Abstract
Accurate segmentation of brain white matter hyperintensities (WMHs) is important for prognosis and disease monitoring. To this end, classifiers are often trained – usually, using T1 and FLAIR weighted MR images. Incorporating additional features, derived from diffusion weighted MRI, could improve classification. However, the multitude of diffusion-derived features requires selecting the most adequate. For this, automated feature selection is commonly employed, which can often be sub-optimal. In this work, we propose a different approach, introducing a semi-automated pipeline to select interactively features for WMH classification. The advantage of this solution is the integration of the knowledge and skills of experts in the process. In our pipeline, a Visual Analytics (VA) system is employed, to enable user-driven feature selection. The resulting features are T1, FLAIR, Mean Diffusivity (MD), and Radial Diffusivity (RD) – and secondarily, \(C_S\) and Fractional Anisotropy (FA). The next step in the pipeline is to train a classifier with these features, and compare its results to a similar classifier, used in previous work with automated feature selection. Finally, VA is employed again, to analyze and understand the classifier performance and results.
You have full access to this open access chapter, Download conference paper PDF
Similar content being viewed by others
Keywords
- White matter hyperintensities (WMHs)
- Visual Analytics (VA)
- Classification
- Interactive feature selection
1 Introduction
White matter hyperintensities of presumed vascular origin (WMHs) are a common finding in MR images of elderly subjects. They are a manifestation of cerebral small vessel disease (SVD) and are associated with cognitive decline and dementia [1]. Accurate segmentation of WMHs is important for prognosis and disease monitoring. To this end, automated WMH classification techniques have been developed [2]. Conventional approaches include raw image intensities from T1 and FLAIR weighted MR images, but recently, it has been suggested that diffusion MRI can improve the segmentation [3, 4]. Multiple features can be derived from this imaging modality; thus, careful feature selection is required.
In this work, we propose a semi-automated approach, to aid the design of WMH classifiers. Our novelty is the introduction of a user-driven, interactive pipeline that provides new insight into the entire classification procedure – especially, in the identification of an adequate feature list, and the analysis of the outcome. Up to now, the knowledge and cognitive skills of experts have not been intensively involved in the process. In the first step of our pipeline, we employ a Visual Analytics (VA) system [5], where expert users select interactively the most important features. In the second step, the resulting feature list is used to train a classifier for WMH segmentation. The performance and results of this classifier can be analyzed and interpreted in the final step of the pipeline, using again VA.
2 Related Work
Visual Analytics (VA) refers to the field that combines, through interaction, visualizations with pattern recognition, data mining and statistics, and focuses on aiding exploration and analytical reasoning [6]. Recently, Raidou et al. proposed a highly interactive VA system for the exploration of intra-tumor tissue characteristics [5]. The system employs a t-Distributed Stochastic Neighbor Embedding [7] of several imaging-derived features, used in tumor diagnosis. It also consists of multiple interactive views, for the exploration and analysis of the underlying structure of the feature space, providing linking to anatomy and ground truth data. Yet, to the best of our knowledge, involving users through VA and interaction in an entire pipeline for feature selection, classification and outcome evaluation for WMH structures has not been addressed before.
3 Materials and Method
3.1 Subjects and MRI Data
We used the subjects of the MRBrainS13 challenge [8], with additional manual WMH delineations. Subjects included patients with diabetes and matched controls (men: 10, age: 71 ± 4 years). All subjects underwent a standardized 3 T MR exam, including a 3D T1-weighted, a multi-slice FLAIR, a multi-slice IR, and a single-shot EPI DTI sequence with 45 directions. All sequences were aligned with the FLAIR sequence [9]. The diffusion images were corrected for subject motion, eddy current induced geometric distortions, and EPI distortions, including the required B-matrix adjustments, using ExploreDTI [10].
The dataset includes T1, FLAIR and IR weighted images, as well as the following diffusion features: Fractional Anisotropy (FA), Mean Diffusivity (MD), Axial Diffusivity (AD), Radial Diffusivity (RD), the Westin measures \(C_L\), \(C_P\), \(C_S\) [11], and MNI152-normalized spatial coordinates [9, 12]. This exact dataset has been previously reported in a study of Kuijf et al. [3], for the investigation of the added value of diffusion features in a WMH classifier. Since we could have access to the exact same data and we share the same goal, we will use the previous work of Kuijf et al., as a baseline for the evaluation of our results.
3.2 Method
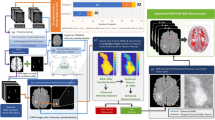
In this section, we describe our new pipelineFootnote 1 for the user-driven, interactive selection of features that can differentiate WMHs from healthy brain tissue. Our pipeline consists of three steps, depicted in Fig. 1. First, the data are interactively explored and analyzed by expert users, in the VA system proposed by Raidou et al. [5]. From this step, we obtain through interaction and visual analysis, a list of features, adequate for WMH detection. These features are subsequently used to train a classifier. After classification, the VA system is used again to evaluate and better understand the classification process and outcome.
Feature Selection Using VA. The VA system of Raidou et al. [5] is employed to interactively explore the data of each one of the available subjects (Fig. 2). Initially, t-Distributed Stochastic Neighbor Embedding (t-SNE) [7] is used to map the high-dimensional feature space of each subject (described in Sect. 3.1) into a reduced 2D abstract embedding view, preserving the local structure of the feature space. Spatial coordinates are excluded, as we are interested in preserving similarities in the feature space, and the voxel positions could introduce bias. In the resulting embedding view (Fig. 2-ii), close-by 2D data points reflect voxels with similar behavior in the high-dimensional feature space. Therefore, voxels from structures with similar imaging characteristics are expected to be grouped together in the embedding, in so-called visual clusters. Having available ground truth data, i.e., manual delineations of the WMHs, allows to associate visual clusters from the feature space to anatomy, and vice versa (Fig. 2-i).
When a WMH-containing visual cluster is interactively selected, its intrinsic feature characteristics are explored; for example, against other structures of the brain, or against WMHs voxels that are not within the selected visual cluster. Then, several linked views (Fig. 2-iii) are interactively updated with complementary data information. This includes feature distributions and correlations, multidimensional data patterns, cluster validity analysis and information on features that help separating visual clusters from each other, as given by the weights of the separation vector of Linear Discriminant Analysis (LDA). In this way, features suitable for the detection of WMHs are interactively identified.
For example, for the subject of Fig. 2, two visual clusters have been selected in the t-SNE of the middle view. As depicted in the anatomical views, one corresponds to the WMH core (green) and the other to the periphery (purple). Together, they represent the biggest part of the structure. Still, several small parts are missed. The separation vector, resulting from LDA between the two visual clusters containing the WMHs against the rest of the brain, is extracted. From the weights of this vector, features adequate for differentiating the detected WMHs from the rest of the brain, are identified. This analysis is subject-specific and has to be performed on a single-subject basis. When all subjects have been explored, the user decides on the most suitable feature list, overall.
Classification. In this step, many different classification approaches could be followed, but comparing to all would be out of scope, for this work. Recently, Kuijf et al. [3] presented an approach for WMH classification, using the same set of diffusion features. To evaluate whether our user-driven feature selection outperforms automated feature selection, we adopt a similar classification approach, as in the previous work of Kuijf et al. The list of features resulting from the VA system is used to train a k-nearest-neighbor classifier for WMH segmentation. For different feature combinations, several classifiers are trained with k = 50, 75, or 100, and the neighbor-weighted is either uniform or distance-based [3].
Evaluation of Classification. In many cases, classifiers are treated as black boxes, and users do not have actual insight into the achieved result. With this step, we want to provide a way for evaluating and understanding both the results of the classifier and the classifier itself. To this end, we import the binary masks resulting from the classification (detected vs. missed WMHs) into the VA system [5]. The user can interactively explore the high-dimensional feature space of the two regions of the mask, and generate hypotheses about why the classifier failed to detect parts of the WMHs, with respect to the imaging features.
4 Results
Feature Selection Using VA. In most of t-SNE embeddings of the subjects, the majority of voxels of the WMHs are grouped together, in one or two visual clusters, similar to the case depicted in Fig. 2. From selecting these visual clusters, we could identify that, for subjects with two visual clusters, these either correspond to the core and the periphery, or to anterior and posterior WMHs. For large WMHs (top 50 %), the visual clusters of the embedding identify 84–98% of the structures. For the rest, the visual clusters can at least detect the core, with a minimum detection percentage of 54 %.
The multiple interactive linked views of the VA system show that there are comparable behaviors, within all cases of visual clusters, especially for larger WMH structures. As mentioned before, the cluster analysis view of the VA system provides the separation vector, resulting from LDA between the visual cluster containing most of WMHs and the visual cluster of the rest of the brain. Table 1 depicts, for all investigated subjects, the weights of separation for these two visual clusters. In all – but three – cases, T1, FLAIR, RD and MD are more important, as they have a considerable weight. For bigger WMHs, \(C_S\) and FA also become important. The contribution of other features such as AD, \(C_L\), \(C_P\) and IR seems not significant.
Considering also the (cor-)relations between diffusion features, we decide on the overall set of features for the classifier: MD, RD, T1 and FLAIR (secondarily, \(C_S\) and FA). Here, we add the MNI152-normalized spatial coordinates (x, y, z) to better represent the brain volume and to suppress non-WMH structures.
Classification. Based on the results of the VA system, the following four combinations of feature sets \(s_i \in S\) are chosen for our k-NN classifier: \(s_1\): MD, RD, T1, FLAIR; \(s_2\): \(s_1 + \) (x, y, z); \(s_3\): \(s_1 + \) \(C_S\), FA; \(s_4\): \(s_3 {+} \) (x, y, z). For each classifier trained on a feature set \(s_i \in S\), we measure the sensitivity and Dice similarity coefficient (mean ± standard deviation), as shown in Table 2. These measurements are performed, with respect to the available manual delineations of the WMH structures. Furthermore, our results are compared to the feature sets \(f_i \in F\), previously used by Kuijf et al. [3]: \(f_1\): T1, IR, FLAIR; \(f_2\): \(f_1 +\) (x, y, z); \(f_3\): \(f_1 +\) FA, MD; \(f_4\): \(f_2 +\) FA, MD; \(f_5\): \(f_4 +\) \(C_L\), \(C_P\), \(C_S\), AD, RD.
The results of Table 2 demonstrate that our proposed VA-guided feature selection can achieve similar or slightly better performance than the automated feature selection, presented by Kuijf et al. [3]. The two best performing feature sets of Kuijf et al. used 8 (\(f_4\)) and 13 (\(f_5\)) features, while our current two best methods use 7 (\(s_2\)) and 8 (\(s_4\)) features only, with comparable results. Our approach allows to discard \(C_L\), \(C_P\), AD and IR, which do not contribute in the classification; hereby, saving scanning and also computational time.
Evaluation of Classification. To evaluate the classification outcome, we introduce the results of the two best performing classifiers, \(s_2\) and \(s_4\), into the VA system. One of the goals is to explore and analyze the parts of the WMHs that are missed, but also to understand better how these classifiers work and how they can be improved. From an initial inspection, it results that classifier \(s_2\) is restricted to the core of the WMHs, while \(s_4\) detects an extension of it. The WMH core is always detected by both classifiers, as it has consistent imaging characteristics and is well-clustered in the t-SNE embeddings. In subjects with bigger WMHs, \(s_4\) misses only small or thin structures and part of the periphery. In subjects with smaller WMHs, there is a tendency to miss periphery parts and posterior structures more often than the anterior. For bigger WMHs, the core differs in T1, MD, RD with the missed structures. Also, the latter are not as good clustered in the t-SNE embeddings as the core, i.e., they are not coherent in their imaging characteristics. As WMHs become smaller, the influence of T1 becomes less strong, while MD and RD seem to become more important.
5 Discussion and Conclusions
We proposed a user-driven pipeline for aiding the design of classifiers, focusing on WMH segmentation. Using VA and the cognitive skills of an expert user, we initially identified the list of features (MD, RD, T1, FLAIR, and secondarily, FA and \(C_S\)) that are suitable for the separation of WMHs. Then, this list was used for WMH classification. In respect of previous work [3], our results are comparable. Yet, our results are not achieved through trial-and-error, but after a justifiable and understandable, interactive feature selection. Additionally, our approach requires less features, which allows to skip several imaging sequences, making the feature calculation less computationally intensive and time consuming. For example, we concluded that \(C_L\), \(C_P\), AD and IR can be omitted, which saves valuable scanning time (IR: 3:49.6 min).
After classification, we evaluated the classifier outcome in the VA system. The periphery is constantly missed. Thin and small structures can be missed due to partial volume effect, while the MNI152-normalized spatial coordinates can influence the separation of posterior or anterior WMHs. For certain subjects, the missed structures have intrinsically different imaging characteristics. In this case, more features, such as texture or tensor information, should be further investigated. The performance of the classifier could be further improved by adding additional post-processing, to remove false positive detection, which was not performed here, to be comparable to Kuijf et al. [3]. Also, it would be interesting to investigate what happens, when our VA-selected features are used with more sophisticated classification algorithms.
In the entire pipeline, the user interacts and guides the analysis. This has the advantage that the cognitive capabilities of the user, which are not easily automatized, can be included in feature selection. However, the results are user-dependent and it remains important to analyze the bias introduced by the user. Although t-SNE is widely used [13] for understanding high dimensional data, errors can also be introduced due to its use. Adding more features for exploration in the VA system, such as textural features or information from tensors, could give interesting results. However, certain visualizations of the VA system do not scale well to a high number of features; thus, new visualizations would be needed to tackle hundreds of features. Finally, evaluating the use of the pipeline with a user study, to define its general usefulness, is another point for future work. Nevertheless, employing VA in the design of classifiers has potential for better understanding the data under exploration, and for obtaining more insight into classifiers and the frequently exploding set of imaging features.
Notes
- 1.
An interactive demo can be found here: https://vimeo.com/170609498
References
Pantoni, L.: Cerebral small vessel disease: from pathogenesis and clinical characteristics to therapeutic challenges. Lancet Neurol. 9(7), 689–701 (2010)
Anbeek, P., Vincken, K.L., van Osch, M.J., Bisschops, R.H., van der Grond, J.: Probabilistic segmentation of white matter lesions in MR imaging. NeuroImage 21(3), 1037–1044 (2004)
Kuijf, H.J., et al.: The added value of diffusion tensor imaging for automated white matter hyperintensity segmentation. In: O’Donnell, L., Nedjati-Gilani, G., Rathi, Y., Reisert, M., Schneider, T. (eds.) Computational Diffusion MRI. Mathematics and Visualization, pp. 45–53. Springer International Publishing, Switzerland (2014)
Maillard, P., Carmichael, O., Harvey, D., Fletcher, E., Reed, B., Mungas, D., DeCarli, C.: FLAIR and diffusion MRI signals are independent predictors of white matter hyperintensities. Am. J. Neuroradiol. 34(1), 54–61 (2013)
Raidou, R.G., Van Der Heide, U.A., Dinh, C.V., Ghobadi, G., Kallehauge, J.F., Breeuwer, M., Vilanova, A.: Visual analytics for the exploration of tumor tissue characterization. Comput. Graph. Forum 34(3), 11–20 (2015)
Keim, D.A., Mansmann, F., Schneidewind, J., Thomas, J., Ziegler, H.: Visual analytics: scope and challenges. In: Simoff, S.J., Böhlen, M.H., Mazeika, A. (eds.) Visual Data Mining: Theory, Techniques and Tools for Visual Analytics. LNCS, vol. 4404, pp. 76–90. Springer, Heidelberg (2008). doi:10.1007/978-3-540-71080-6_6
Van der Maaten, L., Hinton, G.: Visualizing data using t-SNE. J. Mach. Learn. Res. 9(2579–2605), 85 (2008)
Mendrik, A.M., Vincken, K.L., Kuijf, H.J., Breeuwer, M., Bouvy, W.H., de Bresser, J., Jog, A.: MRBrainS challenge: online evaluation framework for brain image segmentation in 3T MRI scans. Comput. Intell. Neurosc. No. 813696, p. 16 (2015). Special issue on Simulation and Validation in Brain Image Analysis
Klein, S., Staring, M., Murphy, K., Viergever, M.A., Pluim, J.P.: Elastix: a toolbox for intensity-based medical image registration. IEEE Trans. Med. Imag. 29(1), 196–205 (2010)
Leemans, A., Jeurissen, B., Sijbers, J., Jones, D.K.: ExploreDTI: a graphical toolbox for processing, analyzing, and visualizing diffusion MR data. In: Proceedings of the 17th Annual Meeting of Intl Soc Mag Reson Med, Vol. 209, p. 3537 (2009)
Westin, C.F., Maier, S.E., Mamata, H., Nabavi, A., Jolesz, F.A., Kikinis, R.: Processing and visualization for diffusion tensor MRI. Med. Image Anal. 6(2), 93–108 (2002)
Fonov, V.S., Evans, A.C., McKinstry, R.C., Almli, C.R., Collins, D.L.: Unbiased nonlinear average age-appropriate brain templates from birth to adulthood. NeuroImage 47, S102 (2009)
Amir, E.A.D., Davis, K.L., Tadmor, M.D., Simonds, E.F., Levine, J.H., Bendall, S.C., Pe’er, D.: viSNE enables visualization of high dimensional single-cell data and reveals phenotypic heterogeneity of leukemia. Nature Biotechnol. 31(6), 545–552 (2013)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing AG
About this paper
Cite this paper
Raidou, R.G. et al. (2016). Employing Visual Analytics to Aid the Design of White Matter Hyperintensity Classifiers. In: Ourselin, S., Joskowicz, L., Sabuncu, M., Unal, G., Wells, W. (eds) Medical Image Computing and Computer-Assisted Intervention – MICCAI 2016. MICCAI 2016. Lecture Notes in Computer Science(), vol 9901. Springer, Cham. https://doi.org/10.1007/978-3-319-46723-8_12
Download citation
DOI: https://doi.org/10.1007/978-3-319-46723-8_12
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-46722-1
Online ISBN: 978-3-319-46723-8
eBook Packages: Computer ScienceComputer Science (R0)