenfMRI [27], an online database with raw brain measurements, mostly from functional magnetic resonance imaging studies. Using WDQS, we query the set of OpenfMRI-linked items using the following query:

A similar scheme is used for a few of the scientific articles associated with data in the neuroinformatics databases Neurosynth [38] and NeuroVault [6].
When bibliographic items exist in Wikidata, they can be used as references to support claims (corresponding to triplets with extra qualifiers) in other items of Wikidata, e.g., a biological claim can be linked to the Wikidata item for a scientific journal.
By using these properties systematically according to an emerging data model,Footnote 13 editors have extended the bibliographic information in Wikidata. Particularly instrumental in this process was a set of tools built by Magnus Manske, QuickStatementsFootnote 14 and Source MetaData,Footnote 15 including the latter’s associated Resolve authors toolFootnote 16 as well as the WikidataIntegratorFootnote 17 associated with the Gene Wiki project [30] and the fatameh toolFootnote 18 based on it. Information can be extracted from, e.g., PubMed, PubMed Central and arXiv and added to Wikidata.
How complete is Wikidata in relation to scientific bibliographic information? Journals and universities are well represented. For instance, 31,902 Wikidata items are linked with their identifier for the Collections of the National Library of Medicine (P1055). This number can be obtained with the following WDQS SPARQL query:

Far less covered are individual articles, individual researchers, university departments and citations between scientific articles. Most of the scientific articles in Wikidata are claimed to be an instance of (P31) the Wikidata item scientific article (Q13442814). With a WDQS query, we can count the number of Wikidata items linked this way to scientific article:

As of 2 August 2017, the query returned the result 2,380,009, see also Tables 1 and 2 (the number of scientific articles has grown considerable since the end of July 2017). In comparison, arXiv states having 1,289,564 e-prints and ACM Digital Library states having 24,668 proceedings.Footnote 19 In 2014, a capture/recapture method estimated the number of scholarly English-language documents on the public web to be “at least 114 million” [13], while researchers found 87,542,370 DOIs in the Crossref database as of 21 March 2017 [32], thus Wikidata currently records only a minor part of all scientific articles. There were 16,327 authors associated with Wikidata items linked through the author property (P50) to items that are instance of scientific article:

The number of citations as counted by triples using the P2860 (cites) property stood at 3,379,786:

The completeness can be fairly uneven. Articles from Public Library of Science (PLOS) journals are much better represented than articles from the journals of IEEE. On 9 August 2017, we counted 160,676 works published in PLOS journals with this WDQS query,

while the equivalent for IEEE (Q131566) only returns 4,595. Note that 160,676 PLOS articles are far more than the 4,553 PLOS articles reported back in 2014 as cited from the 25 largest Wikipedias [17], thus Wikidata has a much better coverage here than Wikipedia.
Given that Wikidata only has around 3.4 million P2860-citations, it is no surprise that the current number of citations is considerable less than the citation counts one finds in other web services, — even for authors with a large part of their published scientific articles listed in Wikidata. Table 3 shows h-index statistics for three such authors. The Wikidata count has been established by WDQS queries similar to the following:

Even for these well-covered researchers, the h-index based on P2860-citations in Wikidata is around two to three times lower than the h-indices obtained with other services.
The sponsor property (P859) has been used extensively for research funded by the National Institute for Occupational Safety and Health (NIOSH), with 52,852 works linking to the organization, 18,135 of which are instance of scientific articles, but apart from NIOSH, the use of the property has been very limited for scientific articles.Footnote 20
3 Scholia
Overview screenshot of part of the Scholia Web page for an author: https://tools.wmflabs.org/scholia/author/Q20980928. Fig. 2 zooms in on one panel.
Scholia provides both a Python package and a Web service for presenting and interacting with scientific information from Wikidata. The code is available via https://github.com/fnielsen/scholia, and a first release has been archived in Zenodo [23].
As a Web service, its canonical site runs from the Wikimedia Foundation-provided service Wikimedia Toolforge (formerly called Wikimedia Tool Labs) at https://tools.wmflabs.org/scholia/, but the Scholia package may be downloaded and run from a local server as well. Scholia uses the Flask Python Web framework [7].
The current Web service relies almost entirely on Wikidata for its presented data. The frontend consists mostly of HTML iframe elements for embedding the on-the-fly-generated WDQS results and uses many of the different output formats from this service: bubble charts, bar charts, line charts, graphs and image lists.
Initially, we used the table output from WDQS to render tables in Scholia, but as links in WDQS tables link back to Wikidata items — and not Scholia items — we have switched to using the DataTablesFootnote 21 Javascript library.
Through a JavaScript-based query to the MediaWiki API, an excerpt from the English Wikipedia is shown on the top of each Scholia page if the corresponding Wikidata item is associated with an article in the English Wikipedia. The label for the item is fetched via Wikidata’s MediaWiki API. While some other information can be fetched this way, Scholia’s many aggregation queries are better handled through SPARQL.
Scholia uses the Wikidata item identifier as its identifier rather than author name, journal titles, etc. A search field on the front page provides a Scholia user with the ability to search for a name to retrieve the relevant Wikidata identifier. To display items, Scholia sets up a number of what we call “aspects”. The currently implemented aspects (see Table 4) are author, work, organization, venue, series, publisher, sponsor, award, topic, disease, protein, chemical and (biological) pathway.
The present selection was motivated by the possibilities inherent in the Wikidata items and properties. We plan to extend this to further aspects. A URL scheme distinguishes the different aspects, so the URL path /scholia/author/Q6365492 will show the author aspect of the statistician Kanti V. Mardia, while /scholia/topic/Q6365492 will show the topic aspect of the person, i.e., articles about Mardia.
Likewise, universities can be viewed, for instance, as organizations or as sponsors. Indeed, any Wikidata item can be viewed in any Scholia aspect, but Scholia can show no data if the user selects a “wrong” aspect, i.e. one for which no relevant data is available in Wikidata.
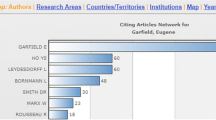
For each aspect, we make multiple WDQS queries based on the Wikidata item for which the results in the panels are displayed. Plots are embedded with HTML iframes. For the author aspect, Scholia queries WDQS for the list of publications, showing the result in a table, displaying a bar chart of the number of publications per year, number of pages per year, venue statistics, co-author graph, topics of the published works (based on the “main theme” property), associated images, education and employment history as timelines, academic tree, map with locations associated with the author, and citation statistics – see Fig. 1 for an example of part of an author aspect page. The citation statistics displays the most cited work, citations by year and citing authors. For the academic tree, we make use of Blazegraph’s graph analytics RDF GAS APIFootnote 22 that is available in WDQS.
The embedded WDQS results link back to WDQS, where a user can modify the query. The interactive editor of WDQS allows users not familiar with SPARQL to make simple modifications without directly editing the SPARQL code.
Related to their work on quantifying conceptual novelty in the biomedical literature [19], Shubhanshu Mishra and Vetle Torvik have set up a website profiling authors in PubMed datasets: LEGOLAS.Footnote 23 Among other information, the website shows the number of articles per year, the number of citations per year, the number of self-citations per year, unique collaborations per year and NIH grants per year as bar charts that are color-coded according to, e.g., author role (first, solo, middle or last author). Scholia uses WDQS for LEGOLAS-like plots. Figure 2 displays one such example for the number of published items as a function of year of publication on an author aspect page, where the components of the bars are color-coded according to author role.
Screenshot of Scholia Web page with the number of papers published per year for Finn Årup Nielsen: https://tools.wmflabs.org/scholia/author/Q20980928. Inspired by LEGOLAS. Colors indicate author role: first, middle, last or solo author. (Color figure online)
Scholia screenshot with page production for a research section (Cognitive Systems at the Technical University of Denmark), where the number of pages per paper has been normalized by the number of authors. The bars are color-coded according to author. The plot is heavily biased, as only a very limited subset of papers from the section is available in Wikidata, and the property for the number of pages is set for only a subset of these papers. From https://tools.wmflabs.org/scholia/organization/Q24283660. (Color figure online)
For the organization aspect, Scholia uses the employer and affiliated Wikidata properties to identify associated authors, and combines this with the author query for works. Scholia formulates SPARQL queries with property paths to identify suborganizations of the queried organization, such that authors affiliated with a suborganization are associated with the queried organization. Figure 3 shows a corresponding bar chart, again inspired by the LEGOLAS style. Here, the Cognitive Systems section at the Technical University of Denmark is displayed with the organization aspect. It combines work and author data. The bar chart uses the P1104 (number of pages) Wikidata property together with a normalization based on the number of authors on each of the work items. The bars are color-coded according to individual authors associated with the organization. In this case, the plot is heavily biased, as only a very limited subset of publications from the organization is currently present in Wikidata, and even the available publications may not have the P1104 property set. Other panels shown in the organization aspect are a co-author graph, a list of recent publications formatted in a table, a bubble chart with most cited papers with affiliated first author and a bar chart with co-author-normalized citations per year. This last panel counts the number of citations to each work and divides it by the number of authors on the cited work, then groups the publications according to year and color-codes the bars according to author.
Screenshot from Scholia’s publisher aspect with number of publications versus number of citations for works published by BioMed Central. The upper right point with many citations and many published works is the journal Genome Biology.
For the publisher aspect, Scholia queries all items where the P123 property (publisher) has been set. With these items at hand, Scholia can create lists of venues (journals or proceedings) ordered according to the number of works (papers) published in each of them, as well as lists of works ordered according to citations. Figure 4 shows an example of a panel on the publisher aspect page with a scatter plot detailing journals from BioMed Central. The position of each journal in the plot reveals impact factor-like information.

For the work aspect, Scholia lists citations and produces a partial citation graph. Figure 5 shows a screenshot of the citation graph panel from the work aspect for a specific article [3]. For this aspect, we also formulate a special query to return a table with a list of Wikidata items where the given work is used as a source for claims. An example query for a specific work is shown with Listing 1. From the query results, it can be seen, for instance, that the article A novel family of mammalian taste receptors [1] supports a claim about Taste 2 receptor member 16 (Q7669366) being present in the cell component (P681) integral component of membrane (Q14327652). For the topic aspect, Scholia uses a property path SPARQL query to identify subtopics.
For a given item where the aspect is not known in advance, Scholia tries to guess the relevant aspect by looking at the instance of property. The Scholia Web service uses that guess for redirecting, so for instance, /scholia/Q8219 will redirect to /scholia/author/Q8219, the author aspect for the psychologist Uta Frith. This is achieved by first making a server site query to establish that Uta Frith is a human and then using that information to choose the author aspect as the most relevant aspect to show information about Uta Frith.
We have implemented a few aspects that are able to display information from two or more specified Wikidata items. For instance, /scholia/organizations/Q1269766,Q193196 displays information from University College London and Technical University of Denmark. One panel lists coauthorships between authors affiliated with the two organizations. Another panel shows a “Works per year” plot for the specified organizations, see Fig. 6. Likewise, an address such as /scholia/authors/Q20980928,Q24290415,Q24390693,Q26720269 displays panels for 4 different authors. With the graph queries in BlazeGraph, Scholia shows co-author paths between multiple authors in a graph plot. Figure 7 shows the co-author path between Paul Erdős and Natalie Portman, which can give an estimate of Portman’s Erdős-number (i.e., the number of coauthorships between a given author and Erdős).
A few redirects for external identifiers are also implemented. For instance, with Uta Frith’s Twitter name ‘utafrith’, /scholia/twitter/utafrith will redirect to /scholia/Q8219, which in turn will redirect to /scholia/author/Q8219. Scholia implements similar functionality for DOI, ORCID, GitHub user identifier as well as for the InChIKey [8] and CAS chemical identifiers.
Screenshot of part of a Scholia Web page at https://tools.wmflabs.org/scholia/work/Q21143764 with the partial citation graph panel of the work aspect for Johan Bollen’s article from 2009 [3].
A co-author path between Paul Erdős and Natalie Portman (Natalie Hershlag) on the page for multiple authors https://tools.wmflabs.org/scholia/authors/Q37876,Q173746.
Aggregation on science awards with respect to gender from the award aspect index page at https://tools.wmflabs.org/scholia/award/ with number of male recipients on the x-axis and number of female recipients on the y-axis.
For the index page for the award aspect, we have an aggregated plot for all science awards with respect to gender, see Fig. 8. The plot gives an overview of awards predominantly given to men (awards close to the x-axis) or predominantly given to women (awards close to the y-axis).
4 Using Wikidata as a Bibliographic Resource
As a command-line tool, Scholia provides a prototype tool that uses Wikidata and its bibliographic data in a  and
and  environment. The current implementation looks up citations in the LaTeX-generated .aux file and queries Wikidata’s MediaWiki API to get cited Wikidata items. The retrieved items are formatted and written to a .bib that bibtex can use to format the bibliographic items for inclusion in the
environment. The current implementation looks up citations in the LaTeX-generated .aux file and queries Wikidata’s MediaWiki API to get cited Wikidata items. The retrieved items are formatted and written to a .bib that bibtex can use to format the bibliographic items for inclusion in the  document. The workflow for a
document. The workflow for a  document with the filename example.tex is
document with the filename example.tex is

Here, the example document could read

In this case, the  command cites Q18507561 (Wikidata: a free collaborative knowledgebase [36]). A DOI can also be used in the
command cites Q18507561 (Wikidata: a free collaborative knowledgebase [36]). A DOI can also be used in the  command: instead of writing
command: instead of writing  , one may write
, one may write  to get the same citation. Scholia matches on the “10.” DOI prefix and makes a SPARQL query to get the relevant Wikidata item.
to get the same citation. Scholia matches on the “10.” DOI prefix and makes a SPARQL query to get the relevant Wikidata item.
The scheme presented above can take advantage of the many available style files of  to format the bibliographic items in the various ways requested by publishers. We have used Scholia for reference management in this paper. This means that all cited papers in this paper are entered in Wikidata.
to format the bibliographic items in the various ways requested by publishers. We have used Scholia for reference management in this paper. This means that all cited papers in this paper are entered in Wikidata.
There are various issues with the translation. Though planned to support UTF-8 encoding at least since 2003 [26], as of 2017,  does not support UTF-8 completely. The problem results in wrong sorting of the bibliographic items as well as wrong extraction of the surname, e.g., “Finn Årup Nielsen” gets extracted as “Årup Nielsen, Finn” instead of “Nielsen, Finn Årup” and sorted among the last items in the bibliography rather than under “N”. A workaround could convert UTF-8 encoded characters to
does not support UTF-8 completely. The problem results in wrong sorting of the bibliographic items as well as wrong extraction of the surname, e.g., “Finn Årup Nielsen” gets extracted as “Årup Nielsen, Finn” instead of “Nielsen, Finn Årup” and sorted among the last items in the bibliography rather than under “N”. A workaround could convert UTF-8 encoded characters to  escapes. A small translation table can handle accented characters, but miss, e.g., non-ASCII non-accented characters like ø, æ, å, ð and Ð. The combination of Biblatex/Biber can handle UTF-8, but required style files might not be available. The current Scholia implementation has a very small translation table to handle a couple of non-ASCII UTF-8 characters that occur in names.
escapes. A small translation table can handle accented characters, but miss, e.g., non-ASCII non-accented characters like ø, æ, å, ð and Ð. The combination of Biblatex/Biber can handle UTF-8, but required style files might not be available. The current Scholia implementation has a very small translation table to handle a couple of non-ASCII UTF-8 characters that occur in names.
5 Discussion
WDQS and Scholia can provide many different scientometrics views of the data available in Wikidata. The bibliographic data in Wikidata are still quite limited, but the number of scientometrically relevant items will likely continue to grow considerably in the coming months and years.
The continued growth of science data on Wikidata can have negative impact on Scholia, making the on-the-fly queries too resource demanding. In the current version, there are already a few queries that run into WDQS’s time out, e.g., it happens for the view of co-author-normalized citations per year for Harvard University. If this becomes a general problem, we will need to redefine the queries. Indeed, the WDQS time out will be a general problem if we want to perform large-scale scientometrics studies. An alternative to using live queries would be to use dumps, which are available in several formats on a weekly basis, with daily increments in between.Footnote 24 The problem is not a limitation of SPARQL, but a limitation set by the server resources. Some queries may be optimized, especially around the item labeling.
Working with Scholia has made us aware of several issues. Some of these are minor limitations in the Wikidata and WDQS systems. The Wikidata label length is limited to 250 characters, whereas the ‘monolingual text’ datatype used for the ‘title’ property (P1476) is limited to 400 characters. There are scholarly articles with titles longer than those limits.
Wikidata fields cannot directly handle subscripts and superscripts, which commonly appear in titles of articles about chemical compounds, elementary particles or mathematical formulas. Other formatting in titles cannot directly be handled in Wikidata’s title property,Footnote 25 and recording a date such as “Summer 2011” is difficult.
Title and names of items can change. Authors can change name, e.g. due to marriage, and journals can change titles, e.g. due to a change of scope or transfer of ownership. For instance, the Journal of the Association for Information Science and Technology has changed name several times over the years.Footnote 26 Wikidata can handle multiple titles in a single Wikidata item and with qualifiers describe the dates of changes in title. For scientometrics, this ability is an advantage in principle, but multiple titles can make it cumbersome to handle when Wikidata is used as a bibliographic resource in document preparation, particularly for articles published near the time when the journal changed its name. One way to alleviate this problem would be to split the journal’s Wikidata item into several, but this is not current practice.
In Wikidata, papers are usually not described to be affiliated with organizations. Scholia’s ability to make statistics on scientific articles published by an organization is facilitated by the fact that items about scientific articles can link to items about authors, which can link to items about organizations. It is possible to link scientific articles to organization directly by using Wikidata qualifiers in connection with the author property. However, this scheme is currently in limited use. This scarcity of direct affiliation annotation on Wikidata items about articles means that scientometrics on the organizational level are unlikely to be precise at present. In the current version, Scholia even ignores any temporal qualifier for the affiliation and employer property, meaning that a researcher moving between several organization gets his/her articles counted under multiple organizations.
Data modeling on Wikidata gives rise to reflections on what precisely a “publisher” and a “work” is. A user can set the publisher Wikidata property of a work to a corporate group, a subsidiary or possibly an imprint. For instance, how should we handle Springer Nature, BioMed Central and Humana Press?
Functional Requirements for Bibliographic Records (FRBR) [12] suggests a scheme for works, expressions, manifestations and “items”. In Wikipedia, most items are described on the work level as opposed to the manifestation level (e.g., book edition), while citations should usually go to the manifestation level. How should one deal with scientific articles that have slightly different “manifestations”, such as preprint, electronic journal edition, paper edition and postprint, or editorials that were co-published in multiple journals with identical texts? An electronic and a paper edition may differ in their dates of publication, but otherwise have the same bibliographic data, while a preprint and its journal edition usually have different identifiers and may also differ in content. From a scientometrics point of view, these difference in manifestation may not matter in some cases, but could be the focus of others. Splitting a scientific article as a work (in the FRBR sense) over multiple Wikidata items seems only to complicate matters.
The initial idea for Scholia was to create a researcher profile based on Wikidata data with list of publications, picture and CV-like information. The inspiration came from a blog post by Lambert Heller: What will the scholarly profile page of the future look like? Provision of metadata is enabling experimentation.Footnote 27 In this blog post, he discussed the different features of several scholarly Web services: ORCID, ResearchGate, Mendeley, Pure, VIVO, Google Scholar and ImpactStory. In Table 5, we have set up a table listing Heller’s features for the Wikidata–Scholia combination. Wikidata–Scholia performs well in most aspects, but in the current version, Scholia has no backend for storing user data, and user features such as forum, Q&A and followers are not available.
Beyond the features listed by Heller, which features set Wikidata–Scholia apart from other scholarly Web services? The collaborative nature of Wikidata means that Wikidata users can create items for authors that do not have an account on Wikidata. In most other systems, the researcher as a user of the system has control over his/her scholarly profile and other researchers/users cannot make amendment or corrections. Likewise, when one user changes an existing item, this change will be reflected in subsequent live queries of that item, and it may still be in future dumps if not reverted or otherwise modified before the dump creation.
With WDQS queries, Scholia can combine data from different types of items in Wikidata in a way that is not usually possible with other scholarly profile Web services. For instance, Scholia generates lists of publications for an organization by combining items for works and authors and can show co-author graphs restricted by affiliation. Similarly, the co-author graph can be restricted to authors publishing works annotated with a specific main theme. Authors are typically annotated with gender in Wikidata, so Scholia can show gender color-coding of co-author graphs. On the topic aspect page, the Scholia panel that shows the most cited works that are cited from works around the topic can point to an important paper for a topic – even if the paper has not been annotated with the topic – by combining the citations data and topic annotation. References for claims are an important part of Wikidata and also singles Wikidata out among other scholarly profile Web service, and it acts as an extra scientometrics dimension. The current version of Scholia has only a few panels where the query uses references, e.g., the “Supports the following statement(s)” on the work aspect page, but it is possible to extend the use of this scientometrics dimension.
Notes
- 1.
- 2.
- 3.
- 4.
- 5.
- 6.
- 7.
- 8.
- 9.
- 10.
The API is at https://www.wikidata.org/w/api.php, and the dump files are available at https://www.wikidata.org/w/api.php.
- 11.
The URI for Wikidata property P50 is http://www.wikidata.org/prop/direct/P50 or with the conventional prefix wdt:P50. Similarly for any other Wikidata property.
- 12.
A Wikidata table lists properties that are commonly used in bibliographic contexts: https://www.wikidata.org/wiki/Template:Bibliographical_properties.
- 13.
- 14.
- 15.
- 16.
- 17.
- 18.
https://tools.wmflabs.org/fatameh/ with documentation available at https://www.wikidata.org/wiki/Wikidata:WikiProject_Source_MetaData/fatameh.
- 19.
As of 2 August 2017 according to https://arxiv.org/ and https://dl.acm.org/contents_guide.cfm.
- 20.
National Institute for Occupational Safety and Health has a Wikimedian-in-Residence program, through which James Hare has added many of the NIOSH works.
- 21.
- 22.
- 23.
- 24.
- 25.
By way of an example, consider the article “A library of 7TM receptor C-terminal tails. Interactions with the proposed post-endocytic sorting proteins ERM-binding phosphoprotein 50 (EBP50), N-ethylmaleimide-sensitive factor (NSF), sorting nexin 1 (SNX1), and G protein-coupled receptor-associated sorting protein (GASP)”, another article with the title “Cerebral 5-HT\(_\mathrm{2A}\) receptor binding is increased in patients with Tourette’s syndrome”, where “2A” is subscripted, and “User’s Guide to the amsrefs Package”, where the “amsrefs” is set in monospaced font.
- 26.
http://onlinelibrary.wiley.com/journal/10.1002/(ISSN)2330-1643/issues records these former titles: Journal of the American Society for Information Science and Technology, Journal of the American Society for Information Science, and American Documentation.
- 27.
References
Adler, E., Hoon, M.A., Mueller, K.L., Chandrashekar, J., Ryba, N.J., Zuker, C.S.: A novel family of mammalian taste receptors. Cell 100, 693–702 (2000)
Alpher, R.A., Bethe, H., Gamow, G.: The origin of chemical elements. Phys. Rev. 73, 803–804 (1948)
Bollen, J., de Sompel, H.V., Hagberg, A., Chute, R.: A principal component analysis of 39 scientific impact measures. PLOS ONE 4, e6022 (2009)
Eom, Y.H., Frahm, K.M., Benczúr, A., Shepelyansky, D.L.: Time evolution of Wikipedia network ranking. Eur. Phys. J. B 86 (2013). Article ID 492
Erxleben, F., Günther, M., Krötzsch, M., Mendez, J., Vrandečić, D.: Introducing Wikidata to the Linked Data web. In: Mika, P., et al. (eds.) ISWC 2014. LNCS, vol. 8796, pp. 50–65. Springer, Cham (2014). https://doi.org/10.1007/978-3-319-11964-9_4
Gorgolewski, K.J., Varoquaux, G., Rivera, G., Schwarz, Y., Ghosh, S.S., Maumet, C., Sochat, V.V., Nichols, T.E., Poldrack, R., Poline, J.B., Yarkoni, T., Margulies, D.S.: NeuroVault.org: a web-based repository for collecting and sharing unthresholded statistical maps of the human brain. Front. Neuroinformatics 9, 8 (2015)
Grinberg, M.: Flask Web Development, April 2014
Heller, S.R., McNaught, A., Stein, S., Tchekhovskoi, D., Pletnev, I.: InChI - the worldwide chemical structure identifier standard. J. Cheminformatics 5, 7 (2013)
Hernández, D., Hogan, A., Krötzsch, M.: Reifying RDF: what works well with Wikidata? In: Proceedings of the 11th International Workshop on Scalable Semantic Web Knowledge Base Systems, September 2015. http://users.dcc.uchile.cl/~dhernand/research/ssws-2015-reifying.pdf
Hernández, D., Hogan, A., Riveros, C., Rojas, C., Zerega, E.: Querying Wikidata: comparing SPARQL, relational and graph databases. In: Groth, P., et al. (eds.) ISWC 2016. LNCS, vol. 9982, pp. 88–103. Springer, Cham (2016). https://doi.org/10.1007/978-3-319-46547-0_10
Hull, D., Pettifer, S., Kell, D.: Defrosting the digital library: bibliographic tools for the next generation web. PLOS Comput. Biol. 4, e1000204 (2008)
IFLA Study Group on the Functional Requirements for Bibliographic Records: Functional Requirements for Bibliographic Records, February 2009. http://www.ifla.org/files/assets/cataloguing/frbr/frbr_2008.pdf
Khabsa, M., Giles, C.L.: The number of scholarly documents on the public web. PLOS ONE 9, e93949 (2014)
Kikkawa, J., Takaku, M., Yoshikane, F.: DOI links on Wikipedia. In: Morishima, A., Rauber, A., Liew, C.L. (eds.) ICADL 2016. LNCS, vol. 10075, pp. 369–380. Springer, Cham (2016). https://doi.org/10.1007/978-3-319-49304-6_40
Kousha, K., Thelwall, M.: Are Wikipedia citations important evidence of the impact of scholarly articles and books? J. Am. Soc. Inf. Sci. 68, 762–779 (2016)
Krötzsch, M., Vrandečić, D., Völkel, M.: Semantic MediaWiki. In: Cruz, I., et al. (eds.) ISWC 2006. LNCS, vol. 4273, pp. 935–942. Springer, Heidelberg (2006). https://doi.org/10.1007/11926078_68
Lin, J., Fenner, M.: An analysis of Wikipedia references across PLOS publications, December 2014. https://figshare.com/articles/An_analysis_of_Wikipedia_references_across_PLOS_publications/1048991/files/1546358.pdf
Maggio, L.A., Willinsky, J., Steinberg, R., Mietchen, D., Wass, J., Dong, T.: Wikipedia as a gateway to biomedical research: the relative distribution and use of citations in the English Wikipedia. bioRxiv.org: the preprint server for biology, July 2017
Mishra, S., Torvik, V.I.: Quantifying conceptual novelty in the biomedical literature. DLib Mag. 22(9/10) (2016). https://doi.org/10.1045/september2016-mishra
Nielsen, F.Å.: Scientific citations in Wikipedia. First Monday 12 (2007). http://firstmonday.org/article/view/1997/1872
Nielsen, F.Å.: Clustering of scientific citations in Wikipedia, December 2008. http://www2.imm.dtu.dk/pubdb/views/edoc_download.php/5666/pdf/imm5666.pdf
Nielsen, F.Å.: Brede Wiki: a neuroinformatics web service with structured information. Front. Neur. Conference Abstract: Neuroinformatics (2009). https://doi.org/10.3389/conf.neuro.11.2009.08.072
Nielsen, F.Å., Mietchen, D., Willighagen, E.: Scholia, March 2017
Nielsen, F.Å., Mietchen, D., Willighagen, E.: Scholia and scientometrics with Wikidata, March 2017. https://arxiv.org/pdf/1703.04222.pdf
Okoli, C., Mehdi, M., Mesgari, M., Nielsen, F.Å., Lanamäki, A.: The people’s encyclopedia under the gaze of the sages: a systematic review of scholarly research on Wikipedia, March 2012. https://papers.ssrn.com/sol3/papers.cfm?abstract_id=2021326
Patashnik, O.: BIBTEX yesterday, today, and tomorrow. TUGboat 24, 25–30 (2003). https://www.tug.org/TUGboat/Articles/tb24-1/patashnik.pdf
Poldrack, R., Barch, D.M., Mitchell, J.P., Wager, T.D., Wagner, A.D., Devlin, J.T., Cumba, C., Koyejo, O., Milham, M.P.: Toward open sharing of task-based fMRI data: the OpenfMRI project. Front. Neuroinformatics 7, 12 (2013)
Pooladian, A., Borrego, Á.: Methodological issues in measuring citations in Wikipedia: a case study in Library and Information Science. Scientometrics 113, 455–464 (2017)
Priem, J., Piwowar, H.A., Hemminger, B.M.: Altmetrics in the wild: using social media to explore scholarly impact, March 2012. https://arxiv.org/html/1203.4745
Putman, T.E., Lelong, S., Burgstaller-Muehlbacher, S., Waagmeester, A., Diesh, C., Dunn, N., Munoz-Torres, M., Stupp, G., Wu, C., Su, A.I., Good, B.M.: WikiGenomes: an open web application for community consumption and curation of gene annotation data in Wikidata. In: Database 2017, March 2017. http://biorxiv.org/content/biorxiv/early/2017/01/21/102046.full.pdf
Putnam, H.: Is semantics possible? Metaphilosophy 1, 187–201 (1970)
Romero, A.R., Tzovaras, B.G., Greene, C.S., Himmelstein, D.S., McLaughlin, S.R.: Sci-Hub provides access to nearly all scholarly literature. PeerJ preprints, July 2017
Taraborelli, D., Dugan, J.M., Pintscher, L., Mietchen, D., Neylon, C.: WikiCite 2016 report, November 2016. https://upload.wikimedia.org/wikipedia/commons/2/2b/WikiCite_2016_report.pdf
Teplitskiy, M., Lu, G., Duede, E.: Amplifying the impact of open access: Wikipedia and the diffusion of science. J. Am. Soc. Inf. Sci. 68(9), 2116–2127 (2017)
Verborgh, R., Sande, M.V., Colpaert, P., Coppens, S., Mannens, E., de Walle, R.V.: Web-scale querying through linked data fragments. In: Proceedings of the Workshop on Linked Data on the Web, July 2014. http://ceur-ws.org/Vol-1184/ldow2014_paper_04.pdf
Vrandečić, D., Krötzsch, M.: Wikidata: a free collaborative knowledgebase. Commun. ACM 57, 78–85 (2014). http://cacm.acm.org/magazines/2014/10/178785-wikidata/fulltext
Watson, J.D., Crick, F.: Molecular structure of nucleic acids: a structure for deoxyribose nucleic acid. Nature 171, 737–738 (1953). http://www.nature.com/nature/dna50/watsoncrick.pdf
Yarkoni, T., Poldrack, R., Nichols, T.E., Essen, D.C.V., Wager, T.D.: Large-scale automated synthesis of human functional neuroimaging data. Nat. Methods 8, 665–670 (2011)
Zhirov, A.O., Zhirov, O.V., Shepelyansky, D.L.: Two-dimensional ranking of Wikipedia articles. Eur. Phys. J. B 77, 523–531 (2010)
Acknowledgements
This work was supported by Innovationsfonden through the DABAI project. The work on Scholia was spawned by the WikiCite project [33]. We would like to thank the organizers of the workshop, particularly Dario Taraborelli. Finn Årup Nielsen’s participation in the workshop was sponsored by an award from the Reinholdt W. Jorck og Hustrus Fund. We would also like to thank Magnus Manske, James Hare, Tom Arrow, Andra Waagmeester, and Sebastian Burgstaller-Muehlbacher for considerable work with Wikidata tools and data in the context of WikiCite. This paper was extended from another paper [24]. We thank Chiara Ghidini and the two other reviewers for providing suggestions for the improvement of that manuscript.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
<SimplePara><Emphasis Type="Bold">Open Access</Emphasis> This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.</SimplePara> <SimplePara>The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.</SimplePara>
Copyright information
© 2017 The Author(s)
About this paper
Cite this paper
Nielsen, F.Å., Mietchen, D., Willighagen, E. (2017). Scholia, Scientometrics and Wikidata. In: Blomqvist, E., Hose, K., Paulheim, H., Ławrynowicz, A., Ciravegna, F., Hartig, O. (eds) The Semantic Web: ESWC 2017 Satellite Events. ESWC 2017. Lecture Notes in Computer Science(), vol 10577. Springer, Cham. https://doi.org/10.1007/978-3-319-70407-4_36
Download citation
DOI: https://doi.org/10.1007/978-3-319-70407-4_36
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-70406-7
Online ISBN: 978-3-319-70407-4
eBook Packages: Computer ScienceComputer Science (R0)