Abstract
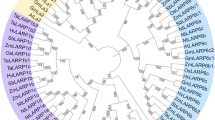
Glutathione (GSH) protects proteins against oxidation of their thiol-containing groups, by alternatively becoming the subject of oxidation, forming glutathione disulfide (GSSG). Appropriate GSH:GSSG levels are maintained by glutathione reductase (GR), a homodimeric flavoprotein which uses NADPH to reduce one GSSG molecule to two of GSH. This enzyme has been characterized in several species, and described as highly conserved, with two isoforms only. Heterogeneity and discticntiveness of plant reproductive tissues led us to investigate the presence of GR sequences. A de novo assembled and annotated olive reproductive transcriptome was subjected to screening, which allowed us to identify at least 11 GR homologues (1 pollen-specific and 10 from pistils). Primers were designed, and full-length sequences were obtained through PCR. In silico analysis, including phylogeny, 3-D modeling of N-terminus, and prediction of cellular localization and post-translational modifications was carried out to shed light into the involvement of olive pollen-intrinsic GR in reproductive development.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Shah, Z.H., Hamooh, B.T., Daur, I., Ha Rehman, H.M., Alghabari, F.: Transcriptomics and biochemical profiling: current dynamics in elucidating the potential attributes of olive. Curr. Issues Mol. Biol. 21, 73–98 (2017)
Kaur, H., Bhatla, S.C.: Melatonin and nitric oxide modulate glutathione content and glutathione reductase activity in sunflower seedling cotyledons accompanying salt stress. Nitric Oxide 59, 42–53 (2016)
Halliwell, B., Foyer, C.H.: Properties and physiological function of a glutathione reductase purified from spinach leaves by affinity chromatography. Planta 139(1), 9–17 (1978)
Edwards, E.A., Rawsthorne, S., Mullineaux, P.M.: Subcellular distribution of multiple forms of glutathione reductase in leaves of pea (Pisum sativum L.). Planta 180, 278–284 (1990)
Foyer, C.H., Halliwell, B.: The presence of glutathione and glutathione reductase in chloroplasts: a proposed role in ascorbic acid metabolism. Planta 133(1), 21–25 (1976)
Rasmusson, A.G., Møller, I.M.: NADP-utilizing enzymes in the matrix of plant mitochondria. Plant Physiol. 94, 1012–1018 (1990)
Jiménez, A., Hernández, J.A., del Río, L.A., Sevilla, F.: Evidence for the presence of the ascorbate-glutathione cycle in mitochondria and peroxisomes of pea leaves. Plant Physiol. 114, 175–284 (1997)
Stevens, R.G., Creissen, G.P., Mullineaux, P.M.: Characterization of pea cytosolic glutathione reductase expressed in transgenic tobacco. Planta 211, 537–545 (2000)
Romero-Puertas, M.C., Corpas, F.J., Sandalio, L.M., Leterrier, M., Rodríguez-Serrano, M., del Río, L.A., Palma, J.M.: Glutathione reductase from pea leaves: response to abiotic stress and characterization of the peroxisomal isozyme. New Phytol. 170, 43–52 (2006)
Creissen, G., Edwards, E.A., Enard, C., Wellburn, A., Mullineaux, P.: Molecular characterization of glutathione reductase cDNAs from pea (Pisum sativum L.). Plant J. 2(1), 129–131 (1992)
Chew, O., Whelan, J., Miller, A.H.: Molecular definition of the ascorbate-glutathione cycle in Arabidopsis mitochondria reveals dual targeting of antioxidant defences in plants. J. Biol. Chem. 278(47), 46869–46877 (2003)
Kaur, N., Reumann, S., Hu, J.: Peroxisome biogenesis and function. The Arabidopsis Book. American Society of Plant Biologists, Rockville, MD (2009). doi:10.1199/tab.0123
Marty, L., Siala, W., Schwarzländer, M., Fricker, M.D., Wirtz, M., Sweetlove, L.J., Meyer, Y., Meyer, A.J., Reichheld, J.P., Hell, R.: The NADPH-dependent thioredoxin system constitutes a functional backup for cytosolic glutathione reductase in Arabidopsis. Proc. Natl. Acad. Sci. U.S.A. 106, 9109–9114 (2009)
Foyer, C.H., Noctor, G.: Redox sensing and signalling associated with reactive oxygen in chloroplasts, peroxisomes and mitochondria. Physiol. Plant. 119, 355–364 (2003)
del Río, L.A., Sandalio, L.M., Corpas, F.J., Palma, J.M., Barroso, J.B.: Reactive oxygen species and reactive nitrogen species in peroxisomes Production scavenging and role in cell signaling. Plant Physiol. 141, 330–335 (2006)
Nyathi, Y., Baker, A.: Plant peroxisomes as a source of signalling molecules. Biochim. Biophys. Acta 1763, 1478–1495 (2006)
Mhamdi, A., Queval, G., Chaouch, S., Vanderauwera, S., Van Breusegem, F., Noctor, G.: Catalase function in plants: a focus on Arabidopsis mutants as stress-mimic models. J. Exp. Bot. 61, 4197–4220 (2010)
Garrett, R.H., Grisham, C.M.: Biochemistry, 3rd edn. Thomson Brooks/Cole, Belmont (2005). ISBN 0534490336
Anjum, N.A., Umar, S., Chan, M.T. (eds.): Ascorbate-Glutathione Pathway and Stress Tolerance in Plants. Springer, Netherlands (2010). ISBN 978-90-481-9404-9
Cejudo, F.J., Meyer, A.J., Reichheld, J.P., Rouhier, N., Traverso, J.A.: Thiol-Based Redox Homeostasis and Signalling. Frontiers Media SA, Lausanne (2014). ISBN 978-2-88919-284-7
Carmona, R., Zafra, A., Seoane, P., Castro, A.J., Guerrero-Fernández, D., Castillo-Castillo, T., Medina-García, A., Cánovas, F.M., Aldana-Montes, J.F., Navas-Delgado, I., Alché, J.D., Claros, M.G.: ReprOlive: a database with linked data for the olive tree (Olea europaea L.) reproductive transcriptome. Frontiers Plant Sci. 6, 625 (2015)
Cruz, F., Julca, I., Gómez-Garrido, J., Loska, D., Marcet-Houben, M., Cano, E., Galán, B., Frias, L., Ribeca, P., Derdak, S., Gut, M., Sánchez-Fernández, M., García, J.L., Gut, I.G., Vargas, P., Alioto, T.S., Gabaldón, T.: Genome sequence of the olive tree, Olea europaea. GigaScience 5, 29 (2016)
Olive genome and annotation files. http://denovo.cnag.cat/genomes/olive/
de Dios Alché, J., M’rani-Alaoui, M., Castro, A.J., Rodríguez-García, M.I.: Ole e 1, the major allergen from olive (Olea europaea L.) pollen, increases its expression and is released to the culture medium during in vitro germination. Plant Cell Physiol. 45, 1149–1157 (2004)
McWilliam, H., Li, W., Uludag, M., Squizzato, S., Park, Y.M., Buso, N., Cowley, A.P., Lopez, R.: Analysis tool web services from the EMBL-EBI. Nucleic Acids Res. 41, W597–W600 (2013)
Gouy, M., Guindon, S., Gascuel, O.: SeaView version 4: a multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol. Biol. Evol. 27, 221–224 (2010)
Darriba, D., Taboada, G.L., Doallo, R., Posada, D.: jModelTest 2: more models, new heuristics and parallel computing. Nat. Meth. 9, 772 (2012)
Chou, K.C., Shen, H.B.: Plant-mPLoc: a top-down strategy to augment the power for predicting plant protein subcellular localization. PLoS ONE 5, e11335 (2010)
Heazlewood, J.L., Durek, P., Hummel, J., Selbig, J., Weckwerth, W., Walther, D., Schulze, W.X.: PhosPhAt: a database of phosphorylation sites in Arabidopsis thaliana and a plant-specific phosphorylation site predictor. Nucleic Acids Res. 36, D1015–D1021 (2008)
Lee, T.-Y., Bretana, N., Lu, C.-T.: PlantPhos: using maximal dependence decomposition to identify plant phosphorylation sites with substrate site specificity. BMC Bioinf. 12, 261 (2011)
Xue, Y., Liu, Z., Gao, X., Jin, C., Wen, L., Yao, X., Ren, J.: GPS-SNO: computational prediction of protein S-nitrosylation sites with a modified GPS algorithm. PLoS ONE 5, e11290 (2010)
Martinez, A., Traverso, J.A., Valot, B., Ferro, M., Espagne, C., Ephritikhine, G., Zivy, M., Giglione, C., Meinnel, T.: Extent of N-terminal modifications in cytosolic proteins from eukaryotes. Proteomics 8, 2809–2831 (2008)
Kelley, L.A., Sternberg, M.J.: Protein structure prediction on the web: a case study using the Phyre server. Nat. Protoc. 4, 363–371 (2009)
Rutley, N., Twell, D.: A decade of pollen transcriptomics. Plant Reprod. 28, 73–89 (2015)
Dukowic-Schulze, S., Chen, C.: The meiotic transcriptome architecture of plants. Frontiers Plant Sci. 5, 220 (2014)
Zechmann, B., Mauch, F., Sticher, L., Müller, M.: Subcellular immunocytochemical analysis detects the highest concentrations of glutathione in mitochondria and not in plastids. J. Exp. Bot. 59, 4017–4027 (2008)
Zechmann, B., Koffler, B.E., Russell, S.D.: Glutathione synthesis is essential for pollen germination in vitro. BMC Plant Biol. 11, 54 (2011)
Zafra, A., Rodriguez-Garcia, M.I., Alche, JdD: Cellular localization of ROS and NO in olive reproductive tissues during flower development. BMC Plant Biol. 10, 36 (2010)
Acknowledgments
This work was supported by ERDF-cofunded projects BFU2016-77243-P, P2011-CVI7487, 201540E065, RTC-2015-4181-2 and RTC2016-4824-2. EGQ thanks the MINECO for FPI grant funding.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this paper
Cite this paper
García-Quirós, E., Carmona, R., Zafra, A., Gonzalo Claros, M., Alché, J.d.D. (2017). Identification and in silico Analysis of Glutathione Reductase Transcripts Expressed in Olive (Olea europaea L.) Pollen and Pistil. In: Rojas, I., Ortuño, F. (eds) Bioinformatics and Biomedical Engineering. IWBBIO 2017. Lecture Notes in Computer Science(), vol 10209. Springer, Cham. https://doi.org/10.1007/978-3-319-56154-7_18
Download citation
DOI: https://doi.org/10.1007/978-3-319-56154-7_18
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-56153-0
Online ISBN: 978-3-319-56154-7
eBook Packages: Computer ScienceComputer Science (R0)