Abstract
Daily (circadian) clocks have evolved to coordinate behaviour and physiology around the 24-h day. Most models of the eukaryotic circadian oscillator have focused principally on transcription/translation feedback loop (TTFL) mechanisms, with accessory cytosolic loops that connect them to cellular physiology. Recent work, however, questions the absolute necessity of transcription-based oscillators for circadian rhythmicity. The recent discovery of reduction-oxidation cycles of peroxiredoxin proteins, which persist even in the absence of transcription, have prompted a reappraisal of current clock models in disparate organisms. A novel mechanism based on metabolic cycles may underlie circadian transcriptional and cytosolic rhythms, making it difficult to know where one oscillation ends and the other begins.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
Introduction
Daily biological clocks provide living organisms with temporal organisation over a 24-h timescale. Organisms from bacteria to humans have evolved these rhythms to adapt their physiology to the solar cycle and anticipate the availability of resources (e.g., food and light; Bass 2012). Despite their presence in evolutionarily disparate organisms, the molecules underlying the clockwork seemed to be different in these organisms. This finding has given rise to the identification of “clock genes” that oscillate with 24-h periods but that are not related in their DNA or protein sequences, except in some cases when comparing fruit flies and mammals. This has led to the notion of the divergent evolution of different clock circuits in various model organisms, the only link being the way in which the components are joined together in a negative feedback loop topology (Rosbash 2009; Fig. 1).
The dominant model organisms used for the study of circadian rhythms are shown. Below are lists of the more important clock genes that are/were thought to mediate negative feedback loops in these organisms. At the DNA or protein sequence levels, there is no evolutionary conversation beyond some in fly and mammals
Challenging Transcriptional Models of the Clockwork
An increasing number of studies, both in higher and lower organisms, have questioned the necessity of a functional transcriptional oscillator for cellular rhythmicity. Circadian transcription is stochastic in mammalian cells (Suter et al. 2011); therefore, one would expect that perturbation of transcription during cell division would result in phase variability, which is not seen. In line with this finding, global inhibition of transcription with actinomycin D and α-amanitin has revealed the robustness of circadian oscillators to such severe perturbations, and single cells exhibit bioluminescence rhythms even when the transcription rate is reduced by ~70 % (Dibner et al. 2009).
Perhaps more importantly, studies showing that constitutive expression, or deletion, of “clock genes” does not abolish circadian rhythms call into question the importance of transcription in current clock models. In flies, expression of both per and tim under the control of a constitutive promoter can affect circadian rhythms. However, ~50 % of the flies still exhibited robust behavioral rhythms (Yang and Sehgal 2001). Similarly, fungi can exhibit conidiation (spore formation) rhythms in the absence of central components of their feedback loop [involving the frequency (frq) gene] (Lakin-Thomas 2006). Moreover, in some organisms, the dominant mechanism regulating circadian rhythms seems to be post-transcriptional, as exemplified by the circadian control of translation of luciferin binding protein (LBP) in the unicellular alga Gonyaulax polyedra (Morse et al. 1989; Mittag et al. 1994).
In mammals, the situation is difficult to dissect since circadian genes often have multiple homologues; therefore, double-mutant animals are generally needed to observe a behavioral phenotype. Bmal1 was thought to be the only exception to this, with its suppression leading to clear behavioural arrhythmicity (Ko et al. 2006). Constitutive brain-specific expression of Bmal1 in knock-out animals is, however, able to restore behavioral rhythmicity, questioning the necessity of rhythmic Bmal1 transcription (McDearmon et al. 2006). In addition, brain-specific knockout of Bmal1 expression produces gross pathology, with a striking abundance of activated microglia in the brains of mice, which gets progressively worse over the first 6 months of life. This finding makes it extremely difficult to dissociate the effects if BMAL1 as a generically important transcription factor from those specifically related to the malfunctioning of a biological clock (Musiek et al. 2013).
Even more importantly, imaging of suprachiasmatic nucleus (SCN) slices from arrhythmic Bmal1−/− and Cry1−/−Cry2−/− animals with bioluminescence reporters revealed the persistence of low amplitude rhythms in individual neurons (Ko et al. 2010; Maywood et al. 2011). As recently shown, it is likely that developmental effects underlie the apparent arrhythmicity that is observed when adult animals are assayed, as is the case in most experimental paradigms (Ono et al. 2013).
There is thus considerable evidence that current transcription-translation feedback loops cannot account for the multiple lines of experimental evidence that have revealed circadian oscillations in the presence of inactivated feedback loops or indeed in their absence.
Non-transcriptional Clock Mechanisms
The experimental anomalies highlighted above suggest that other mechanisms are required to fully explain the molecular basis of circadian timekeeping. It is indeed worth underscoring that transcriptional mechanisms were regarded as only one of the several possibilities that were investigated before the discovery of “clock genes” (Edmunds 1988).
An instructive outlier in clock research is the macroscopic unicellular alga Acetabularia, which can maintain self-sustained circadian rhythms in photosynthetic activity when its nucleus is removed by cutting off its nucleus-containing rhizoid process (Sweeney and Haxo 1961). Intriguingly, its nucleus is able to dictate the phase of oscillation but is dispensable for entrainment and phase shifting (Schweiger et al. 1964). Moreover, inhibition of transcription with Actinomycin D did not suppress rhythms in either nucleated or enucleated Acetabularia cells, although the former surprisingly lost rhythmicity after 2 weeks under these conditions (Mergenhagen and Schweiger 1975). Similarly, platelets were used to show that glutathione exhibited circadian oscillations relying on de novo synthesis of this important cellular reductant (Radha et al. 1985), again in the absence of a nucleus. These examples point to the fact that current circadian models cannot explain issues raised almost 40 years ago, in some cases, suggesting the existence of non-transcriptional rhythms.
How can we reconcile these seemingly opposite views? One way is to view transcription and translation in the current models as having limited roles in setting the pace of the oscillator and to note that they are needed to maintain the levels of clock proteins and to control circadian output functions. Accordingly, post-translational modifications of known clock proteins could be the fundamental oscillator, but the transcriptional oscillator would be important for robustness and could amplify post-translational oscillations. In fact, such a model exists in cyanobacteria, in which the master transcriptional regulator KaiC is part of its post-translational oscillator.
An alternative point of view is that circadian timekeeping might have evolved more than one clock in the cell to meet the requirements of precision, robustness and stability. In this case, the known transcriptional oscillator would be coupled to a post-translational oscillator. Post-translational modifications are an integral feature of the current transcription-translation feedback models, but a definitive post-translational oscillator has not yet been identified in eukaryotic species. The recent discovery of oxidation cycles in peroxiredoxin proteins (PRDXs) offers a new window on non-transcriptional rhythms in higher organisms (O’Neill and Reddy 2011; O’Neill et al. 2011; Edgar et al. 2012; Olmedo et al. 2012). More importantly, this finding immediately suggests a common phylogenetic origin for circadian timekeeping mechanisms in virtually all species relying on oxygen for energy metabolism (Edgar et al. 2012).
PRDX Rhythms
PRDXs are an antioxidant protein family involved in hydrogen peroxide metabolism and signalling (Hall et al. 2009). Their catalytic mechanism involves the oxidation of a catalytic cysteine residue in the enzymes’ active site to sulfenic acid (Cys-SOH), which then forms a disulfide bond with another non-catalytic (and so-called ‘resolving’) cysteine residue. The thioredoxin system usually completes the cycle by reducing this disulfide bond while oxidising a molecule of NADPH. This catalytic loop has rapid turnover and allows the maintenance of low levels of intracellular hydrogen peroxide.
So-called ‘typical 2-Cys PRDXs,’ a subclass of PRDXs whose basic functional unit is a homodimer in which catalytic and resolving cysteine residues belong to different molecules of PRDX, are the main players implicated in circadian cycles. These can undergo further oxidation of their catalytic cysteine to sulfinic and sulfonic acid forms (Cys-SO2/3H). The ‘over-oxidised’ Cys-SO2H residues can be slowly recycled through adenosine triphosphate (ATP)-dependent reduction by sulfiredoxin (Rhee et al. 2007), whereas further oxidation to Cys-SO3H (termed ‘hyper-oxidation’) is thought to be irreversible.
Human red blood cells do not undergo transcription since they lack nuclei in their mature form. In these unique cells, PRDXs exhibit circadian accumulation of their dimeric over-oxidised form (PRDX-SO2H) over several days (O’Neill and Reddy 2011). Such rhythms fulfil all criteria for circadian rhythms: (1) persistence in constant conditions; (2) the ability to be entrained (via temperature cycles in this case); and (3) temperature compensation (the clock does not run faster in higher temperatures). In addition, these redox rhythms are accompanied by oscillations in haemoglobin oxidation and metabolic variables, including NADH and NADPH (Fig. 2). Similar results have also recently been found in mouse red blood cells (Cho et al. 2014).
A range of circadian oscillations in human red blood cells. (a) Oxidation of PRDXs occurs on a circadian basis in cells maintained in constant conditions (in the absence of external temporal cues) for at least 3 consecutive days. (b) Dynamic changes in the equilibrium of haemoglobin forms (tetramer vs. dimer states) oscillate according to a 24-h rhythm in vitro. (c) Oscillation of the key cellular reductants, NADH and NADPH, in red blood cells
Rhythms similar to these are also present in the unicellular alga Ostreococcus tauri, even when transcription is inhibited by prolonged darkness (O’Neill et al. 2011); they are autophototrophic, requiring light for synthesis of most cellular substrates including RNA. Moreover, the deep phylogenetic conservation of PRDX redox rhythms extends to include fungal, plant, bacterial and even archaeal species. Critically, such rhythms are not dependent on previously identified clock genes, since mutants lacking circadian components maintain redox oscillations, albeit slightly phase-shifted (Edgar et al. 2012).
The phylogenetic conservation of PRDX rhythms suggests that primordial redox oscillators probably evolved following the Great Oxidation Event 2.5 billion years ago. At this time, photosynthetic bacteria are thought to have acquired the ability to produce oxygen from water, which caused a dramatic rise in Earth’s atmospheric oxygen. Rhythmic production of oxygen and reactive oxygen species (ROS) by sunlight may therefore have been a critical driving force in the co-evolution of clock mechanisms and ROS removal systems that could anticipate, and thus resonate with, externally driven redox cycles (Bass 2012; Edgar et al. 2012; Fig. 3).
Phylogenetic origins of circadian oscillatory systems. A timeline is shown at the top of the schematic, with the geological era illustrated. A schematic phylogenetic tree shows the origins of each organism studied, stemming from the last universal common ancestor (LUCA). The putative epoch over which each oscillator system has existed is illustrated by the labeled bars. CK1/2, casein kinase 1 or 2; GSK3, glycogen synthase kinase 3; SOD, superoxide dismutase (Adapted from Edgar et al. 2012)
Redox and Metabolic Clocks in Eukaryotes
There is clearly an interplay between circadian and metabolic cycles, and there is good evidence of reciprocal effects that disruption of one cycle has on the other at physiological and molecular levels (Bass 2012; Rey and Reddy 2013). High-fat diet, for example, lengthens the behavioral period of rhythms in mice and changes the expression pattern of clock genes (Kohsaka et al. 2007). Conversely, healthy patients subjected to 3 weeks of circadian disruption exhibit pre-diabetic symptoms (Buxton et al. 2012). The growing evidence suggesting that circadian rhythms are fundamentally metabolic requires that currently understood transcriptional oscillations are tightly coupled to metabolic cycles. This hypothesis is strongly supported by the numerous examples of accessory loops embedding the circadian transcriptional clock within cellular metabolism (Fig. 4).
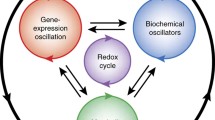
Links between transcriptional, cytosolic, and metabolic cycles. Cytosolic processes are thought to be part of the transcription/translation feedback loop (TTFL). The latter are involved mainly in redox and energy metabolism and form accessory loops that are controlled by the TTFL oscillator and, in turn, feed back to it. Abbreviations: CREB cAMP response element-binding protein, GRE glucose response element, PARP poly(ADP-ribose) polymerase, PTMs posttranslational modifications (Adapted from Reddy and Rey 2014)
An accessory loop involving NAD+/NADH is likely to play an important role in connecting cytosolic and compartment-specific redox states to transcriptional clock components such as PER2 (Asher et al. 2008) and CLOCK/BMAL1 (Rutter et al. 2001; Nakahata et al. 2008; Asher et al. 2010; Yoshii et al. 2013). In addition, other redox-sensitive mechanisms have been identified in the clockwork and, in particular, the heme-sensing transcriptional regulators (Dioum et al. 2002; Yin et al. 2007; Gupta et al. 2011).
Even in early molecular studies of the circadian clock, before “clock genes” had been discovered in any model organism, rhythms in redox had been reported. For example, in plants, NADP+:NADPH ratio exhibited circadian cycles in seedlings kept in constant darkness (Wagner and Frosch 1974). Several studies in rodents showed that redox parameters, including the glutathione redox ratio, were diurnally regulated in the liver, although it is possible that these oscillations might have been partially driven by food intake (Isaacs and Binkley 1977a, b; Robinson et al. 1981; Kaminsky et al. 1984; Belanger et al. 1991). Nevertheless, human platelets kept in vitro showed circadian rhythms in glutathione content (Radha et al. 1985), suggesting that feeding cycles might internally resonate with these cell-autonomous biochemical rhythms.
The hypothesis that metabolic cycles might be a fundamental mechanism underlying biological clocks has been proposed based on both theoretical and experimental observations (Roenneberg and Merrow 1999). Potential evidence for this hypothesis in mammals has come from the McKnight and Sagami groups, who have shown that BMAL1/CLOCK DNA-binding activity can be modulated in vitro by the redox poise of NAD(P)+/NAD(P)H coenzymes (Rutter et al. 2001; Yoshii et al. 2013). In addition, the action of BMAL1/CLOCK on the NAD+-producing enzymes lactate dehydrogenase (Ldh) and Nampt could potentially feed back onto intracellular redox balance (Rutter et al. 2001; Nakahata et al. 2009; Ramsey et al. 2009). These results still require in vivo confirmation, given the relatively high concentration (millimolar range) of the coenzymes used in in vitro assays previously (Rutter et al. 2001; Yoshii et al. 2013). The recent discovery of PRDX oscillations in non-transcriptional systems, however, offers supportive evidence that redox cycles can function as circadian oscillators in their own right.
It is evident that, in organisms in which metabolic oscillations have been found but transcription-translation feedback loops have not, as in the worm Caenorhabditis elegans, insights into metabolic oscillatory mechanisms may be easier to come by. It is thus possible that metabolic oscillations could drive PRDX oscillations in the absence of known transcriptional feedback oscillators (Olmedo et al. 2012). So-called accessory loops, including NAD+/NADH and NADP+/NADPH cycles, are potential candidates for self-sustained metabolic oscillators, but further studies of their oscillatory properties in clock mutant backgrounds will be of great interest to identify bona fide components of metabolic oscillators. However, this assumes that deletion of important circadian-relevant transcription factors itself does not lead to abhorrent redox changes in cells and tissues, as is the case in Bmal1−/− animals (Kondratov et al. 2006), which could compromise redox oscillations indirectly.
Conclusion
Metabolic non-transcriptional cycles clearly interlock with transcriptional processes in the circadian system. The peroxiredoxin system could be part of an uncharacterised metabolic oscillator, given its broad phylogenetic conservation and its slow kinetics, which is compatible with 24-h rhythmicity. Establishing the molecular links between fundamental cellular redox metabolism and transcriptional components of the clockwork remains an exciting challenge in the field.
References
Asher G, Gatfield D, Stratmann M, Reinke H, Dibner C, Kreppel F, Mostoslavsky R, Alt FW, Schibler U (2008) SIRT1 regulates circadian clock gene expression through PER2 deacetylation. Cell 134:317–328
Asher G, Reinke H, Altmeyer M, Gutierrez-Arcelus M, Hottiger MO, Schibler U (2010) Poly(ADP-ribose) polymerase 1 participates in the phase entrainment of circadian clocks to feeding. Cell 142:943–953
Bass J (2012) Circadian topology of metabolism. Nature 491:348–356
Belanger PM, Desgagné M, Bruguerolle B (1991) Temporal variations in microsomal lipid peroxidation and in glutathione concentration of rat liver. Drug Metab Dispos 19:241–244, Available at: http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=1673408&retmode=ref&cmd=prlinks
Buxton OM, Cain SW, O’Connor SP, Porter JH, Duffy JF, Wang W, Czeisler CA, Shea SA (2012) Adverse metabolic consequences in humans of prolonged sleep restriction combined with circadian disruption. Sci Translat Med 4:129ra43
Cho CS, Yoon HJ, Kim JY, Woo HA, Rhee SG (2014) Circadian rhythm of hyperoxidized peroxiredoxin II is determined by hemoglobin autoxidation and the 20S proteasome in red blood cells. Proc Natl Acad Sci USA 111:12043–12048
Dibner C, Sage D, Unser M, Bauer C, d'Eysmond T, Naef F, Schibler U (2009) Circadian gene expression is resilient to large fluctuations in overall transcription rates. EMBO J 28:123–134
Dioum EM, Dioum EM, Rutter J, Tuckerman JR, Gonzalez G, Gilles-Gonzalez M-A, McKnight SL (2002) NPAS2: a gas-responsive transcription factor. Science 298:2385–2387
Edgar RS, Green EW, Zhao Y, van Ooijen G, Olmedo M, Qin X, Xu Y, Pan M, Valekunja UK, Feeney KA, Maywood ES, Hastings MH, Baliga NS, Merrow M, Millar AJ, Johnson CH, Kyriacou CP, O’Neill JS, Reddy AB (2012) Peroxiredoxins are conserved markers of circadian rhythms. Nature 485:459–464
Edmunds LNJ (1988) Cellular and molecular bases of biological clocks. Springer, New York, p 514
Gupta N, Gupta N, Ragsdale SW, Ragsdale SW (2011) Thiol-disulfide redox dependence of heme binding and heme ligand switching in nuclear hormone receptor rev-erb. J Biol Chem 286:4392–4403
Hall A, Karplus PA, Poole LB (2009) Typical 2-Cys peroxiredoxins—structures, mechanisms and functions. FEBS J 276:2469–2477
Isaacs J, Binkley F (1977a) Glutathione dependent control of protein disulfide-sulfhydryl content by subcellular fractions of hepatic tissue. Biochim Biophys Acta 497:192–204, Available at: http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=557349&retmode=ref&cmd=prlinks
Isaacs JT, Binkley F (1977b) Cyclic AMP-dependent control of the rat hepatic glutathione disulfide-sulfhydryl ratio. Biochim Biophys Acta 498:29–38, Available at: http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=18207&retmode=ref&cmd=prlinks
Kaminsky YG, Kosenko EA, Kondrashova MN (1984) Analysis of the circadian rhythm in energy metabolism of rat liver. Int J Biochem 16:629–639, Available at: http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=6468728&retmode=ref&cmd=prlinks
Ko CH, Ko CH, Takahashi JS (2006) Molecular components of the mammalian circadian clock. Hum Mol Genet 15:R271–R277
Ko CH, Yamada YR, Welsh DK, Buhr ED, Liu AC, Zhang EE, Ralph MR, Kay SA, Forger DB, Takahashi JS (2010) Emergence of noise-induced oscillations in the central circadian pacemaker. PLoS Biol 8:e1000513
Kohsaka A, Laposky AD, Ramsey KM, Estrada C, Joshu C, Kobayashi Y, Turek FW, Bass J (2007) High-fat diet disrupts behavioral and molecular circadian rhythms in mice. Cell Metab 6:414–421
Kondratov RV, Kondratova AA, Gorbacheva VY, Vykhovanets OV, Antoch MP (2006) Early aging and age-related pathologies in mice deficient in BMAL1, the core component of the circadian clock. Genes Dev 20:1868–1873
Lakin-Thomas PL (2006) Transcriptional feedback oscillators: maybe, maybe not. J Biol Rhythms 21:83–92
Maywood ES, Chesham JE, O’Brien JA, Hastings MH (2011) A diversity of paracrine signals sustains molecular circadian cycling in suprachiasmatic nucleus circuits. Proc Natl Acad Sci USA 108:14306–14311
McDearmon EL, Patel KN, Ko CH, Walisser JA, Schook AC, Chong JL, Wilsbacher LD, Song EJ, Hong H-K, Bradfield CA, Takahashi JS (2006) Dissecting the functions of the mammalian clock protein BMAL1 by tissue-specific rescue in mice. Science 314:1304–1308
Mergenhagen D, Schweiger HG (1975) The effect of different inhibitors of transcription and translation on the expression and control of circadian rhythm in individual cells of Acetabularia. Exp Cell Res 94:321–326, Available at: http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=1193133&retmode=ref&cmd=prlinks
Mittag M, Lee DH, Hastings JW (1994) Circadian expression of the luciferin-binding protein correlates with the binding of a protein to the 3′ untranslated region of its mRNA. Proc Natl Acad Sci USA 91:5257–5261
Morse D, Milos PM, Roux E, Hastings JW (1989) Circadian regulation of bioluminescence in Gonyaulax involves translational control. Proc Natl Acad Sci USA 86:172–176
Musiek ES, Lim MM, Yang G, Bauer AQ, Qi L, Lee Y, Roh JH, Ortiz-Gonzalez X, Dearborn JT, Culver JP, Herzog ED, Hogenesch JB, Wozniak DF, Dikranian K, Giasson BI, Weaver DR, Holtzman DM, Fitzgerald GA (2013) Circadian clock proteins regulate neuronal redox homeostasis and neurodegeneration. J Clin Invest 123:5389–5400
Nakahata Y, Kaluzova M, Grimaldi B, Sahar S, Hirayama J, Chen D, Guarente LP, Sassone-Corsi P (2008) The NAD+-dependent deacetylase SIRT1 modulates CLOCK-mediated chromatin remodeling and circadian control. Cell 134:329–340
Nakahata Y, Sahar S, Astarita G, Kaluzova M, Sassone-Corsi P (2009) Circadian control of the NAD+ salvage pathway by CLOCK-SIRT1. Science 324:654–657
O’Neill JS, Reddy AB (2011) Circadian clocks in human red blood cells. Nature 469:498–503
O’Neill JS, van Ooijen G, Dixon LE, Troein C, Corellou F, Bouget F-Y, Reddy AB, Millar AJ (2011) Circadian rhythms persist without transcription in a eukaryote. Nature 469:554–558
Olmedo M, O’Neill JS, Edgar RS, Valekunja UK, Reddy AB, Merrow M (2012) Circadian regulation of olfaction and an evolutionarily conserved, nontranscriptional marker in Caenorhabditis elegans. Proc Natl Acad Sci USA 109:20479–20484
Ono D, Honma S, Honma K-I (2013) Cryptochromes are critical for the development of coherent circadian rhythms in the mouse suprachiasmatic nucleus. Nat Commun 4:1666
Radha E, Hill TD, Rao GH, White JG (1985) Glutathione levels in human platelets display a circadian rhythm in vitro. Thromb Res 40:823–831, Available at: http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=4089839&retmode=ref&cmd=prlinks
Ramsey KM, Yoshino J, Brace CS, Abrassart D, Kobayashi Y, Marcheva B, Hong HK, Chong JL, Buhr ED, Lee C, Takahashi JS, Imai SI, Bass J (2009) Circadian clock feedback cycle through NAMPT-mediated NAD+ biosynthesis. Science 324:651–654
Reddy AB, Rey G (2014) Metabolic and nontranscriptional circadian clocks: eukaryotes. Annu Rev Biochem 83:165–189
Rey G, Reddy AB (2013) Connecting cellular metabolism to circadian clocks. Trends Cell Biol 23:234–241
Rhee SG, Jeong W, Chang TS, Woo HA (2007) Sulfiredoxin, the cysteine sulfinic acid reductase specific to 2-Cys peroxiredoxin: its discovery, mechanism of action, and biological significance. Kidney Int Suppl 106:S3–S8
Robinson JL, Foustock S, Chanez M, Bois-Joyeux B, Peret J (1981) Circadian variation of liver metabolites and amino acids in rats adapted to a high protein, carbohydrate-free diet. J Nutr 111:1711–1720, Available at: http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=7288496&retmode=ref&cmd=prlinks
Roenneberg T, Merrow M (1999) Circadian systems and metabolism. J Biol Rhythms 14:449–459
Rosbash M (2009) The implications of multiple circadian clock origins. PLoS Biol 7:e62
Rutter J, Reick M, Wu LC, McKnight SL (2001) Regulation of clock and NPAS2 DNA binding by the redox state of NAD cofactors. Science 293:510–514. doi:10.1126/science.1060698
Schweiger E, Wallraff HG, Schweiger HG (1964) Endogenous circadian rhythm in cytoplasm of Acetabularia: influence of the nucleus. Science 146:658–659
Suter DM, Suter DM, Molina N, Molina N, Gatfield D, Gatfield D, Schneider K, Schneider K, Schibler U, Schibler U, Naef F, Naef F (2011) Mammalian genes are transcribed with widely different bursting kinetics. Science 332:472–474
Sweeney BM, Haxo FT (1961) Persistence of a photosynthetic rhythm in enucleated Acetabularia. Science 134:1361–1363
Wagner E, Frosch S (1974) Cycles in plants. J Interdis Cycle Res 5:231–239
Yang Z, Sehgal A (2001) Role of molecular oscillations in generating behavioral rhythms in Drosophila. Neuron 29:453–467
Yin L, Wu N, Curtin JC, Qatanani M, Szwergold NR, Reid RA, Waitt GM, Parks DJ, Pearce KH, Wisely GB, Lazar MA (2007) Rev-erbalpha, a heme sensor that coordinates metabolic and circadian pathways. Science 318:1786–1789
Yoshii K, Ishijima S, Sagami I (2013) Effects of NAD(P)H and its derivatives on the DNA-binding activity of NPAS2, a mammalian circadian transcription factor. Biochem Biophys Res Commun 437:386–391
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is distributed under the terms of the Creative Commons Attribution-Noncommercial 2.5 License (http://creativecommons.org/licenses/by-nc/2.5/) which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
The images or other third party material in this chapter are included in the work’s Creative Commons license, unless indicated otherwise in the credit line; if such material is not included in the work’s Creative Commons license and the respective action is not permitted by statutory regulation, users will need to obtain permission from the license holder to duplicate, adapt or reproduce the material.
Copyright information
© 2016 The Author(s)
About this chapter
Cite this chapter
Reddy, A.B. (2016). Redox and Metabolic Oscillations in the Clockwork. In: Sassone-Corsi, P., Christen, Y. (eds) A Time for Metabolism and Hormones. Research and Perspectives in Endocrine Interactions. Springer, Cham. https://doi.org/10.1007/978-3-319-27069-2_6
Download citation
DOI: https://doi.org/10.1007/978-3-319-27069-2_6
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-27068-5
Online ISBN: 978-3-319-27069-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)