Abstract
The development of new adaptations of CRISPR-based genome editing platforms, such as base editing and prime editing, made it possible to broaden the scope and applications of genome editing in plants. First base editing and, more recently, prime editing evade the creation of double-stranded breaks in deoxyribonucleic acid (DNA) and the requirement of donor template of DNA for repair while enhancing editing efficiency and product purity over CRISPR/Cas9. As base-pair changes in genomic DNA determine many significant agronomic traits, crop varieties can be developed by precisely converting specific single bases in plant genomes. While base editing can introduce specific nucleotide changes, such as transition and transversion mutations in the targeted region, prime editing can create precise insertions, deletions, and all 12 types of point mutations using the “search-and-replace” method.
This chapter provides the basic principles of base editing and prime editing technologies and their practical applications in plants. The chapter also summarizes the recent breakthroughs in applying base and prime editors in diverse plant species, including their use in improving disease resistance, herbicide resistance, nutritional quality, crop yield, and quality. Finally, this chapter aims to clearly understand base editing and prime editing in plants by outlining potential developments.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
1 Base Editing
Base editing is a novel genome editing method that creates transition and transversion mutations at the single-base level without double-stranded DNA breaks, donor templates, or undesirable effects of NHEJ and HDR mechanisms [1, 2] (Fig. 2.1a). Since base editors (BEs) considerably minimize unintended modifications, they show great potential in plant genome editing applications [3]. Base editor combines a catalytically impaired Cas protein with a nucleotide deaminase to convert one base to another at a target locus in DNA or RNA [4, 5]. First, the Cas protein-gRNA complex binds to its target locus in DNA, and then the Cas protein denatures the double-stranded DNA resulting in an R-loop that exposes a small segment of single-stranded DNA [6]. Next, the deaminase enzyme catalyzes the specific base conversion in this single-stranded DNA. Finally, the permanent introduction of single-base substitutions resulted in the target region through DNA repair and replication [5].
Schematic diagrams of base editors. (a) Graphical overview of the transition and transversion base-pair substitutions in base editing, (b) Cytosine base editors (CBE) mediate C-to-T conversion, (c) Adenine base editors (ABE) mediate A-to-G conversion, (d) Glycosylase base editors (CGBE) mediate transversion mutations
The first developed base editors, cytosine base editors (CBEs), convert a cytosine (C) to thymine (T) and guanine (G) to adenine (A) in the opposite strand) in the target region [5] (Fig. 2.1b). In 2016, David Liu group created the first-generation base-editor (CBE1) by fusing a rat cytidine deaminase (rAPOBEC1) to the N terminus of dCas9 (dead Cas9) via a 16 amino acid XTEN linker [5, 7]. Although CBE1 successfully converts C:G to T:A in vitro, the base excision repair mechanism (BER) recognizes any G:U base pair as a mismatch and removes the uracils with the help of uracil N-glycosylases (UNGs) in vivo. To address this limitation and improve its efficiency, CBE2 was developed by adding a uracil DNA glycosylase inhibitor (UGI) to the C-terminus of dCas9 (dead Cas9) via a 4-amino acid linker [5, 8]. With this new version, editing efficiency was increased three times compared to CBE1 due to the inhibition of uracil DNA glycosylase (UDG) in the organism by UGI. To increase editing efficiency, CBE3 was developed by replacing the dCas9 with a nCas9 (H840, HNH catalytic domain) nickase variant. In this new version, nCas9 would induce a nick in the G-containing DNA strand, which activated the cellular mismatch repair (MMR) mechanism [5, 7]. This MMR mechanism replaces the G on the nicked strand with an A, forming a U:A pair with the target strand. The U:A pair was later corrected, leading to the desired T:A substitution. While inhibition of BER by using UGI in CBE2 enhanced editing efficiencies approximately threefold, the nicking strategy of CBE3 increased efficiencies by up to sixfold compared to CBE2 in human cells [5]. CBE in plants was first implemented in rice [9,10,11] and then adapted to various plants species such as wheat [11, 12], Arabidopsis [13, 14], maize [11], potato [15, 16], tomato [15, 16], cotton [17], watermelon [18], soybean [19], apple [20], pear [20], strawberry [21], rapeseed [22], P. patens [23] and poplar [24] quickly CBE-mediated genome editing used for different purposes, such as obtaining disease and herbicide resistance, accelerating crop domestication, and increasing yield and nutrient use efficiency in various plants, has been summarized in Table 2.1 (also reviewed in [25, 26]).
1.1 Improving the Base Editing Efficiency
With a better understanding of the molecular functions of deaminases, adenine base editors (ABEs) are developed for inducing A•T to G•C conversions with high efficiency [27] (Fig. 2.1c). ABEs use engineered transfer RNA adenosine deaminases (TadA) derived from E. coli, which bind to ssDNA and deaminates A into inosine I. The use of ABEs overcomes the limitation of CBEs, which can only edit C or G bases, and provides a broader range of base transformation options. Unlike CBEs, ABEs do not need to suppress the activity of alkyl adenine DNA glycosylase (AAG) [28, 29]. Over time, various optimization strategies were implemented, including TadA mutations and using varying lengths of the linker between TadA and nCas9 (D10A) to enhance the editing efficiencies of ABEs [25, 27, 30]. Various variants (ABE 6.3, ABE 7.8, ABE 7.9, ABEmax, etc.) of ABEs have been developed and implemented in mammalian cells [8, 27] and then rapidly adapted to plant cells. Li et al. [31] used ABE-mediated base editing to edit rice’s acetyl-coenzyme A carboxylase (ACC) gene to confer herbicide resistance. To create A·T to G·C conversion in OsMPK6, OsSERK2, and OsWRKY45 in rice, a florescence-tracking ABE was developed and successfully implemented by obtaining up to 62.3% editing efficiency [32]. In a proof-of-concept study, Kang et al. [33] edited the PDS gene in Arabidopsis thaliana and Brassica napus by creating a single amino acid substitution using ABE. Hua et al. [34] compared two different ABE versions (ABE-P1S and ABE-P1) to increase efficiency in rice. Like CBEs, ABEs were also implemented in various plants, as shown in Table 2.1.
Since CBEs and ABEs mainly generate base transitions, C-to-G base editors (CGBEs) were developed by modifying CBEs to generate a new tool suitable for C·G to-G·C transversion (Fig. 2.1d). Instead of the UGI inhibitor used in CBEs, CGBEs include Uracil-N-glycosylase (UNG), which locates U in the DNA and eliminates it [35], promoting the BER pathway and improving uracil glycosylation. CGBEs were first developed in human cells, including a UNG fused to nCas9 nickase (D10A) and a cytidine deaminase rAPOBEC1 or its engineered form rAPOBEC1 (R33A). Because of the promising editing results of CBEs and ABEs in plants, Sretenovic et al. [24] modified the CGBE method for plants which is proven to work in human cells [36,37,38]. They tested three versions of CGBEs in rice, tomato, and poplar and obtained different editing efficiencies from CGBEs in different plants. Tian et al. [39] developed CGBEs for rice by optimizing the codon of UNG and by using three highly active deaminases, hAID, hA3A, and Anc689. They tested optimized CGBEs in five different rice genes and obtained successful C-to-G conversions with an average frequency of 21.3% [39]. Another CGBE-mediated genome editing in rice was reported by Zeng et al. using the highly active cytidine deaminase evoFENRY and the PAM-relaxed Cas9-nickase variant Cas9n-NG with rice and human UNG [40]. Although their CGBEs achieved C-to-G conversions up to 27.3% in rice, they did not achieve significant C-to-G performance, contrary to previous studies on mammalian cells [40]. Recently, monocot plant-compatible CGBEs were developed in rice protoplasts, and low editing efficiency was obtained [41]. Similar to previous CGBE studies in plants, this study also emphasizes that further improvements are necessary to enhance the editing efficiency of plant CGBEs for versatile applications [41].
As a result of several attempts to increase base editing efficiency, dual base editing technology which combines both ABEs and CBEs into a single base editor, was developed. Dual base editors convert C-G to T-A and A-T to G-C mutations simultaneously in the target site using a single gRNA [42]. Various dual-base editing platforms have been developed for mammalian cells (SPACE, A&C-BEmax, Target-ACEmax, and ACBE) [42,43,44,45] and plants (STEMEs) [46]. STEMEs (saturated targeted endogenous mutagenesis editors), a fusion of nCas9 with both deaminases, APOBEC3A/ecTadA, was first tested in the OsACC gene to obtain herbicide-resistant rice mutants [46].
Applications of base editors in plants are presented in Table 2.1 by highlighting the plant species, target genes, type of BE, purpose of the targeted mutation, and delivery technique of the reagents.
2 Prime Editing
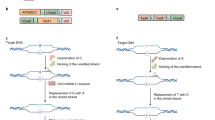
In 2019, David Liu’s group introduced prime editing, a ‘search and replace’ tool that can perform any intended changes, including all 12 possible base-to-base conversions, insertions, and deletions without requiring DSBs or donor DNA templates [78] (Fig. 2.2a). Prime editor is a versatile, precise genome editing tool that is composed of a Cas9 nickase (nCas9; H840A mutation) fused to an engineered reverse transcriptase (RT) and a prime editing guide RNA (pegRNA) containing a primer binding site (PBS) and an RT template (Fig. 2.2b). When pegRNA is delivered into a cell, Cas9 nickase (nCas9; H840A mutation) recognizes and breaks the non-complementary strand of the DNA three bases upstream of the PAM site. The PBS hybridizes with the bases upstream of the nCas9 (H840A)-generated nick and RT template encodes desired edits and directs reverse transcription. Then the new DNA containing the desired edit is integrated, and the unedited strand is repaired to match the edited strand by a cellular DNA repair system [78].
Anzalone et al. [78] presented three versions of prime editing system in their first article on prime editing. The first prime editor (PE1) incorporates wild-type reverse transcriptase from commercial Moloney murine leukemia virus (M-MLV) with Cas9 (H840A) nickase and a pegRNA. Various RT mutations have been investigated to increase the efficiency of prime editing by altering thermostability, processivity, DNA–RNA substrate affinity, and RNaseH activity. PE1 efficiency was increased by harboring engineered M-MLV-RT pentamutant (M-MLV RT (D200N/L603W/T330P/T306K/ W313F)) after which it is called “prime editor 2” (PE2). PE2 enhanced the editing efficiency by 1.6- to 5.1-fold to harbor point mutations on average. In addition, it showed higher editing efficiency in indels and reported that it was compatible with shorter PBS sequences. To further increase the efficiency, an optimized prime editor called PE3 used an additional sgRNA to direct Cas9 (H840A) nickase to produce a nick in the non-edited DNA strand and increased approximately three times the editing efficiency of PE2. Since a high level of indels could be formed depending on the location of the additional sgRNA, the authors resolved this issue by designing the additional sgRNA to target the edited strand but not the original one. This variant of the PE3 system is called PE3b, as it achieved a 13-fold reduction in the average number of indels in human cell lines compared to PE3 while maintaining editing efficiency [78].
2.1 Improving the PE Efficiency
Since it was first published, the possibilities of the use of PE have advanced and broadened. Prime editing mechanism is a complex process influenced by multiple factors, including prime editor structure, pegRNA design, and cellular determinants [79]. Different groups have been developing new strategies to increase the editing efficiency of prime editing in animal and human cells [80]. For example, it is known that the 3′ extension of a pegRNA is critical for priming reverse transcription and templating the desired edit. Nelson et al. [81] discovered that exonucleases could hinder prime editing efficiency by degrading the 3′ extension of pegRNA [81]. pegRNA optimization by incorporating structured RNA motifs to the 3′ terminus of pegRNAs enhanced stability by preventing degradation of the 3′ extension. This strategy is called engineered pegRNAs (epegRNAs) that improved prime editing efficiency three to fourfold in human cells without increasing off-target editing activity [81].
Another strategy was manipulating the DNA repair pathway to increase PE efficiency and reduce indels [82]. The temporary inhibition of DNA mismatch repair (MMR) by MLH1dn significantly improves the effectiveness of PE and reduces the occurrence of indels in various cell types [82]. Transient co-expression of MLH1dn (a dominant-negative variant of the MMR protein MLH1) with PE2 and PE3 yielded PE4 (PE2 + MLH1dn) and PE5 (PE3 + MLH1dn), respectively [82]. PE4 and PE5 versions enhanced the editing efficiency by sevenfold over PE2 and twofold over PE3, respectively [79, 82]. With further efforts to enhance prime editing efficiency, an improved prime editor architecture, “PEmax,” was obtained by optimizing RT codon usage, Cas9 mutations, linker length /composition, and nuclear localization signals (NLS) tags based on the PE2 protein. The combination of PEmax with PE4/PE5 systems (known as PE4max and PE5max, respectively) along with epegRNAs significantly improved editing efficiency [80, 82, 83]. Additional optimization strategies were used in various cell types, including prime editing protein engineering, pegRNA structure, stability improvements, repair mechanism suppression, and two-pegRNA implementation [80].
Most pioneering studies in prime editing have been implemented in animal and human cells, and applications of prime editing in plants are mostly proof of concept and optimization studies (Table 2.2). The first report on prime editing in plants was published by Lin et al. [84]. They obtained a variety of edits, including insertions up to 15 bp in wheat and rice protoplast, by optimizing codon, promoter, and editing conditions. This study was followed by studies demonstrating the applicability of prime editors in various plant species, including tomato [85], potato [86], maize [87], Arabidopsis [88], N. benthamiana [88], and rice [89, 90]. However, these studies showed that the application of prime editing is limited by the low efficiency and optimization studies required to reach its full potential. Therefore, various approaches have been rapidly developed and applied in plants to overcome the limitations of prime editors [91, 92].
Dual-pegRNA strategy employs two pegRNAs in trans to simultaneously encode the same edits, increasing PE efficiency by expanding the size and type of genomic mutations in rice [93]. Lin et al. [93] also optimized the melting temperature (Tm) of the PBS combined with a dual-pegRNA strategy and increased the editing efficiency from 2.9-fold to 17.4-fold in rice protoplasts [93]. Xu et al. [94] reported that changing the C-terminal reverse transcriptase Cas9 nickase fusion with N- terminal fusion improved the editing efficiency at some target sites in rice and maize [94]. In addition to this modification, codon optimization of M-MLV RT by introducing multiple-nucleotide substitutions enhanced editing frequency up to 24.3% and 6.2% in rice and maize, respectively [94]. Zong et al. [95] indicated that engineering the M-MLV reverse transcriptase by deletion of the RT RNase H domain and the addition of a virus-derived protein which is called engineered plant prime editor (ePPE) improved prime editing efficiency by ~1.8–3.4-fold in rice and wheat [95]. Zou et al. [96] optimized prime editing by combining PE3 system and epegRNAs, which include a structured RNA motif (evopreQ1 or mpknot) with an 8 bp linker at the 3′ terminus of the pegRNA [96]. Their PPE3-evopreQ1 and PPE3-mpknot systems improved the prime editing efficiencies in rice protoplast, and PPE3-evopreQ1system showed a more significant increase compared to PPE3-mpknot system [96]. This study also increased PE efficiency by at least 2.8 times by applying an appropriate high-temperature treatment. Although each modification enhances PE efficiency, combining these approaches could result in even more significant efficiency improvements [81].
Different prime editing systems in plants, targeted genes, the purpose of the study, and plant delivery technique are summarized in Table 2.2 and also reviewed in [97, 98].
3 Future Prospects and Limitations
3.1 Base Editors
The precise and efficient conversion of single bases at targeted genomic sites is made possible by the CRISPR/Cas base editing technology, which has found wide applications across various plants, as shown in Table 2.1. Although this technology holds great promise for plant trait development, it needs to be improved in order to overcome several limitations, including off-target activity, editing window length, PAM site compatibility, bystander effect, sequence preferences, and the limited capability in editing only four types of base changes [4, 7, 25]. In recent years, substantial efforts have been dedicated to reducing these limitations and enhancing the specificity of base editors in mammalian cells and plants [2, 7].
Comprehensive whole-genome sequencing studies have revealed that base editors can induce gRNA-dependent and gRNA-independent off-target mutations throughout the entire genome [98, 107]. Several effective strategies have been reported to reduce gRNA-dependent off-target effects, such as employing alternative Cas9 variants, enlarging the gRNA sequence, and delivering base editors through RNP (ribonucleoprotein) complexes [98, 108, 109]. Moreover, gRNA-independent off-target mutations were observed in mice and rice using cytosine base editors (CBEs) but not adenine base editors (ABEs). This occurrence is likely attributed to the excessive expression of the deaminase, resulting in random mutations throughout the genome, particularly in gene-enriched regions. Effective strategies to mitigate gRNA-independent off-target effects involve employing alternative deaminases instead of rAPOBEC1 or modifying the deaminase domain through engineering [110].
Jin et al. [111] reported unexpected and unpredictable genome-wide off-target mutations induced by CBEs BE3 and high-fidelity BE3 (HF1-BE3) in rice [111]. The study emphasized the need to optimize the cytidine deaminase domain and/or UGI components to minimize the occurrence of off-target mutations. Additionally, improved variants of CBEs, such as YEE-BE3, were suggested as a potential approach for reducing off-target edits in plants [111]. In another study, upon analyzing off-targets of ABE, considering the predicted top off-target sites with 1- or 2-nt mismatches, it was found that the TadA* (modified version of TadA) deaminases displayed negligible off-target activity (0–4.65% frequency). Furthermore, they suggested that TadA variants exhibit minimal off-target effects dependent on sgRNA [71]. Target selection can be restricted in base editing applications because of the limitations of PAM site compatibility and editing window length [98]. In order to surpass these limitations, various Cas orthologs and engineered variants with altered PAM specificities have been employed to expand the scope of base editors [62, 112]. However, although these variants expand the scope of base editors, they can decrease editing efficiency and enhance the target dependence [98]. In addition to these limitations, large genomes of plants with duplicated regions and genes could pose additional obstacles in selecting target genes and plant transformation steps in base editing [98].
3.2 Prime Editors
While prime editing represents a significant advancement in plant genome editing, the technology is still in its early stages, necessitating further research and studies to unlock its capabilities and potential. One significant challenge with prime editing is its relatively low efficiency [84, 97]. The editing efficiency frequencies observed in plants were considerably lower than those reported in mammalian cells, and numerous sites exhibited a lack of editing, particularly in dicot species [113]. It is also reported that editing efficiencies for insertions were lower than for deletions and substitutions [114]. Although it is possible to obtain targeted mutation in stable transgenic lines by prime editing, as shown in rice and tomato, the occurrence of homozygous and biallelic edits is infrequent, highlighting the inefficiency of prime editing in plants [84, 90, 102, 113]. Researchers have devised various strategies to overcome these limitations, including using engineered prime-editing proteins, enhancements in prime-editing guide RNA design, manipulation of the mismatch repair pathway, and optimization of delivery strategies [92, 97]. These approaches aim to enhance the effectiveness and efficiency of prime editing for more precise and robust genome modifications. Ensuring high efficiency in prime editing relies heavily on the careful design of the pegRNA [78, 93]. Selecting an appropriate combination of the primer binding site (PBS) and reverse transcriptase (RT) template is crucial for optimizing prime editing efficiency. Typically, efficient PBSs range from 8 to 15 nucleotides, while RT templates are between 10 and 20 nucleotides long [78, 93]. Although the specific matrix of optimal PBS and RT template combinations is determined through empirical observations, several factors contribute to selecting the ideal pegRNA design, including GC content, primary sequence motifs, and secondary structures within the pegRNA 3′ extensions [84]. The design of pegRNA is considerably more complex than sgRNA design for other CRISPR-based editing techniques, as it requires adherence to multiple fundamental rules and the various combinations of PBS and RT templates [92, 93]. As a result, its manual design is time-consuming, error-prone, and challenging in high-throughput applications [92]. Several design tools have been developed to overcome this limitation [93, 115, 116].
In conclusion, the development of base editing and prime editing technologies has revolutionized the field of plant genome editing, providing efficient and precise tools for targeted genetic modifications. In addition, the ability to introduce single nucleotide changes without the need for double-stranded DNA breaks has opened new possibilities for crop improvement, disease resistance, and trait engineering. However, despite the significant progress made in this field, challenges still need to be addressed, such as improving editing efficiency and specificity, optimizing delivery methods, and addressing off-target effects. Nevertheless, with continued research and development, base editing, and prime editing hold great promise for advancing the field of plant biotechnology and crop improvement.
References
Hess, G.T., et al.: Methods and applications of CRISPR-mediated base editing in eukaryotic genomes. Mol. Cell. 68(1), 26–43 (2017)
Molla, K.A., Yang, Y.: CRISPR/Cas-mediated base editing: technical considerations and practical applications. Trends Biotechnol. 37(10), 1121–1142 (2019)
Molla, K.A., et al.: Precise plant genome editing using base editors and prime editors. Nat. Plants. 7(9), 1166–1187 (2021)
Rees, H.A., Liu, D.R.: Base editing: precision chemistry on the genome and transcriptome of living cells. Nat. Rev. Genet. 19(12), 770–788 (2018)
Komor, A.C., et al.: Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature. 533, 420–424 (2016)
Jiang, F., et al.: Structures of a CRISPR-Cas9 R-loop complex primed for DNA cleavage. Science. 351(6275), 867–871 (2016)
Anzalone, A.V., Koblan, L.W., Liu, D.R.: Genome editing with CRISPR–Cas nucleases, base editors, transposases and prime editors. Nat. Biotechnol. 38(7), 824–844 (2020)
Koblan, L.W., et al.: Improving cytidine and adenine base editors by expression optimization and ancestral reconstruction. Nat. Biotechnol. 36(9), 843–846 (2018)
Li, J., et al.: Generation of targeted point mutations in rice by a modified CRISPR/Cas9 system. Mol. Plant. 10(3), 526–529 (2017)
Lu, Y., Zhu, J.-K.: Precise editing of a target base in the rice genome using a modified CRISPR/Cas9 system. Mol. Plant. 10(3), 523–525 (2017)
Zong, Y., et al.: Precise base editing in rice, wheat and maize with a Cas9-cytidine deaminase fusion. Nat. Biotechnol. 35(5), 438–440 (2017)
Zhang, R., et al.: Generation of herbicide tolerance traits and a new selectable marker in wheat using base editing. Nat. Plants. 5(5), 480–485 (2019)
Chen, Y., et al.: CRISPR/Cas9-mediated base-editing system efficiently generates gain-of-function mutations in Arabidopsis. Sci. China Life Sci. 60(5), 520–523 (2017)
Xue, C., et al.: Manipulating mRNA splicing by base editing in plants. Sci. China Life Sci. 61, 1293–1300 (2018)
Veillet, F., et al.: Transgene-free genome editing in tomato and potato plants using agrobacterium-mediated delivery of a CRISPR/Cas9 cytidine base editor. Int. J. Mol. Sci. 20(2), 402 (2019)
Veillet, F., et al.: Expanding the CRISPR toolbox in P. patens using SpCas9-NG variant and application for gene and base editing in solanaceae crops. Int. J. Mol. Sci. 21(3), 1024 (2020)
Qin, L., et al.: High-efficient and precise base editing of C•G to T•A in the allotetraploid cotton (Gossypium hirsutum) genome using a modified CRISPR/Cas9 system. Plant Biotechnol. J. 18(1), 45–56 (2020)
Tian, S., et al.: Engineering herbicide-resistant watermelon variety through CRISPR/Cas9-mediated base-editing. Plant Cell Rep. 37(9), 1353–1356 (2018)
Cai, Y., et al.: Target base editing in soybean using a modified CRISPR/Cas9 system. Plant Biotechnol. J. 18(10), 1996 (2020)
Malabarba, J., et al.: New strategies to overcome present CRISPR/Cas9 limitations in apple and pear: efficient dechimerization and base editing. Int. J. Mol. Sci. 22(1), 319 (2020)
Xing, S., et al.: Fine-tuning sugar content in strawberry. Genome Biol. 21(1), 1–14 (2020)
Wu, J., et al.: Engineering herbicide-resistant oilseed rape by CRISPR/Cas9-mediated cytosine base-editing. Plant Biotechnol. J. 18(9), 1857 (2020)
Guyon-Debast, A., et al.: A blueprint for gene function analysis through base editing in the model plant Physcomitrium (Physcomitrella) patens. New Phytol. 230(3), 1258–1272 (2021)
Sretenovic, S., et al.: Exploring C-To-G base editing in rice, tomato, and poplar. Front. Genome Editing. 3, 24 (2021)
Azameti, M.K., Dauda, W.P.: Base editing in plants: applications, challenges, and future prospects. Front. Plant Sci. 12, 664997 (2021)
Jiang, Z., et al.: CRISPR base editing and prime editing: DSB and template-free editing systems for bacteria and plants. Synth. Syst. Biotechnol. 5(4), 277–292 (2020)
Gaudelli, N.M., et al.: Programmable base editing of T to G·C in genomic DNA without DNA cleavage. Nature. 551(7681), 464–471 (2017)
Rees, H.A., et al.: Analysis and minimization of cellular RNA editing by DNA adenine base editors. Sci. Adv. 5(5), eaax5717 (2019)
Kim, H.S., et al.: Adenine base editors catalyze cytosine conversions in human cells. Nat. Biotechnol. 37(10), 1145–1148 (2019)
Hua, K., Tao, X., Zhu, J.K.: Expanding the base editing scope in rice by using Cas9 variants. Plant Biotechnol. J. 17(2), 499–504 (2019)
Li, C., et al.: Expanded base editing in rice and wheat using a Cas9-adenosine deaminase fusion. Genome Biol. 19, 1–9 (2018)
Yan, F., et al.: Highly efficient A·T to G·C base editing by Cas9n-guided tRNA adenosine deaminase in rice. Mol. Plant. 11(4), 631–634 (2018)
Kang, B.-C., et al.: Precision genome engineering through adenine base editing in plants. Nat. Plants. 4(7), 427–431 (2018)
Hua, K., et al.: Simplified adenine base editors improve adenine base editing efficiency in rice. Plant Biotechnol. J. 18(3), 770–778 (2020)
Cortizas, E.M., et al.: UNG protects B cells from AID-induced telomere loss. J. Exp. Med. 213(11), 2459–2472 (2016)
Chen, L., et al.: Programmable C: G to G: C genome editing with CRISPR-Cas9-directed base excision repair proteins. Nat. Commun. 12(1), 1384 (2021)
Kurt, I.C., et al.: CRISPR C-to-G base editors for inducing targeted DNA transversions in human cells. Nat. Biotechnol. 39(1), 41–46 (2021)
Zhao, D., et al.: Glycosylase base editors enable C-to-A and C-to-G base changes. Nat. Biotechnol. 39(1), 35–40 (2021)
Tian, Y., et al.: Efficient C-to-G editing in rice using an optimized base editor. Plant Biotechnol. J. 20(7), 1238 (2022)
Zeng, D., et al.: Exploring C-to-G and A-to-Y Base editing in Rice by using new vector tools. Int. J. Mol. Sci. 23(14), 7990 (2022)
Lee, J., et al.: Application of CRISPR-Based C-to-G Base editing in rice protoplasts. Appl. Biol. Chem. 66(1), 1–5 (2023)
Zhang, X., et al.: Dual base editor catalyzes both cytosine and adenine base conversions in human cells. Nat. Biotechnol. 38(7), 856–860 (2020)
Grünewald, J., et al.: A dual-deaminase CRISPR base editor enables concurrent adenine and cytosine editing. Nat. Biotechnol. 38(7), 861–864 (2020)
Sakata, R.C., et al.: Base editors for simultaneous introduction of C-to-T and A-to-G mutations. Nat. Biotechnol. 38(7), 865–869 (2020)
Xie, J., et al.: ACBE, a new base editor for simultaneous C-to-T and A-to-G substitutions in mammalian systems. BMC Biol. 18(1), 1–14 (2020)
Li, C., et al.: Targeted, random mutagenesis of plant genes with dual cytosine and adenine base editors. Nat. Biotechnol. 38(7), 875–882 (2020)
Bottero, E., et al.: Generation of a multi-herbicide-tolerant alfalfa by using base editing. Plant Cell Rep. 41(2), 493–495 (2022)
Li, Z., et al.: Gene disruption through base editing-induced messenger RNA missplicing in plants. New Phytol. 222(2), 1139–1148 (2019)
Bastet, A., et al.: Mimicking natural polymorphism in eIF 4E by CRISPR-Cas9 base editing is associated with resistance to potyviruses. Plant Biotechnol. J. 17(9), 1736–1750 (2019)
Nakazato, I., et al.: Targeted base editing in the plastid genome of Arabidopsis thaliana. Nat. Plants. 7(7), 906–913 (2021)
Huang, X., Wang, Y., Wang, N.: Base editors for citrus gene editing. Front. Genome Editing. 4, 852867 (2022)
Wang, G., et al.: Development of an efficient and precise adenine base editor (ABE) with expanded target range in allotetraploid cotton (Gossypium hirsutum). BMC Biol. 20(1), 45 (2022)
Luo, J., et al.: Cytosine base editors (CBEs) for inducing targeted DNA base editing in Nicotiana benthamiana. BMC Plant Biol. 23(1), 1–10 (2023)
Zong, Y., et al.: Efficient C-to-T base editing in plants using a fusion of nCas9 and human APOBEC3A. Nat. Biotechnol. 36(10), 950–953 (2018)
Shimatani, Z., et al.: Targeted base editing in rice and tomato using a CRISPR-Cas9 cytidine deaminase fusion. Nat. Biotechnol. 35(5), 441–443 (2017)
Ren, B., et al.: A CRISPR/Cas9 toolkit for efficient targeted base editing to induce genetic variations in rice. Sci. China Life Sci. 60(5), 516–519 (2017)
Ren, B., et al.: Improved base editor for efficiently inducing genetic variations in rice with CRISPR/Cas9-guided hyperactive hAID mutant. Mol. Plant. 11(4), 623–626 (2018)
Hao, L., et al.: CRISPR/Cas9-mediated adenine base editing in rice genome. Rice Sci. 26(2), 125–128 (2019)
Zhong, Z., et al.: Improving plant genome editing with high-fidelity xCas9 and non-canonical PAM-targeting Cas9-NG. Mol. Plant. 12(7), 1027–1036 (2019)
Endo, M., et al.: Genome editing in plants by engineered CRISPR–Cas9 recognizing NG PAM. Nat. Plants. 5(1), 14–17 (2019)
Negishi, K., et al.: An adenine base editor with expanded targeting scope using SpCas9-NGv1 in rice. Plant Biotechnol. J. 17(8), 1476 (2019)
Wang, M., et al.: Targeted base editing in rice with CRISPR/ScCas9 system. Plant Biotechnol. J. 18(8), 1645–1647 (2020)
Liu, X., et al.: A CRISPR-Cas9-mediated domain-specific base-editing screen enables functional assessment of ACCase variants in rice. Plant Biotechnol. J. 18(9), 1845 (2020)
Kuang, Y., et al.: Base-editing-mediated artificial evolution of OsALS1 in planta to develop novel herbicide-tolerant rice germplasms. Mol. Plant. 13(4), 565–572 (2020)
Zhang, C., et al.: Expanding the base editing scope to GA and relaxed NG PAM sites by improved xCas9 system. Plant Biotechnol. J. 18(4), 884 (2020)
Zeng, D., et al.: Engineered Cas9 variant tools expand targeting scope of genome and base editing in rice. Plant Biotechnol. J. 18(6), 1348 (2020)
Zeng, D., et al.: PhieCBEs: plant high-efficiency Cytidine Base editors with expanded target range. Mol. Plant. 13(12), 1666–1669 (2020)
Li, J., et al.: Optimizing plant adenine base editor systems by modifying the transgene selection system. Plant Biotechnol. J. 18(7), 1495 (2020)
Xu, Z., et al.: SpRY greatly expands the genome editing scope in rice with highly flexible PAM recognition. Genome Biol. 22(1), 6 (2021)
Liu, T., et al.: The ScCas9(++) variant expands the CRISPR toolbox for genome editing in plants. J. Integr. Plant Biol. 63(9), 1611–1619 (2021)
Yan, D., et al.: High-efficiency and multiplex adenine base editing in plants using new TadA variants. Mol. Plant. 14(5), 722–731 (2021)
Wei, C., et al.: Efficient generation of homozygous substitutions in rice in one generation utilizing an rABE8e base editor. J. Integr. Plant Biol. 63(9), 1595–1599 (2021)
Li, R., et al.: High-efficiency plastome base editing in rice with TAL cytosine deaminase. Mol. Plant. 14(9), 1412–1414 (2021)
Xu, R., et al.: Development of an efficient plant dual cytosine and adenine editor. J. Integr. Plant Biol. 63(9), 1600–1605 (2021)
Tan, J., et al.: PhieABEs: a PAM-less/free high-efficiency adenine base editor toolbox with wide target scope in plants. Plant Biotechnol. J. 20(5), 934–943 (2022)
Wang, H., et al.: Development of plant cytosine base editors with the Cas12a system. Crop. J. 11(5), 1451–1457 (2023)
Ren, B., et al.: Three novel alleles of OsGS1 developed by base-editing-mediated artificial evolution confer glufosinate tolerance in rice. Crop. J. 11(2), 661–665 (2023)
Anzalone, A.V., et al.: Search-and-replace genome editing without double-strand breaks or donor DNA. Nature. 576(7785), 149–157 (2019)
Doman, J.L., et al.: Designing and executing prime editing experiments in mammalian cells. Nat. Protoc. 17, 2431–2468 (2022)
Huang, Z., Liu, G.: Current advancement in the application of prime editing. Front. Bioeng. Biotechnol. 11, 1039315 (2023)
Nelson, J.W., et al.: Engineered pegRNAs improve prime editing efficiency. Nat. Biotechnol. 40, 1–9 (2021)
Chen, P.J., et al.: Enhanced prime editing systems by manipulating cellular determinants of editing outcomes. Cell. 184(22), 5635–5652 (2021). e29
Chen, P.J., Liu, D.R.: Prime editing for precise and highly versatile genome manipulation. Nat. Rev. Genet. 24(3) 161–177 (2022)
Lin, Q., et al.: Prime genome editing in rice and wheat. Nat. Biotechnol. 38(5), 582–585 (2020)
Lu, Y., et al.: Precise genome modification in tomato using an improved prime editing system. Plant Biotechnol. J. 19(3), 415 (2021)
Perroud, P.F., et al.: Prime editing in the model plant Physcomitrium patens and its potential in the tetraploid potato. Plant Sci. 316, 111162 (2022)
Jiang, Y.Y., et al.: Prime editing efficiently generates W542L and S621I double mutations in two ALS genes in maize. Genome Biol. 21(1), 257 (2020)
Wang, L., et al.: Spelling changes and fluorescent tagging with prime editing vectors for plants. Front. Genome Editing. 3, 617553 (2021)
Butt, H., et al.: Engineering herbicide resistance via prime editing in rice. Plant Biotechnol. J. 18(12), 2370–2372 (2020)
Hua, K., et al.: Precision genome engineering in rice using prime editing system. Plant Biotechnol. J. 18(11), 2167–2169 (2020)
Sretenovic, S., Qi, Y.: Plant prime editing goes prime. Nat. Plants. 8(1), 20–22 (2022)
Hua, K., Han, P., Zhu, J.-K.: Improvement of base editors and prime editors advances precision genome engineering in plants. Plant Physiol. 188(4), 1795–1810 (2022)
Lin, Q., et al.: High-efficiency prime editing with optimized, paired pegRNAs in plants. Nat. Biotechnol. 39(8), 923–927 (2021)
Xu, W., et al.: A design optimized prime editor with expanded scope and capability in plants. Nat. Plants. 8(1), 45–52 (2022)
Zong, Y., et al.: An engineered prime editor with enhanced editing efficiency in plants. Nat. Biotechnol. 40(9), 1394–1402 (2022)
Zou, J., et al.: Improving the efficiency of prime editing with epegRNAs and high-temperature treatment in rice. Sci. China Life Sci. 65(11), 2328–2331 (2022)
Hassan, M.M., et al., Prime editing technology and its prospects for future applications in plant biology research. BioDesign Res. 2020, 1–14 (2020)
Li, Y., et al.: Applications and prospects of CRISPR/Cas9-Mediated Base editing in plant breeding. Curr. Issues Mol. Biol. 45(2), 918–935 (2023)
Biswas, S., et al.: Optimization of prime editing in rice, peanut, chickpea, and cowpea protoplasts by restoration of GFP activity. Int. J. Mol. Sci. 23(17), 9809 (2022)
Xu, R., et al.: Development of plant prime-editing systems for precise genome editing. Plant. Commun. 1(3), 100043 (2020)
Xu, W., et al.: Versatile nucleotides substitution in plant using an improved prime editing system. Mol. Plant. 13(5), 675–678 (2020)
Li, H., et al.: Precise modifications of both exogenous and endogenous genes in rice by prime editing. Mol. Plant. 13(5), 671–674 (2020)
Jiang, Y., et al.: Optimized prime editing efficiently generates glyphosate-resistant rice plants carrying homozygous TAP-IVS mutation in EPSPS. Mol. Plant. 15(11), 1646–1649 (2022)
Li, J., et al.: Development of a highly efficient prime editor 2 system in plants. Genome Biol. 23(1), 161 (2022)
Li, H., et al.: Multiplex precision gene editing by a surrogate prime editor in rice. Mol. Plant. 15(7), 1077–1080 (2022)
Liang, Z., et al.: Addition of the T5 exonuclease increases the prime editing efficiency in plants. J. Genet. Genomics. 50, 582 (2023)
Slaymaker, I.M., et al.: Rationally engineered Cas9 nucleases with improved specificity. Science. 351(6268), 84–88 (2016)
Bharat, S.S., et al.: Base editing in plants: current status and challenges. Crop. J. 8(3), 384–395 (2020)
Rees, H.A., et al.: Improving the DNA specificity and applicability of base editing through protein engineering and protein delivery. Nat. Commun. 8(1), 15790 (2017)
Yu, Y., et al.: Cytosine base editors with minimized unguided DNA and RNA off-target events and high on-target activity. Nat. Commun. 11(1), 2052 (2020)
Jin, S., et al.: Cytosine, but not adenine, base editors induce genome-wide off-target mutations in rice. Science. 364(6437), 292–295 (2019)
Ren, B., et al.: Cas9-NG greatly expands the targeting scope of the genome-editing toolkit by recognizing NG and other atypical PAMs in rice. Mol. Plant. 12(7), 1015–1026 (2019)
Lu, Y., et al.: Precise genome modification in tomato using an improved prime editing system. Plant Biotechnol. J. 19(3), 415–417 (2020)
Wang, L., et al., Spelling changes and fluorescent tagging with prime editing vectors for plants. Front. Genome Editing 3, 617553 (2021)
Siegner, S.M., et al.: PnB designer: a web application to design prime and base editor guide RNAs for animals and plants. BMC Bioinformatics. 22, 1–12 (2021)
Hsu, J.Y., et al.: PrimeDesign software for rapid and simplified design of prime editing guide RNAs. Nat. Commun. 12(1), 1034 (2021)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2024 The Author(s)
About this chapter
Cite this chapter
Kaya, H.B. (2024). Base Editing and Prime Editing. In: Ricroch, A., Eriksson, D., Miladinović, D., Sweet, J., Van Laere, K., Woźniak-Gientka, E. (eds) A Roadmap for Plant Genome Editing . Springer, Cham. https://doi.org/10.1007/978-3-031-46150-7_2
Download citation
DOI: https://doi.org/10.1007/978-3-031-46150-7_2
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-46149-1
Online ISBN: 978-3-031-46150-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)