Abstract
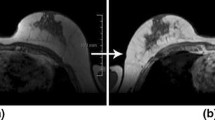
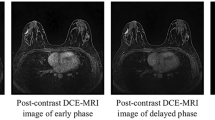
Dynamic contrast-enhanced magnetic resonance imaging (DCE-MRI) and its fast variant, ultrafast DCE-MRI, are useful for the management of breast cancer. Segmentation of breast lesions is necessary for automatic clinical decision support. Despite the advantage of acquisition time, existing segmentation studies on ultrafast DCE-MRI are scarce, and they are mostly fully supervised studies with high annotation costs. Herein, we propose a semi-supervised segmentation approach that can be trained with small amounts of annotations for ultrafast DCE-MRI. A time difference map is proposed to incorporate the distinct time-varying enhancement pattern of the lesion. Furthermore, we present a novel loss function that efficiently distinguishes breast lesions from non-lesions based on triple loss. This loss reduces the potential false positives induced by the time difference map. Our approach is compared to that of five competing methods using the dice similarity coefficient and two boundary-based metrics. Compared to other models, our approach achieves better segmentation results using small amounts of annotations, especially for boundary-based metrics relevant to spatially continuous breast lesions. An ablation study demonstrates the incremental effects of our study. Our code is available on GitHub (https://github.com/yt-oh96/SSL-CTL).
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Sung, H., et al.: Global cancer statistics 2020: Globocan estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clinic. 71, 209–249 (2021)
Maicas, G., Bradley, A.P., Nascimento, J.C., Reid, I., Carneiro, G.: Pre and post-hoc diagnosis and interpretation of malignancy from breast DCE-MRI. Med. Image Anal. 58, 101562 (2019)
Lauby-Secretan, B., et al.: Breast-cancer screening-viewpoint of the IARC working group. N. Engl. J. Med. 372, 2353–2358 (2015)
Morgan, M.B., Mates, J.L.: Applications of artificial intelligence in breast imaging. Radiol. Clinics 59, 139–148 (2021)
Ayatollahi, F., Shokouhi, S.B., Mann, R.M., Teuwen, J.: Automatic breast lesion detection in ultrafast DCE-MRI using deep learning. Med. Phys. 48, 5897–5907 (2021)
Gubern-Mérida, A., et al.: Automated localization of breast cancer in DCE-MRI. Med. Image Anal. 20, 265–274 (2015)
Pisano, E.D., et al.: Diagnostic accuracy of digital versus film mammography: exploratory analysis of selected population subgroups in DMIST. Radiology 246, 376 (2008)
Onishi, N., et al.: Ultrafast dynamic contrast-enhanced breast MRI may generate prognostic imaging markers of breast cancer. Breast Cancer Res. 22, 1–13 (2020)
Mus, R.D., et al.: Time to enhancement derived from ultrafast breast MRI as a novel parameter to discriminate benign from malignant breast lesions. Eur. J. Radiol. 89, 90–96 (2017)
Jing, X., Dorrius, M.D., Wielema, M., Sijens, P.E., Oudkerk, M., van Ooijen, P.: Breast tumor identification in ultrafast MRI using temporal and spatial information. Cancers 14, 2042 (2022)
Kuhl, C.K.: A call for improved breast cancer screening strategies, not only for women with dense breasts. JAMA Netw. Open 4, e2121492 (2021)
Mann, R.M., Kuhl, C.K., Moy, L.: Contrast-enhanced MRI for breast cancer screening. J. Magn. Reson. Imaging 50, 377–390 (2019)
Abe, H., et al.: Kinetic analysis of benign and malignant breast lesions with ultrafast dynamic contrast-enhanced MRI: comparison with standard kinetic assessment. AJR Am. J. Roentgenol. 207, 1159 (2016)
Kim, E.S., et al.: Added value of ultrafast sequence in abbreviated breast MRI surveillance in women with a personal history of breast cancer: a multireader study. Eur. J. Radiol. 151, 110322 (2022)
Onishi, N., et al.: Differentiation between subcentimeter carcinomas and benign lesions using kinetic parameters derived from ultrafast dynamic contrast-enhanced breast MRI. Eur. Radiol. 30, 756–766 (2020)
Honda, M., et al.: New parameters of ultrafast dynamic contrast-enhanced breast MRI using compressed sensing. J. Magn. Reson. Imaging 51, 164–174 (2020)
Goto, M., et al.: Diagnostic performance of initial enhancement analysis using ultra-fast dynamic contrast-enhanced MRI for breast lesions. Eur. Radiol. 29, 1164–1174 (2019)
Gubern-Mérida, A., et al.: Automated localization of breast cancer in DCE-MRI. Med. Image Anal. 20, 265–274 (2015)
Ayatollahi, F., Shokouhi, S.B., Mann, R.M., Teuwen, J.: Automatic breast lesion detection in ultrafast DCE-MRI using deep learning. Med. Phys. 48, 5897–5907 (2021)
Dalmis, M.U., et al.: Artificial intelligence-based classification of breast lesions imaged with a multiparametric breast MRI protocol with ultrafast DCE-MRI, t2, and DWI. Invest. Radiol. 54, 325–332 (2019)
Oh, Y.T., Ko, E., Park, H.: TDM-stargan: Stargan using time difference map to generate dynamic contrast-enhanced MRI from ultrafast dynamic contrast-enhanced MRI. In: 2022 IEEE 19th International Symposium on Biomedical Imaging (ISBI), pp. 1–5. IEEE (2022)
Militello, C., et al.: Semi-automated and interactive segmentation of contrast-enhancing masses on breast DCE-MRI using spatial fuzzy clustering. Biomed. Signal Process. Control 71, 103113 (2022)
Piantadosi, G., Sansone, M., Sansone, C.: Breast segmentation in MRI via U-Net deep convolutional neural networks. In: 2018 24th International Conference on Pattern Recognition (ICPR), pp. 3917–3922. IEEE (2018)
Maicas, G., Carneiro, G., Bradley, A.P., Nascimento, J.C., Reid, I.: Deep reinforcement learning for active breast lesion detection from DCE-MRI. In: Descoteaux, M., Maier-Hein, L., Franz, A., Jannin, P., Collins, D.L., Duchesne, S. (eds.) MICCAI 2017. LNCS, vol. 10435, pp. 665–673. Springer, Cham (2017). https://doi.org/10.1007/978-3-319-66179-7_76
Zhang, J., Saha, A., Zhu, Z., Mazurowski, M.A.: Hierarchical convolutional neural networks for segmentation of breast tumors in MRI with application to radiogenomics. IEEE Trans. Med. Imaging 38, 435–447 (2018)
Van Engelen, J.E., Hoos, H.H.: A survey on semi-supervised learning. Mach. Learn. 109, 373–440 (2020)
Cheplygina, V., de Bruijne, M., Pluim, J.P.: Not-so-supervised: a survey of semi-supervised, multi-instance, and transfer learning in medical image analysis. Med. Image Anal. 54, 280–296 (2019)
Chen, X., Yuan, Y., Zeng, G., Wang, J.: Semi-supervised semantic segmentation with cross pseudo supervision. In: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, pp. 2613–2622 (2021)
Tarvainen, A., Valpola, H.: Mean teachers are better role models: weight-averaged consistency targets improve semi-supervised deep learning results. In: Advances in Neural Information Processing Systems 30 (2017)
Yu, L., Wang, S., Li, X., Fu, C.-W., Heng, P.-A.: Uncertainty-aware self-ensembling model for semi-supervised 3D left atrium segmentation. In: Shen, D., et al. (eds.) MICCAI 2019. LNCS, vol. 11765, pp. 605–613. Springer, Cham (2019). https://doi.org/10.1007/978-3-030-32245-8_67
Berthelot, D., Carlini, N., Goodfellow, I., Papernot, N., Oliver, A., Raffel, C.A.: MixMatch: a holistic approach to semi-supervised learning. In: Advances in Neural Information Processing Systems 32 (2019)
Vu, T.H., Jain, H., Bucher, M., Cord, M., Pérez, P.: Advent: adversarial entropy minimization for domain adaptation in semantic segmentation. In: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, pp. 2517–2526 (2019)
Lee, D.H., et al.: Pseudo-label: The simple and efficient semi-supervised learning method for deep neural networks. In: Workshop on Challenges in Representation Learning, vol. 3, pp. 896. ICML (2013)
Ge, W., Huang, W., Dong, D., Scott, M.R.: Deep metric learning with hierarchical triplet loss. In: Ferrari, V., Hebert, M., Sminchisescu, C., Weiss, Y. (eds.) ECCV 2018. LNCS, vol. 11210, pp. 272–288. Springer, Cham (2018). https://doi.org/10.1007/978-3-030-01231-1_17
Cakir, F., He, K., Xia, X., Kulis, B., Sclaroff, S.: Deep metric learning to rank. In: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) (2019)
Chopra, S., Hadsell, R., LeCun, Y.: Learning a similarity metric discriminatively, with application to face verification. In: 2005 IEEE Computer Society Conference on Computer Vision and Pattern Recognition (CVPR2005), vol. 1, pp. 539–546 (2005)
Do, T.T., Tran, T., Reid, I., Kumar, V., Hoang, T., Carneiro, G.: A theoretically sound upper bound on the triplet loss for improving the efficiency of deep distance metric learning. In: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) (2019)
Park, T., Efros, A.A., Zhang, R., Zhu, J.-Y.: Contrastive learning for unpaired image-to-image translation. In: Vedaldi, A., Bischof, H., Brox, T., Frahm, J.-M. (eds.) ECCV 2020. LNCS, vol. 12354, pp. 319–345. Springer, Cham (2020). https://doi.org/10.1007/978-3-030-58545-7_19
Ronneberger, O., Fischer, P., Brox, T.: U-Net: convolutional networks for biomedical image segmentation. In: Navab, N., Hornegger, J., Wells, W.M., Frangi, A.F. (eds.) MICCAI 2015. LNCS, vol. 9351, pp. 234–241. Springer, Cham (2015). https://doi.org/10.1007/978-3-319-24574-4_28
Wu, Z., Xiong, Y., Yu, S.X., Lin, D.: Unsupervised feature learning via non-parametric instance discrimination. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR) (2018)
Luo, X.: SSL4MIS. https://github.com/HiLab-git/SSL4MIS (2020)
Oh, Y.T., Ko, E., Park, H.: SSL-CTL. https://github.com/yt-oh96/SSL-CTL (2022)
Yeghiazaryan, V., Voiculescu, I.D.: Family of boundary overlap metrics for the evaluation of medical image segmentation. J. Med. Imaging 5, 1–19 (2018)
Acknowledgments
This research was supported by the National Research Foundation (NRF-2020M3E5D2A01084892), Institute for Basic Science (IBSR015- D1), Ministry of Science and ICT (IITP-2020-2018-0-01798), AI Graduate School Support Program (2019-0-00421), ICT Creative Consilience program (IITP-2020-0-01821), and Artificial Intelligence Innovation Hub (2021-0-02068).
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Oh, Yt., Ko, E., Park, H. (2023). Semi-supervised Breast Lesion Segmentation Using Local Cross Triplet Loss for Ultrafast Dynamic Contrast-Enhanced MRI. In: Wang, L., Gall, J., Chin, TJ., Sato, I., Chellappa, R. (eds) Computer Vision – ACCV 2022. ACCV 2022. Lecture Notes in Computer Science, vol 13846. Springer, Cham. https://doi.org/10.1007/978-3-031-26351-4_13
Download citation
DOI: https://doi.org/10.1007/978-3-031-26351-4_13
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-26350-7
Online ISBN: 978-3-031-26351-4
eBook Packages: Computer ScienceComputer Science (R0)