Abstract
Image-based sequencing of RNA molecules directly in tissue samples provides a unique way of relating spatially varying gene expression to tissue morphology. Despite the fact that tissue samples are typically cut in micrometer thin sections, modern molecular detection methods result in signals so densely packed that optical “slicing” by imaging at multiple focal planes becomes necessary to image all signals. Chromatic aberration, signal crosstalk and low signal to noise ratio further complicates the analysis of multiple sequences in parallel. Here a previous 2D analysis approach for image-based gene decoding was used to show how signal count as well as signal precision is increased when analyzing the data in 3D instead. We corrected the extracted signal measurements for signal crosstalk, and improved the results of both 2D and 3D analysis. We applied our methodologies on a tissue sample imaged in six fluorescent channels during five cycles and seven focal planes, resulting in 210 images. Our methods are able to detect more than 5000 signals representing 140 different expressed genes analyzed and decoded in parallel.
You have full access to this open access chapter, Download conference paper PDF
Similar content being viewed by others
Keywords
- 2D and 3D signal detection
- Microscopy based in situ sequencing
- Image processing & analysis
- Crosstalk compensation
1 Introduction
Digital pathology is making its way into modern clinical diagnosis, increasing the need for automated digital image analysis methods for fast and reproducible quantification of tissue morphology [1]. In multi-cellular organisms, all cells have the same genes, while at the same time different cell types have different functions. The identity and function of a cell is defined by the gene expression (i.e., transcription). Thus, analysis of gene expression provides valuable information on health and disease, e.g. by identifying different types of immune cells or metastatic tumor cells. Most existing approaches for analysis of gene expression are based on bulk analysis of larger tissue samples, making it impossible to correlate gene expression with individual cells. More recently, analysis of individual cells has been made possible [2], but requires cells to be removed from the tissue architecture, resulting in loss of spatial information.
Amplified expressed genes (here enhanced for visualization purposes) in a tissue sample imaged in five sequencing cycles. In each cycle, four different fluorescent probes target each of the four letters of the genetic code. In this illustration, cyan=A, orange=C, magenta=G, green=T. The sequence of colors in a given position reveals the barcode of a unique expressed gene. (Color figure online)
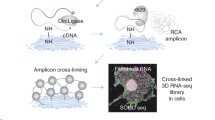
Our previously published methods for image-based in situ sequencing of expressed genes allow multiplexed gene expression profiling at cellular resolution in intact tissue samples, and thus opens up for detailed large-scale comparison of genotype and phenotype [3]; similar approaches have later been developed by others [4, 5]. Expressed genes are detected by molecular probes, locally amplified by rolling circle amplification, and decoded by sequential staining and imaging cycles. Each cycle targets the four letters of the genetic code with different fluorescent colors (see Fig. 1). By controlled design of probes, such that each probe contains a known “barcode” (i.e., sequence of nucleotide bases), it is known a priori what sequences of signals to expect across fluorescent colors and sequencing cycles, and only the number of signals as well as their location are unknown. Multiple molecular probes, targeting genes are typically used in parallel, and as little as five cycles of decoding can detect as many as 4\(^5\) = 1024 distinct barcodes in the same tissue sample. By comparing the number of expected barcodes to the number of unexpected barcodes (most likely originating from random noise and autofluorescence), it is possible to evaluate precision as well as efficiency (number of detected signals) of an image analysis approach.
Considering image size and richness of information, computerized image processing provides tools for enabling spatially resolved information and quantitative measurements. Tissue samples are typically cut in slices of a few micrometers prior to analysis, yet the data is typically collected by imaging the sample at multiple focal planes, acquiring a stack of images representing a 3D volume. An argument for this is that the different micrometer-sized signals often lie in different focal planes, making it impossible to collect an image where all are focused at once. Despite the data being 3D, all analysis approaches previously described were based on 2D projections of such 3D volumes. This is true also for our own previous approach [3, 6] implemented in TissueMaps [7], a platform for 2D giga-pixel image analysis and visualization built on free and open-source software.
As more genes are targeted in parallel, and the efficiency of the molecular methods increases, the signals in the tissue samples become denser. This means that a lot of information will be lost when relying on 2D projections for signal decoding. To avoid over-crowding, one has to limit the amplification step, meaning that a more complete analysis of gene expression comes at the cost of lower signal-to-noise ratios and signals close to the resolution limits of the microscope. Images are shifted between imaging cycles due to the manual staining/washing procedures, and signals from different fluorophores may be shifted due to chromatic aberration which further complicates the data analysis. Correction for chromatic aberrations has been suggested for similar methods by others: Briefly, the methods of [4, 5] first correct the effects of chromatic aberrations, respectively, through deconvolution and morphological opening followed by background subtraction. Then, the alignment is done on the maximum-intensity projection (MIP) along the z-dimension and using brick-based algorithm and cross-correlation of MIP along the c-dimension. Finally, the signal detection is completed by a per-pixel base calling and barcoding evaluation for maxima above a specified threshold value in a log-filtered version of the aligned images.
In this study, we approached the challenge of analyzing a full 3D data set with four color channels and five sequencing cycles. We compared the output to our previously published 2D approach [3, 6] applied to the same dataset projected to 2D. Finally, these results were improved using a post-processing crosstalk compensation to better separate the different color channels, and thus, correct some unexpected transcripts to expected barcodes. The methods were evaluated by comparing the number of detected signals by each method as well as the ratio of expected versus unexpected barcodes of targeted genes.
2 Image Acquisition
A total of 140 gene transcripts were targeted within a 10 \(\upmu \)m thick tissue sample, and subjected to five cycles of sequencing by ligation as previously described [3]. Images were acquired at seven different focal depths, 1.4 \(\upmu \)m apart, to create a 3D image volume using an Axioplan II epifluorescence microscope with a numerical aperture of 0.8 and a nominal magnification of 20.0 at 610 \(\upmu \text {m}\) distance. In each sequencing cycle, the four letters of the genetic code, A, C, G, and T, were fluorescence stained with Cy5, Texas Red, Cy3, and Fluorescein respectively. Furthermore, a general stain (AF750) marking all targets and a nuclear stain (DAPI) were also added to visualize signal distribution and tissue morphology, resulting in a total of six color channels. The resulting image volume is 2048 \(\times \) 2048 pixels, with a z-dimension of seven, a color dimension of six and a time dimension (=sequencing cycles or t) of five, for a total of 210 images to process. A cut out volume of 63 \(\times \) 66 \(\times \) 7 voxels from one color channel at one time point is shown in Fig. 2, illustrating signal size, noise and resolution in different spatial dimensions. Note that signals located in the same (x,y) position, but different z-positions will be merged when working with projected data in 2D.
3D visualizations of the image showing that more than one signal can appear along the z-direction (blue axis). The background images are the maximum-intensity projection of the slices in the general stain channel. The left image shows an example of the spatial distribution of the signals in the general stain. The right image shows the spatial distribution of the individual signal detection separated channel-wise (one per color). This illustrates the need for a detection in 3D since some signals are merged during the projection. (Color figure online)
3 Image Analysis
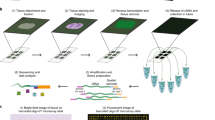
The global workflow aims to align and normalize the data prior to sequence decoding, as illustrated in Fig. 3. The challenges lie in (i) image registration, compensating for alignment shifts along the sequencing cycles and chromatic shift, (ii) signal detection and normalization, and (iii) signal decoding. The signal-to-noise ratio (SNR) in the images is limited by the trade off between exposure time during image acquisition and bleaching of the stains. The longer the exposure time, the higher the SNR, but at the same time there is an increased risk of bleaching signals in neighboring focal planes. In order to detect as many true signals as possible, we have decided to have a more inclusive approach for signal detection. Following signal decoding noise and true signals were discriminated using a quality measurement as described in Sect. 3.3.
Workflow of the original 2D method and the proposed 3D approach. The signal decoding relies on measuring the signal intensity at the same position for all fluorescent channels at each sequencing step. Therefore, registration is needed, in the 2D case it is a registration of the image data, while in 3D a registration of detected signals. Post-processing including crosstalk compensation increase signal confidence, as defined by the quality measurement.
3.1 2D Approach
For the 2D approach, we used our previously published method [3, 6], implemented in the TissueMaps workflow [7]. TissueMaps is built on free and open source tools, and the analysis workflow makes use of the CellProfiler software [8]. The 2D analysis started with the MIP of the image stack (reducing the z-dimension to one). Following the MIP, for each cycle (t), each image channel (\(I_{tc}\), c representing either the general stain or one of the four letters of the genetic code) was first enhanced by a top-hat transformation (\(F_{tc}\)) with a structuring element (B) consisting of a disc with radius 10 pixels:
where \(\circ \) is a morphological opening. Individual signals were then defined by a labeled mask (L) in the general stain channel (D) of the first sequencing cycle (\(t=1\)), by a fixed intensity threshold, low enough to detect the signals after the top-hat (here, equal to 0.5). Finally, clustered signals were separated by shape-based watershed segmentation, i.e., a watershed applied on the negated distance transform, since signals are relatively circular [9]. Filtered images (\(F_{tc}\)) from the same sequencing cycle were thereafter registered (\(R_{tc}= registered(F_{tc}\))) towards the general stain using a rigid-body transformation (preserving the distance between every pair of points), from the “MultiStackReg” plugin for Fiji [10]. We applied the final mask representing the signals (L) on \(R_{tc}\), so that \(L_s\) is the set of pixels representing signal s in L. Finally, the intensity (\(S_{stc}\)) for each signal (s) in each channel (c) and time step (t) is defined in the 2D method as the maximum fluorescence intensity:
We specifically extracted its (x, y) location as well as the intensity of this location in each of the five color channels (general stain and four letters of the genetic code), and five time steps in order to later decode and evaluate the signal as described in Sect. 3.3.
3.2 3D Approach
In the 3D approach, signals were separately detected in all color channels at all time steps using a local thresholding approach referred to as Per Object Ellipsefit (POE) [11]. The POE method computes local adaptive thresholds for each individual object (signal) where the threshold values are set to optimize the ellipse (ellipsoid in 3D) fit. This is done by creating a component tree [12] and traversing the pixels in order of decreasing intensities. Ellipsoid fit is defined by computing the moment matrix M for each object, extracting the axes from the eigenvalues of M, and computing the ratio between the actual object volume and an ideal ellipsoid with the dimensions given by these axes. The search for the best ellipsoid fit is done within given ranges for object volume (36–96 voxels), major and minor axis length (3–8 pixels), and value of the ellipsoid fit (\(\ge \)0.5).
Following signal detection, 3D spatial coordinates of detected signals were aligned and grouped. Within each time step (sequencing cycle) the color channels representing A, C, G, and T were affinely registered to the general stain of that same time step, using Iterative Closest Point (ICP) [13], followed by a spline based ICP version [14, 15], with a grid of 6 \(\times \) 6 \(\times \) 5 control points, that further corrects any chromatic aberration. Once the channels were aligned within each cycle, the general stain of each cycle was aligned with the general stain of the first cycle, used as a reference, utilizing rigid ICP registration. The associated channels of the time step were aligned using the same transformation.
Due to digitization effects and noise, slight shifts in the detected signals, for different color channels and time steps, remain also after the registration. Detected signals closer to each other than 3.4 pixels were merged together as one spot. As (x, y) location of a merged signal s we use the centroid of the corresponding cluster of (registered) signals. The intensity values of all merged signals were extracted from the smoothed (gaussian filter with \(\sigma =0.5\)) and dilated (ball-shaped structural element with five pixels diameter) original images, utilizing the inverse of the respective registration transformations.
Intensity measures were normalized separately for each channel (c) and time (t), such that signals with a brightness equal to the mode of the respective image volume gets the value zero, and the mean detected signal intensity is mapped to the value one:
where \(S_{stc}\) is the intensity of signal s in channel c and time t for the 3D method, and \(N_s\) is the total number of detected signals.
Due to the inclusive intensity threshold used for the signal detection, artifacts from random background noise may have been detected as well. After normalization, a quality check based on the general stain was applied to reduce such noise. We require that the general stain channel (D), for each cycle (c), presents each signal detected (s), so that the following condition holds for all cycles:
This step reduces the number of signals by approximately 2%.
3.3 Sequence Decoding and Quality Measurement
We measured the respective quality value for the 2D and 3D methods to evaluate the consistency of the signals detected. For each sequencing cycle, every location containing a signal is assigned the base, A, C, G or T, decided on the highest image intensity (following top-hat (2D) or normalization according to Eq. (3) (3D)). Autofluorescence may result in false signals that have a high intensity across all sequencing cycles, but always display the same color (that is, always appear in the same color channel). Such signals will appear as “homopolymers”, e.g. barcodes consisting of a single letter, such as ‘AAAAA’ or ‘GGGGG’. No such signals were included in the expected barcodes, and they are removed from our set of detected signals, reducing the number of signals by 0.6%.
To evaluate the signals detected, a quality \(Q_{st}\) of a signal s, in the cycle t was defined as:
The quality score of the full sequence \(Q_{s}\) of signal s is further defined by the quality of its “weakest” cycle:
The quality score ranges from \(\frac{1}{N_t}\) (i.e., all signals equal) to 1 (all non-max signals equal to 0).
3.4 Crosstalk Compensation
Intensity values detected from each of the five sequencing cycles were crosstalk compensated in order to color-correct the intensities and determine the real dye concentration present in each signal. The sequencing cycles can not be assumed to be independent from each other, but the sequencing process and the image acquisition is affected by several kinds of cycle-dependent noise (e.g., focus, imperfect image registration, chromatic aberration, photobleaching, and other experimental conditions), meaning that the crosstalk between channels may vary cycle to cycle. Therefore a separate crosstalk compensation matrix for each of the sequencing cycles was estimated. Each crosstalk matrix \(X_t\) was estimated as in Sect. 2.2.6 of Li and Speed [16], inverted and multiplied by the matrix of the intensities of all signals s of cycle t, producing crosstalk compensated intensity values:
We measured a new quality value for the methods by replacing the intensity \(S_{stc}\) in Eq. 5 by the compensated intensity value \(X_{stc}\).
3.5 Validation Approach
The only “ground truth” available for this type of image data is the a priori knowledge of the barcodes of the probes applied to the tissue section. In this particular experiment, 140 different probes were applied. The barcode length is five letters, meaning that our decoding approach may find 4\(^5\) = 1024 different codes, but only 140 out of these codes are to be expected (TP), and it can be assumed that any other code found is noise due to poor signal detection/decoding and is considered as unexpected (FP). There are of course also other sources of error, such as actual errors in the probes, but these will affect the 2D and 3D approach equally. Using the quality measure described in Sect. 3.3 an acceptance threshold can be set to balance the signal count vs. the signal precision (TP/(TP+FP)).
4 Results
4.1 Validation
Out of the 1024 possible barcodes, only 140 correspond to the barcodes of our targeted gene transcripts. If decoded signals were completely random (and the four homopolymers removed), a precision of 140/(1024−4) = 0.14 would be expected. From Fig. 4, showing number of detected TP versus number of detected FP depending on the quality threshold value, we can see that the new 3D approach detects more signals than the 2D method with a higher ratio of TP over FP (respectively red curve and blue curve). The alignment method in the 2D workflow produces part of the FP signals due to its difficulty to find control points to define the transformation, especially in this noisy dataset. Moreover, the MIP tends to overcrowd the working plane so that two signals may overlap and corrupt the decoding process. The crosstalk compensation improves the 2D workflow by correcting some of the unexpected barcodes and thus, improving these results (blue dashed curve). On the other hand, the 3D approach is able to extract more robust information through the z-dimension which helps for both the registration process and for the spatial localization of the signal as they are better separated. These better results are also improved through the crosstalk compensation (red dashed curve). Consequently, assuming an acceptable ratio of one FP for four TP, i.e. a precision of 0.8, then we obtained respectively 2641 and 2968 TP for the 2D and 3D method, which increase to 3622 and 4742 TP with the crosstalk compensation (black square markers).
Comparison of the original 2D method and the proposed 3D approach by plotting true positive signals (TP) against the false positive signals (FP) at various quality threshold settings. The red and blue curves show the signals detected by the 3D and 2D approaches respectively, before compensation for crosstalk. The dashed curves show the results after the application of crosstalk compensation. Precision, i.e., TP/(TP+FP), increases for both the 2D and the 3D approach when crosstalk compensation is applied as shown by the black square markers for a precision of 0.8. (Color figure online)
4.2 Visualisation
We confirmed the spatial localization of the transcripts detected by using the TissueMaps platform. Currently, this platform allows the display of 54 different symbols to localize the genes on a 2D image at different resolutions. We chose the projected general stain image as background and displayed the 54 most common barcodes (sum of the two methods) among the total 140 genes detected by our methods (Fig. 5).
Visualization of the 54 most common transcripts among the total 140 expected barcodes using the TissueMaps platform. The top image is the result of the 2D approach while the middle visualization corresponds to the 3D method. The bottom bar plot represents these 54 transcript counts for the 2D (in blue) and 3D (in red) methods. (Color figure online)
5 Discussions
Digital microscopes can capture 3D images of signals emitted by molecular detection probes by recording data at multiple focal planes of the imaged tissue samples. While the current 2D method of gene decoding, by applying a MIP, provides an overview of the stack with a good SNR, it tends to overcrowd the signals and lose some of the individuality. The 3D approach, presented in this study, analyzes the different slices of the tissue volume to detect more of separated signals. The individual transcripts are in the same proportion (in respect to the total number detected) and present the same global pattern in the tissue (Fig. 5).
The advantages of the 3D method also come from the improvement and use of new steps. The images were normalized based on their mode and mean, and the segmentation was applied to each 3D volume (four channels \(\times \) five sequencing cycles) individually rather than on the general stain. This allows the 3D method to compensate the SNR channel-wise, similarly to the top-hat in the 2D approach, but also to have a better definition of the individuality in each channel where signal overlap could occur in the general stain.
We also improved the general quality measurement and gene decoding by incorporating crosstalk compensation. This allows us to correct some of the unexpected barcodes based on the signal intensities (in each channel) and the general tendency of the signals to switch from one base to another. For both methods, the crosstalk compensation as a post-process converts around a thousand of false positive signals into true positive signals (Fig. 4) and increases the precision of our results.
References
Gurcan, M.N., Boucheron, L.E., Can, A., Madabhushi, A., Rajpoot, N.M., Yener, B.: Histopathological image analysis: a review. IEEE Rev. Biomed. Eng. 2, 147–171 (2009)
Darmanis, S., Sloan, S.A., Zhang, Y., Enge, M., Caneda, C., Shuer, L.M., Gephart, M.G.H., Barres, B.A., Quake, S.R.: A survey of human brain transcriptome diversity at the single cell level. Proc. Nat. Acad. Sci. 112(23), 7285–7290 (2015)
Ke, R., Mignardi, M., Pacureanu, A., Svedlund, J., Botling, J., Wählby, C., Nilsson, M.: In situ sequencing for RNA analysis in preserved tissue and cells. Nat. Methods 10(9), 857–860 (2013)
Lee, J.H., Daugharthy, E.R., Scheiman, J., Kalhor, R., Yang, J.L., Ferrante, T.C., Terry, R., Jeanty, S.S., Li, C., Amamoto, R., Peters, D.T., Turczyk, B.M., Marblestone, A.H., Inverso, S.A., Bernard, A., Mali, P., Rios, X., Aach, J., Church, G.M.: Highly multiplexed subcellular RNA sequencing in situ. Science 343(6177), 1360–1363 (2014)
Shah, S., Lubeck, E., Zhou, W., Cai, L.: In situ transcription profiling of single cells reveals spatial organization of cells in the mouse hippocampus. Neuron 92(2), 342–357 (2016)
Pacureanu, A., Ke, R., Mignardi, M., Nilsson, M., Wählby, C.: Image based in situ sequencing for RNA analysis in tissue. In: 2014 IEEE 11th International Symposium on Biomedical Imaging (ISBI), pp. 286–289. IEEE (2014)
Ranefall, P., Pacureanu, A., Avenel, C., Carpenter, A.E., Wählby, C.: The giga-pixel challenge: full resolution image analysis without losing the big picture: an open-source approach for multi-scale analysis and visualization of slide-scanner data. In: SSBA 2014, Symposium of the Swedish Society for Automated Image Analysis, Luleå, Sweden (2014)
Carpenter, A.E., Jones, T.R., Lamprecht, M.R., Clarke, C., Kang, I.H., Friman, O., Guertin, D.A., Chang, J.H., Lindquist, R.A., Moffat, J., Golland, P., Sabatini, M.: Cell profiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 7(10), R100 (2006)
Malpica, N., Ortiz de Solorzano, C., Vaquero, J.J., Santos, A., Vallcorba, I., Garcia-Sagredo, J.M., Pozo, F.D.: Applying watershed algorithms to the segmentation of clustered nuclei. Cytometry 28, 289–297 (1997)
Thevenaz, P., Ruttimann, U.E., Unser, M.: A pyramid approach to subpixel registration based on intensity. IEEE Trans. Image Process. 7(1), 27–41 (1998)
Ranefall, P., Sadanandan, S.K., Wählby, C.: Fast adaptive local thresholding based on ellipse fit. In: International Symposium on Biomedical Imaging (ISBI 2016), Prague, Czech Republic (2016)
Najman, L., Couprie, M.: Building the component tree in quasi-linear time. IEEE Trans. Image Process. 15(11), 3531–3539 (2006)
Amberg, B., Romdhani, S., Vetter, T.: Optimal step nonrigid ICP algorithms for surface registration. In: 2007 IEEE Conference on Computer Vision and Pattern Recognition, pp. 1–8. IEEE (2007)
Rueckert, D., Sonoda, L.I., Hayes, C., Hill, D.L., Leach, M.O., Hawkes, D.J.: Nonrigid registration using free-form deformations: application to breast MR images. IEEE Trans. Med. Imaging 18(8), 712–721 (1999)
Lee, S., Wolberg, G., Shin, S.Y.: Scattered data interpolation with multilevel B-splines. IEEE Trans. Vis. Comput. Graph. 3(3), 228–244 (1997)
Li, L., Speed, T.P.: An estimate of the crosstalk matrix in four-dye fluorescence-based DNA sequencing. Electrophoresis 20(7), 1433–1442 (1999)
Acknowledgments
The authors would like to thank Johan Oelrich for contributions to the development of TissueMaps, and the European Research council for funding via ERC Consolidator grant 682810 to C. Wählby.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this paper
Cite this paper
Bombrun, M. et al. (2017). Decoding Gene Expression in 2D and 3D. In: Sharma, P., Bianchi, F. (eds) Image Analysis. SCIA 2017. Lecture Notes in Computer Science(), vol 10270. Springer, Cham. https://doi.org/10.1007/978-3-319-59129-2_22
Download citation
DOI: https://doi.org/10.1007/978-3-319-59129-2_22
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-59128-5
Online ISBN: 978-3-319-59129-2
eBook Packages: Computer ScienceComputer Science (R0)