Abstract
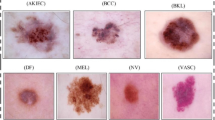
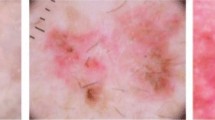
Melanoma is the most severe type of skin cancer due to its ability to cause metastasis. It is more common in black people, often affecting acral regions: palms, soles, and nails. Deep neural networks have shown tremendous potential for improving clinical care and skin cancer diagnosis. Nevertheless, prevailing studies predominantly rely on datasets of white skin tones, neglecting to report diagnostic outcomes for diverse patient skin tones. In this work, we evaluate supervised and self-supervised models in skin lesion images extracted from acral regions commonly observed in black individuals. Also, we carefully curate a dataset containing skin lesions in acral regions and assess the datasets concerning the Fitzpatrick scale to verify performance on black skin. Our results expose the poor generalizability of these models, revealing their favorable performance for lesions on white skin. Neglecting to create diverse datasets, which necessitates the development of specialized models, is unacceptable. Deep neural networks have great potential to improve diagnosis, particularly for populations with limited access to dermatology. However, including black skin lesions is necessary to ensure these populations can access the benefits of inclusive technology.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
Notes
- 1.
If you have skin, you can get skin cancer.
- 2.
- 3.
Clinical images can be captured with standard cameras, while dermoscopic images are captured with a device called dermatoscope, that normalize the light influence on the lesion, allowing to capture deeper details.
References
American Cancer Society. Key statistics for melanoma skin cancer (2022). https://www.cancer.org/cancer/melanoma-skin-cancer/about/key-statistics.html
AIM at Melanoma Foundation. What is acral lentiginous melanoma? https://www.aimatmelanoma.org/melanoma-101/types-of-melanoma/cutaneous-melanoma/acral-lentiginous-melanoma/
Memorial Sloan Kettering Cancer Center. Types of melanoma (2022). https://www.mskcc.org/cancer-care/types/melanoma/types-melanoma
Caetano, Y.A., Quinteiro Ribeiro, A.M., da Silva Albernaz, B.R., de Paula Eleutério, I., Fleury Fróes, L.F.: Melanoma acral-estudo clínico e epidemiológico. Surgical & Cosmetic Dermatology (2020)
Rabin, R.C.: Dermatology has a problem with skin color (2020). https://www-nytimes-com.cdn.ampproject.org/c/s/www.nytimes.com/2020/08/30/health/skin-diseases-black-hispanic.amp.html
Menegola, A., Fornaciali, M., Pires, R.: Flávia Vasques Bittencourt, Sandra Avila, and Eduardo Valle. Knowledge transfer for melanoma screening with deep learning, International Symposium on Biomedical Imaging (2017)
Chaves, L., Bissoto, A., Valle, E., Avila, S.: An evaluation of self-supervised pre-training for skin-lesion analysis. In: European Conference on Computer Vision Workshops (2022)
Singh, N.: Decolonising dermatology: why black and brown skin need better treatment. The Guardian, 13 (2020)
DermNet. Fitzpatrick skin phototype (2012). https://dermnetnz.org/topics/skin-phototype
Dermatology Learning Network. Skin cancer in African-Americans (2004). https://www.hmpgloballearningnetwork.com/site/thederm/article/2547
Groh, M., et al.: Evaluating deep neural networks trained on clinical images in dermatology with the fitzpatrick 17k dataset. In: Conference on Computer Vision and Pattern Recognition (2021)
Chanki, Yu., et al.: Acral melanoma detection using a convolutional neural network for dermoscopy images. PloS one (2018)
Lee, S., et al.: Augmented decision-making for acral lentiginous melanoma detection using deep convolutional neural networks. J. Eur. Acad. Dermatology Venereology (2020)
Abbas, Q., Ramzan, F., Ghani, M.U.: Acral melanoma detection using dermoscopic images and convolutional neural networks. Visual Computing for Industry, Biomedicine, and Art (2021)
Alipour, N., Burke, T., Courtney, J.: Skin type diversity: a case study in skin lesion datasets (2023)
Pacheco, A., et al. PAD-UFES-20: a skin lesion dataset composed of patient data and clinical images collected from smartphones. Data in Brief (2020)
Daneshjou, R., et al.: Disparities in dermatology AI performance on a diverse, curated clinical image set. Science Advances (2022)
Bevan, P.J., Atapour-Abarghouei, A.: Detecting melanoma fairly: skin tone detection and debiasing for skin lesion classification. In: MICCAI Workshop on Domain Adaptation and Representation Transfer, pp. 1–11 (2022)
Pakzad, A., Abhishek, K., Hamarneh, G.: Circle: color invariant representation learning for unbiased classification of skin lesions. In: European Conference on Computer Vision (2022)
Rezk, E., Eltorki, M., El-Dakhakhni, W., et al.: Improving skin color diversity in cancer detection: deep learning approach. JMIR Dermatology 5(3), e39143
Deng, J., Dong, W., Socher, R., Li, L.-J., Li, K., Fei-Fei, L.: Imagenet: a large-scale hierarchical image database. In: Conference on Computer Vision and Pattern Recognition (2009)
ISIC Archive (2023). https://www.isic-archive.com
Valle, E., et al.: Data, depth, and design: Learning reliable models for skin lesion analysis. Neurocomputing (2020)
He, K., Zhang, X., Ren, S., Sun, J.: Deep residual learning for image recognition. In: IEEE Conference on Computer Vision and Pattern Recognition, pp. 770–778 (2016)
Grill, J.-B., et al.: Bootstrap your own latent - a new approach to self-supervised learning. In: Advances in Neural Information Processing Systems (2020)
Tian, Y., Sun, C., Poole, B., Krishnan, D., Schmid, V., Isola, P.: What makes for good views for contrastive learning? In: Advances in Neural Information Processing Systems (2020)
He, K., Fan, H., Wu, Y., Xie, S., Girshick, R.: Momentum contrast for unsupervised visual representation learning. In: Conference on Computer Vision and Pattern Recognition (2020)
Chen, T., Kornblith, S., Norouzi, M., Hinton, G.: A simple framework for contrastive learning of visual representations. In: International Conference on Machine Learning (2020)
Caron, M., Misra, I., Mairal, J., Goyal, P., Bojanowski, P., Joulin, A.: Unsupervised learning of visual features by contrasting cluster assignments. Advances in Neural Information Processing Systems (2020)
Kawahara, J., Daneshvar, S., Argenziano, G., Hamarneh., H.: Seven-point checklist and skin lesion classification using multitask multimodal neural nets. IEEE J. Biomed. Health Inf. (2019)
Richard, P.: Usatine and Brian D. Madden. Interactive dermatology atlas (2023). https://www.dermatlas.net
Dermis.net: Dermatology information service available on the internet (2023). https://www.dermis.net/dermisroot/pt/home/index.htm
Dermnet resource (2023). https://dermnetnz.org
Argenziano, G., Fabbrocini, G., Carli, P., De Giorgi, V., Sammarco, E., Delfino, M.: Epiluminescence microscopy for the diagnosis of doubtful melanocytic skin lesions: comparison of the ABCD rule of dermatoscopy and a new 7-point checklist based on pattern analysis. Arch. Dermatol. 134(12), 1563–1570 (1998)
AlKattash, J.A.: Dermaamin. https://www.dermaamin.com
da Silva, S.F.: Atlas dermatologico. http://atlasdermatologico.com.br
Acknowledgments
L. Chaves is funded by Becas Santander/Unicamp - HUB 2022, Google LARA 2021, in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior - Brasil (CAPES) - Finance Code 001. S. Avila is funded by CNPq 315231/2020-3, FAEPEX, FAPESP 2013/08293-7, 2020/09838-0, H.IAAC 01245.013778/2020-21, and Google Award for Inclusion Research Program 2022 (“Dark Skin Matters: Fair and Unbiased Skin Lesion Models”).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2024 Springer Nature Switzerland AG
About this paper
Cite this paper
Barros, L., Chaves, L., Avila, S. (2024). Assessing the Generalizability of Deep Neural Networks-Based Models for Black Skin Lesions. In: Vasconcelos, V., Domingues, I., Paredes, S. (eds) Progress in Pattern Recognition, Image Analysis, Computer Vision, and Applications. CIARP 2023. Lecture Notes in Computer Science, vol 14470. Springer, Cham. https://doi.org/10.1007/978-3-031-49249-5_1
Download citation
DOI: https://doi.org/10.1007/978-3-031-49249-5_1
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-49248-8
Online ISBN: 978-3-031-49249-5
eBook Packages: Computer ScienceComputer Science (R0)