Abstract
Endoglin (CD105) is an auxiliary receptor of transforming growth factor (TGF)-β family members that is expressed in human melanomas. It is heterogeneously expressed by primary and metastatic melanoma cells, and endoglin targeting as a therapeutic strategy for melanoma tumors is currently been explored. However, its involvement in tumor development and malignancy is not fully understood. Here, we find that endoglin expression correlates with malignancy of primary melanomas and cultured melanoma cell lines. Next, we have analyzed the effect of ectopic endoglin expression on two miRNAs (hsa-mir-214 and hsa-mir-370), both involved in melanoma tumor progression and endoglin regulation. We show that compared with control cells, overexpression of endoglin in the WM-164 melanoma cell line induces; (i) a significant increase of hsa-mir-214 levels in small extracellular vesicles (EVs) as well as an increased trend in cells; and (ii) significantly lower levels of hsa-mir-370 in the EVs fractions, whereas no significant differences were found in cells. As hsa-mir-214 and hsa-mir-370 are not just involved in melanoma tumor progression, but they can also target endoglin-expressing endothelial cells in the tumor vasculature, these results suggest a complex and differential regulatory mechanism involving the intracellular and extracellular signaling of hsa-mir-214 and hsa-mir-370 in melanoma development and progression.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
The mechanism involved in tumor development and dissemination of cancer cells is still poorly understood and numerous proteins, miRNAs and signaling pathways have been reported to regulate this process [1, 2]. Among these, endoglin, an auxiliary receptor of the transforming growth factor β (TGF-β) family, has emerged as a promising therapeutic target [3, 4]. Endoglin (Eng; CD105) is a 180-kDa disulfide-linked homodimeric transmembrane glycoprotein [5, 6] highly expressed by proliferating endothelial cells in tumor associated neoangiogenesis [7], as well as in a large number of cancers with poor prognosis [8,9,10,11,12,13]. The role of endoglin in tumor progression and metastasis has been studied in several cancer cell types using in vitro and in vivo models [14,15,16,17,18,19,20,21]. In this regard, an active role of endothelial endoglin in extracellular extravasation of healthy and metastatic tumor cells has been postulated [22, 23]. Furthermore, endoglin-targeted therapy for malignant melanoma is currently been investigated with promising results [24,25,26]. While endoglin is heterogeneously expressed by primary and metastatic melanoma cells, its involvement in the malignant and metastasis processes is not fully understood [8, 27,28,29,30]. Given the high mortality rate of this type of skin cancer and the unresponsiveness of some patients to current immunological treatments, a better knowledge of the mechanisms and active players involved in melanoma growth and development, including endoglin, is a subject of scientific and clinical interest [31, 32].
While endoglin is a type 1 transmembrane glycoprotein with cytoplasmic, transmembrane and extracellular regions, almost 90% of the protein is encompassed within its extracellular region [5]. For this reason, the extracellular region of endoglin has focused many structural and functional studies [6, 33]. Structurally, the extracellular region of endoglin contains two distinct domains: (i) a conserved Zona Pellucida (ZP) juxtamembrane domain at the C-terminus consisting of ~ 260 amino acids (Lys362-Asp561) with eight conserved cysteine residues and divided in two well-defined subdomains (ZP-C and ZP-N); and (ii) a domain at the N-terminus named orphan (orphan domain; OD) due to its lack of significant homology with other protein families [34, 35]. The orphan domain is involved in recognition of TGF-β family ligands [35, 36], whereas the ZP domain is involved in the interaction with members of the integrin family via its arginine-glycine-aspartic acid (RGD) motif located within the ZP-N subdomain [37]. The cellular and pathophysiological function of endoglin has been widely studied in endothelial cells, which are the target in hereditary hemorrhagic telangiectasia type 1 (HHT1), a vascular disease caused by heterozygous mutations in the endoglin gene. HHT1 is associated with telangiectases in skin and mucosa, as well as with arteriovenous malformations in lung, liver, and brain [38, 39]. As an auxiliary receptor of the TGF-β system, endoglin can bind with high affinity to bone morphogenetic protein (BMP)-9 and BMP-10 ligands [36] and interact with the type I and II serine/threonine kinase TGF-β receptors, including ALK1 and ALK5 (type I receptors) and the type II TβRII [40, 41] to modulate cellular responses to different TGF-β family members. Several lines of experimental evidence suggest that binding of BMP9 to endoglin potentiates ALK1 signaling, including the fact that mutations in the gene coding for ALK1 (ACVRL1) are responsible for a second form of HHT (HHT2), whereas heterozygous and homozygous mutations in GDF2, the gene encoding BMP-9, lead to an HHT-like variant [38, 42]. Signaling triggered by BMP-9 through the endoglin/ALK1 route mediates, via the Smad1/5/8 pathway, the expression of a wide range of genes, including the gene for the helix-loop-helix transcription factor inhibitor of differentiation 1 (ID1), a negative transcriptional regulator which is involved in the development of malignant melanoma [43,44,45]. Beyond the TGF-β/BMP-related functions, endoglin is also involved in integrin-mediated cell adhesion via its RGD motif in its extracellular ZP-N subdomain. Thus, endoglin has shown functional binding activity to integrins, such as α5β1 or αIIbβ3 from leukocytes, smooth muscle cells and platelets [22, 37, 46]. Of note, integrins, the major family of cell adhesion receptors in humans, play a key role in tumor growth and metastasis and several studies have investigated the contribution of integrins to the phenotypic aggressiveness of melanoma [47, 48]. In this line, differential expression of integrins in primary cutaneous melanoma has been used to distinguish indolent from aggressive, prometastatic melanoma. Also, some integrins preferentially direct circulating melanoma cells to specific organs, promoting the development of metastases. For example, melanoma cells expressing β1 or β3 integrins, both endoglin interactors, tend to metastasize to the lungs or generate lymph node metastases, respectively. In addition to their relevant role in mediating invasion and metastasis, integrins are not only promising biomarkers, but also attractive therapeutic targets in melanoma [47, 48]. Given the role of integrins in tumor angiogenesis, tumor cell migration and proliferation, and organ-specific metastasis in malignant melanoma, it can be postulated that endoglin, as integrin counter-receptor, will have a relevant impact in melanoma development.
In addition to the membrane-bound form of endoglin, a circulating form of endoglin packed into small extracellular vesicles (EVs) has been described in several pathological conditions, such as preeclampsia, liver disease or thromboembolic pulmonary hypertension [49,50,51,52]. Heterogeneous EVs, including exosomes, can be secreted by all cell types carrying various bioactive cargos such as proteins, RNAs, lipids or metabolites [53]. They are emerging as key regulators of inter-cellular communication in health and disease with potential relevance as biomarkers and therapeutic strategies in different pathological conditions [54, 55]. EVs can transfer their bioactive cargo from donor to recipient cells and influence the biological function of the target cell. In this regard, a functional role for circulating endoglin in EVs has been postulated in several studies, including a protective mechanism supporting endothelial cell survival and angiogenesis [49]. In addition, endoglin+ EVs have been proposed as a biomarker for preeclampsia and metastatic breast cancer [10, 50]. Among the different bioactive cargos of EVs are microRNAs (miRNAs, miRs), which are small endogenous non-coding RNAs that regulate gene expression. During the last decade, compelling evidences support the involvement of cellular and EVs miRNAs in cancer. Among others, miRNAs may act as either tumor suppressors or oncogenes, activating invasion and metastasis, or inducing angiogenesis; as therapeutic targets; and as potential biomarkers for cancer diagnosis, and prognosis [56,57,58,59]. Aberrant expression of miRNAs occurs in several human cancers, including melanoma. Thus, dysregulation of miRNAs has been linked to suppression, progression, differentiation, development, and prognosis of melanoma [60,61,62]. Some miRNAs are specific for one or more skin cancer type, such as hsa-mir-21 and hsa-mir-221, which are observed in cutaneous melanoma and squamous carcinoma; while hsa-mir-155 has been detected in melanoma and cutaneous lymphoma. In this work, we have focused our studies on the pleiotropic hsa-mir-214 and hsa-mir-370, as they are predicted and have been shown to target endoglin [63, 64]. Both, hsa-mir-214 and hsa-mir-370 are dysregulated in several other tumors, besides skin cancers, displaying contrasting behavior. Regardless of whether hsa-mir-214 levels are upregulated or downregulated in skin cancer and melanoma, its dysregulation always correlates with metastasis or poor progression [65, 66]. In the case of hsa-mir-370, controversial findings have also been reported since its upregulation correlates with progression and poor prognosis in breast and prostate cancer [67, 68], as well as promotion of cell apoptosis and inhibition of proliferation in human gastric cancer [69]. By contrast, (i) downregulation of hsa-mir-370 in esophageal squamous-cell carcinoma is associated with cell proliferation and cancer progression [70], and (ii) hsa-mir-370 acts as a tumor suppressor in hepatocellular carcinoma [71]. Interestingly, enforced expression of hsa-mir-370 in melanoma cell lines promotes proliferation, inhibits apoptosis and enhances invasion [72]. Overall, these contradictory results suggest that the function of these miRNAs is highly dependent on the cancer cell context, likely due to their differential cell source, cell target, expression level and/or specific mRNA targeting in each case.
Here we have delved into role of endoglin in human melanoma. We find a correlation between expression levels of endoglin with malignancy in primary melanomas and cultured melanoma cell lines. In addition, overexpression of endoglin in a melanoma cell line leads to dysregulated levels of hsa-mir-214 and hsa-mir-370, mRNAs involved in melanoma tumor progression and endoglin regulation. These results suggest that endoglin is actively involved in development and dissemination of malignant melanoma, and identify endoglin as a potential therapeutic target to block tumor progression.
2 Methods
Immunohistochemistry of melanoma tissues: A total of 73 human specimens (3 benign nevi, 73 malignant melanomas) were analyzed with the corresponding informed consent and ethical protocols approved by the Clinical Investigation Ethical Committee. Immunohistochemistry was performed on 4-μm-thick sections of formalin-fixed, paraffin-embedded tissue samples using an anti-endoglin monoclonal antibody (SN6h, Dako). The staining results were independently analyzed by two expert pathologists who were blinded to the staging and clinical features of the subjects.
Cell culture: WM-164, SK-Mel-28, SK-Mel-103, and SK-Mel-147 cell lines were kindly provided by Dr. María S. Soengas (Spanish National Cancer Research Centre (CNIO), Madrid, Spain). Cells were cultured in DMEM (Lonza BE12-604F) supplemented with 10% heat-inactivated filtered fetal bovine serum (FBS) (Gibco) and 20 µg/mL gentamycin (Lonza 17-519Z). This melanoma cell line was routinely tested for mycoplasma contamination.
Lentiviral production and generation of human ENG stably overexpressing WM-164 cells: Lentiviral plasmids expressing human endoglin containing a hemagglutinin (HA) tag (pLV-CMV-IRES-Puro/hEng) and the corresponding empty vector (pLV-CMV-IRES-Puro/Ø) were kindly provided by Professor Peter ten Dijke (LUMC, Leiden, The Netherlands). These vectors were used in conjunction with the packaging plasmids p8.91 and pSVCG. HEK 293T cells were seeded in a 10-cm plate and transfected with 5 µg p8.91, 2.5 µg pSVCG and 5 µg pLV-CMV-IRES-Puro/Ø or pLV-CMV-IRES-Puro/hENG, using Lipofectamine® 2000 (Thermo Fisher Scientific), according to the manufacturer’s instructions. After 10–12 h, medium was changed by fresh culture medium (DMEM) and cells were incubated for additional 48 h. Culture supernatants containing lentiviral particles were harvested, clarified by centrifugation at 1500 rpm for 5 min, and filtered through a 0.45 µm filter. Lentiviral particles at 1:3 dilution were used to infect WM-164 cells in suspension in the presence of 4 µg/mL polybrene (Sigma). After incubation for 24 h, medium was replaced by fresh culture medium. Twenty four hours later, infected cells were selected in the presence of 0.4 µg/mL puromycin (Sigma), and the resulting endoglin-overexpressing WM-164 cells (WM-164 ENG) were validated by immunoblot and flow cytometry analyses.
Immunoblot assays: Cells were washed twice with PBS and lysed in cold lysis solution containing 50 mM HEPES pH 7.5, 0.4 M KCl, 10% glycerol, 1% NP-40 and protease inhibitors (PhosSTOP™, Sigma Aldrich). Lysates were sonicated for 1 min and centrifuged at 13,000 rpm for 10 min at 4 °C. Supernatant fractions were used for Western blot analyses. Protein extracts or purified EVs were quantified for protein content using the bicinchoninic acid assay (Pierce™ BCA Protein Assay kit, Thermo Scientific). Equal amounts of extracted protein or purified EVs from each sample were resuspended in Laemmli buffer and, subsequently, incubated at 95 °C for 10 min. Samples were separated by SDS-PAGE and then transferred onto a PVDF membrane (Invitrogen). Protein-bound membranes were blocked with 0.1% Tween-20 (Sigma-Aldrich) in Tris-buffered saline (TBS) containing 5% BSA or 2.5–3% milk (TBS-T), and phosphatase inhibitor cocktail (0.2 mM sodium orthovanadate, 5 mM sodium beta-glycerophosphate and 10 mM sodium fluoride) for 2 h at room temperature. Membranes were then incubated overnight at 4 °C with the following primary antibodies specific for: human endoglin (1:1000 in TBS-T/BSA, Abcam #169545); ALIX (1:1000 in TBS-T/milk, Cell Signal #2171); MEK 1/2 (1:1000 in milk; Cell Signaling #8727S); AKT (1:1000 in milk, Cell Signaling #9272); β-actin (1 μg/mL in TBS-T/BSA, Sigma #A1978); and GAPDH (1:500 in TBS-T/BSA, Abcam #9484). Then, membranes were washed with TBT-T and incubated for 1 h at room temperature with the corresponding secondary HRP-linked antibodies. After rinsing with TBS-T, protein bands were revealed using SuperSignal™ West Pico PLUS Chemiluminescent substrate (Thermo Scientific) to enhance HRP luminescence, followed by analysis using the Molecular Imager® Gel Doc™ XR+ System with Image Lab™ software (Bio-Rad).
Immunofluorescence flow cytometry: Cell surface expression of endoglin in WM164 cells was analyzed by flow cytometry. After collecting and washing transfected cells in PBS by soft centrifugation at 1000 rpm 8 °C for 5 min, non-specific binding was blocked for 20 min at 4 °C with sterile-filtered 1% BSA in PBS (PBS-BSA). Cells were then incubated for 1 h at 4 °C with a mouse monoclonal antibody against human endoglin (P4A4, anti-CD105, 1/100; Developmental Studies Hybridoma Bank-DSHB-University of Iowa, USA) or against the hemagglutinin (HA) tag (1/100; MilliporeSigma). As a negative control, cells were stained with isotype control antibodies (Immunostep, Salamanca, Spain) at the same concentration as the corresponding primary antibody. Following incubation with primary antibodies, cells were washed with PBS, and incubated with Alexa-Fluor-488-conjugated anti-mouse antibody (1/200, Molecular Probes) for an additional period of 45 min. Samples were then washed, resuspended in cold PBS, and analyzed with a FC500 Beckman Coulter flow cytometer using the FlowLogic software. Endoglin protein levels were measured using the fluorescence intensity mean and expressed as fold induction relative to empty-transfected cells.
EVs isolation by sequential ultracentrifugation, characterization and analyses: Cells were cultured in media supplemented with 10% EVs-depleted FBS. Serum was depleted of bovine EVs by ultracentrifugation at 100,000 g for 70 min at 10 °C and then filtered. Supernatant fractions collected from 48 to 72 h exponentially growing cell cultures were pelleted by centrifugation at 500 g for 10 min at 4 °C to remove any cell contamination. In addition, possible apoptotic bodies and large cell debris were removed from supernatants by centrifugation at 12,000 g for 20 min at 10 °C. EVs, including exosomes were then collected by spinning at 100,000 g for 70 min at 10 °C. The pellet with EVs was then washed in 20 mL of PBS and collected again by ultracentrifugation at 100,000 for 70 min at 10 °C (Beckman, L100 X-P). The final pellet of EVs was resuspended in PBS. EVs size and particle number were analyzed using Nanosight (Nanoparticle Tracking Analysis-NTA) and its protein content was measured by BCA.
RNA isolation, cDNA synthesis and quantitative RT-PCR: microRNA (miRNA) and total RNA were isolated from cells using the miRNeasy Micro kit (Qiagen), according to the manufacturer´s instructions. To quantify specific microRNAs, first they were reversed transcribed using TaqMan™ MicroRNA Reverse Transcription kit and then, PCR was performed using Taqman Universal PCR Master mix (Applied Biosystems) and specific and pre-designed Taqman® MicroRNA assays (hsa-miR-214: ID 002306 and hsa-miR-370: ID 002275). For quantification of gene expression, RT-PCR was performed with SuperScript™ II (Invitrogen) and FastStart Essential DNA Green Master (Roche) using the primers shown in Table 14.1. qRT-PCR was performed on Light Cycler 96 (Roche), according to the following PCR settings: initial denaturation for 10 min at 95 °C, 40 cycles of 15 s at 95 °C and 60 s at 60 °C for miRNA assays and 30 s at 60 °C in the case of gene expression assays. Both miRNA and total RNA quantifications were performed in triplicates. Gene and miRNA expressions were analyzed using the delta-deltaCT method for relative quantification and all samples were normalized to the corresponding housekeeping gene, hsa-miR-16 ID 000391 and human mRNA β-actin.
3 Results
Endoglin expression in primary melanomas and cultured cells
Endoglin expression was assessed in a cohort of primary melanomas and dermal nevi by immunohistochemistry. As expected, staining of endoglin was observed in endothelial cells from primary melanomas and dermal nevi. While endoglin staining was not detected in dermal nevi, 41.4 and 18.6% of primary melanomas showed low or high endoglin expression, respectively, in tumor cells (Fig. 14.1). These data suggest that, compared to normal nevi, melanoma tumors show markedly increased levels of endoglin. This prompted us to study whether endoglin expression correlates with melanoma malignancy or metastatic potential. Endoglin expression levels were also analyzed by immunoblotting in a panel of different melanoma cell lines (Fig. 14.2). While the non-metastatic or low metastatic melanoma cell lines (WM-164 and SK-Mel-28, respectively) showed low levels of endoglin, the more metastatic cell lines (SK-Mel-147 and SK-Mel-103) had higher levels of endoglin expression (Fig. 14.2). These results suggest that endoglin expression correlates with malignancy in primary melanomas as well as in cultured melanoma cell lines.
Characterization of ectopically overexpressed endoglin in the WM-164 cell line
To investigate the impact of endoglin in the malignant phenotype of melanoma cells, the low metastatic melanoma cell line WM-164 was transduced with a lentivirus encoding HA-tagged human endoglin. Following cell infection and puromycin selection, we verified the ectopic expression of endoglin by immunoblot analysis in cellular extracts and EVs fractions (Fig. 14.3a). As expected, endoglin-transduced cells and derived EVs showed a clear signal of ectopic endoglin relative to mock-transduced cells. To confirm the correct isolation of the EVs, the expression of ALIX, a broad biomarker of EVs, was tested. ALIX was not detected in cellular extracts, whereas a weak band was observed in EVs from mock- and endoglin-transduced cells (Fig. 14.3a), confirming the proper quality of purified EVs. To assess whether endoglin overexpression could be affecting other relevant signaling pathways, we analyzed total protein levels of MEK and AKT by immunoblotting. No significant differences were found in MEK and AKT protein levels between mock- and endoglin-transduced cells (Fig. 14.3b). The expression of ectopic endoglin in transduced WM-164 cells was also analyzed by flow cytometry using anti-HA or anti-endoglin (P4A4) monoclonal antibodies (Fig. 14.4). The strong expression of cell surface endoglin was demonstrated in endoglin-transduced WM-164 cells compared to the weak labelling of mock-transduced cells, as evidenced by the histograms obtained with anti-HA (Fig. 14.4a, left panel) and P4A4 anti-endoglin (Fig. 14.4a, right panel and Fig. 14.4b) monoclonal antibodies. Taken together, immunoblot (Fig. 14.3) and flow cytometry (Fig. 14.4) analyses demonstrate that lentiviral transduction of WM-164 cells efficiently yields endoglin overexpression at their cell surface. The results from Figs. 14.1 and 14.2 suggesting that endoglin expression correlates with malignancy of melanomas prompted us to analyze the malignant phenotype of the endoglin-expressing WM-164 cells by measuring the levels of PMEL and TYR2, two well-known melanoma markers. PMEL (Premelanosome protein) is expressed by melanocytes and melanoma cells, and is widely used as a melanoma marker in serum samples. Compared with normal melanocytes, PMEL is over-expressed at all stages of melanoma progression [73, 74]. Tyrosinase TYR2 is involved in melanogenesis and mediates anti-apoptotic effects in human melanoma cells [75]. Analysis by qRT-PCR of endoglin-transduced WM-164 cells and their derived EVs showed that PMEL and TYR2 mRNA levels were significantly higher than those of mock-transduced WM-164 controls (Fig. 14.5). These results further support the involvement of endoglin in melanoma progression.
Western blot analysis of endoglin-expressing WM-164 cells. a Analysis of cellular lysates and EVs from mock-transduced (ø) or endoglin-transduced (hENG+) WM-164 cells using antibodies to endoglin (anti-HA), the EVs marker ALIX or GAPDH, as a loading control. b Analysis of cellular lysates from mock-transduced (ø) or endoglin-transduced (hENG+) WM-164 cells using antibodies to endoglin (anti-HA), total MEK, total AKT or β-actin, as a loading control. Representative Western blots are shown
Flow cytometry analysis of endoglin-expressing WM-164 cells. a Cell surface expression of endoglin was analyzed in mock-transduced (ø) or endoglin-transduced (hENG+) WM-164 cells using anti-HA or P4A4 antibodies that recognize the recombinant endoglin. Representative flow cytometry histograms are shown. b Endoglin protein levels were measured in cells stained with P4A4 anti-endoglin antibody (n = 4 per condition) as in panel (a). **p < 0.01; by two-tailed student’s t-test
Analysis of the melanoma marker genes PMEL and TYR2 in endoglin-expressing WM-164 cells. Cells transduced with endoglin (hENG+) or an empty vector (ø) were analyzed by qRT-PCR using primers specific for PMEL and TYR2. Representative qRT-PCR from cells (upper panels) or their derived EVs (lower panels) are shown (n = 3 per condition). *p < 0.05; **p < 0.01; ***p < 0.001; by two-tailed student’s t-test
Effect of endoglin expression on additional markers of melanoma cancer cells
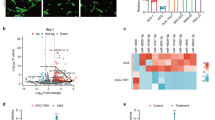
To further assess whether endoglin upregulation exerted a functional effect during melanoma development and progression, three additional markers were analyzed: (i) MLANA (also known as MART-1, Melanoma antigen recognized by T-cells 1); (ii) MITF (Microphthalmia-associated transcription factor); and (iii) VEZF-1 (Vascular endothelial zinc finger 1). Of note, MLANA and MITF are relevant proteins involved in melanocyte and melanoma biology. MLANA is a cytoplasmic protein expressed by normal melanocytes and benign nevi and it is used in the clinic to detect and confirm melanocytic tumors [76, 77]. In addition, MITF has been described as the main transcription factor regulating key processes in melanoma cell development, growth, survival, proliferation, differentiation and invasion [78, 79]. Also, VEZF-1 is a Krüppel-like zinc finger protein that contributes to cancer pathogenesis [80, 81]. qRT-PCR analysis showed that MLANA, MITF and VEZF-1 mRNA levels were significantly increased in endoglin-expressing WM-164 cells, but not in EVs, both compared to mock-transduced cells (Fig. 14.6). These findings further support the active role of endoglin in melanoma development and progression.
Analysis of MLANA, MITF and VEZF-1 genes in WM-164 cells transduced with empty (ø) or endoglin-expressing (hENG+) vectors. qRT-PCR of ENG, MLANA, MITF and VEZF-1 genes was carried out using specific primers in cells (a) or their derived EVs (b) (n = 3 per condition). *p < 0.05; **p < 0.01 and ***p < 0.001; by two-tailed student’s t-test
Effect of ectopic endoglin expression in the levels of hsa-miR-214 and hsa-miR-370
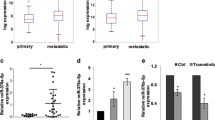
Emerging evidence support the involvement of cellular and EVs miRNAs in cancer progression, diagnosis, and prognosis [56,57,58,59], including melanoma [60,61,62]. Therefore, we investigated the effect of endoglin in the levels of miR-214 and miR-370 as they (i) have been found dysregulated in several cancer types, including skin cancers, (ii) are predicted to target endoglin [64], and (iii) differential expression of circulating miR-370 has been reported in plasma from patients with hereditary hemorrhagic telangiectasia type 1 (HHT1), an autosomal dominant disorder due to mutations in the endoglin gene [64]. We then measured by qRT-PCR hsa-miR-214 and hsa-miR-370 levels in cells and EVs from endoglin-expressing WM-164 cells (Fig. 14.7). Levels of hsa-miR-370 were similar in control and endoglin-expressing WM-164 cells, whereas the expression of hsa-miR-214 showed a non-significant increased trend in endoglin-positive WM-164 cells compared to controls (Fig. 14.7, left panels). In EVs from endoglin-transduced WM-164 cells, the levels of hsa-miR-370 displayed a significant reduction, while those of hsa-miR-214 showed a significant increase compared to mock-transduced WM-164 cells (Fig. 14.7, right panels). These results suggest that endoglin expression in melanoma involves the dysregulation of hsa-miR-214 and hsa-miR-370, which in turn could modulate melanoma progression.
Analysis of miRNAs hsa-mir-370 and hsa-mir-214 in endoglin-expressing WM-164 cells. Cells transduced with endoglin (hENG+) or an empty vector (ø) were analyzed by qRT-PCR using primers specific for hsa-mir-370 and hsa-mir-214. Representative qRT-PCR from cells (left panels) or their derived EVs (right panels) are shown (n = 4 per condition). The miRNA expression levels are displayed relative to WM-164 cells transduced with the empty vector. *p < 0.05; **p < 0.01 by two-tailed student’s t-test
4 Discussion
In this work, we demonstrate a correlation between endoglin expression and tumor malignancy in primary melanoma and cultured melanoma cell lines. We have also deepened into the underlying endoglin-dependent molecular mechanisms, mainly focusing on the role of microRNAs in this process. Besides its physiological role in angiogenesis, endoglin has also emerged as a promising therapeutic target in recent years since endoglin expression has been reported either in tumor vessels or neoplasm in tumor cells, including melanoma, renal cell carcinoma (RCC), leukemias, certain subtypes of sarcomas, and breast, ovarian, endometrial, and prostate cancer. The role of endoglin in tumor cells depends on the cellular context. In this regard, and in line with our results obtained in melanoma, endoglin would be promoting tumor development and progression, playing an important role in oncogenic signalling (Fig. 14.8); whereas in other cases it has been associated with tumor suppression [4, 15, 82, 83]. In melanoma, endoglin has been pointed out to be essential for tumor plasticity, playing a key role in the interplay between TGF-β and BMP signalling pathways. Accordingly, endoglin downregulation hinders anchorage-independent growth and invasiveness and abrogates tumor growth in preclinical models of melanoma [8]. Moreover, experiments with shRNA against endoglin have shown to significantly reduce proliferation, survival and migration of melanoma cells [26, 30]. Recently, the therapeutic efficacy of a fusion protein containing endoglin single-chain variable fragment and IP10 (Endoglin-scFv/IP10) has been demonstrated. Indeed, this fusion protein inhibited proliferation and angiogenesis, while stimulating apoptosis within melanoma tissue [25]. In this context, our results further support the hypothesis that endoglin mediates malignant melanocyte transformation in WM-164, as the levels of the well-known melanoma markers PMEL and TYR2 increase upon endoglin overexpression. Furthermore, an increased trend of hsa-mir-214 levels is observed in endoglin-transduced melanoma cells. Interestingly, hsa-mir-214 dysregulation has been widely described in several tumors, including melanoma.
Hypothetical model of the role of endoglin in melanoma progression. Endoglin expression is upregulated in malignant melanoma compared to primary melanocytes, contributing to the malignant phenotype, at least by dysregulating hsa-mir-214 and hsa-mir-370. EVs released by malignant melanocytes and loaded with abnormal levels of hsa-mir-214 and hsa-mir-370 can target cells from the tumor microenvironment, including primary melanocytes, endothelial cells from the tumor vasculature, melanoma-associated fibroblasts (MAFs), lymphocytes or tumor-associated macrophages (TAMs), leading to enhanced melanoma tumor growth and development. Created with BioRender.com
Cancer-derived extracellular vesicles, including EVs, can target different cell types in the tumor microenvironment modulating tumor growth and metastasis [84,85,86]. Of note, cellular endoglin expression significantly regulates both hsa-mir-214 and hsa-mir-370 in EVs, of which endoglin is also a component. Thus, compared to EVs from control cells, endoglin overexpressing cells show reduced levels of hsa-mir-370 while increased content of hsa-mir-214 in EVs. We hypothesize that these dysregulated microRNAs in EVs may play a relevant role in tumor development and metastasis (Fig. 14.8). For example, the reduction of hsa-mir-370 levels in EVs from endoglin-expressing melanoma cells could favour the process of neo-angiogenesis, which is necessary for tumor growth. This can be achieved because endoglin is negatively regulated by hsa-mir-370 [63], and endoglin is highly expressed by actively proliferating endothelial cells of the tumor vasculature [7]. Consequently, EVs from the primary melanoma tumors carrying lower levels of hsa-mir-370 would favour migration, proliferation, differentiation and adhesion of endothelial cells. Given the reported role of hsa-mir-214 in tumor progression [65, 66], increased levels of hsa-mir-214 in EVs from melanoma cells may act in a paracrine manner once taken up by neighbour melanocyte cells, thereby transforming them and contributing to tumor growth and development (Fig. 14.8). The EVs-mediated targeting of hsa-mir-370 and hsa-mir-214 may not be limited to neoangiogenic vessels or melanocytes, as an effect on additional non-cancer cells from the tumor environment is expected as well [87]. Apart from malignant cells, non-cancerous cells, including adipocytes, endothelial cells of tumor vessels, lymphocytes, tumor-associated macrophages (TAMs), and cancer-associated fibroblasts (CAFs), as well as molecules produced and released by them, constitute the tumor microenvironment [88, 89]. Active and mutual interactions, through a paracrine signalling or circulatory and lymphatic systems, between tumor cells and the tumor microenvironment have been described to play decisive roles in tumor initiation, development and progression, metastasis, and response to therapies [90, 91]. Consequently, the tumor environment has received increased attention in the recent cancer literature [92, 93]. For instance, melanoma-associated fibroblasts (MAFs) have been described to have a role in melanoma progression, therapy resistance and immunosurveillance [94,95,96]. Moreover, a variety of immune cells, i.e., T and B lymphocytes, macrophages, neutrophils, dendritic and natural killer cells support the growth and invasiveness of melanoma cells, using multiple mechanisms. Among them, it is remarkable the downregulation in T lymphocytes of anti-apoptotic proteins, including Bcl-2, caused by melanoma-derived EVs containing miRNAs, such as hsa-mir-690 [97, 98]. A recent study has shown that hsa-mir-125b-5p transferred by cutaneous melanoma-derived EVs induces a tumor-promoting TAM phenotype in macrophages [99]. A role for EVs carrying hsa-mir-370 or hsa-mir-214 on malignant progression has been outlined. Breast cancer cells-secreted EVs with hsa-mir-370-3p cargo can aggravate breast cancer through downregulation of the cylindromatosis (CYLD) tumor suppressor in fibroblasts concomitantly with activation of the NF-κB signaling pathway, thereby promoting the tumor cell functions [100]. Interestingly, expression of endoglin, a target of hsa-mir-370, in CAFs regulates invasion and stimulates colorectal cancer metastasis [101]. Also, by sponging hsa-mir-370-3p, the circular RNA (circRNA) circ_0020710 can promote melanoma cell proliferation, migration and invasion in vitro, as well as tumor growth in vivo through the upregulated expression of the CXCL12 [102], a chemokine known to regulate melanoma metastasis to distant sites [103]. In the case of hsa-mir-214, its downregulation in CAFs contributes to migration and invasion of gastric cancer cells through induction of epithelial-mesenchymal transition (EMT) [104]. Accordingly, hsa-mir-214-3p has been proposed as a novel therapeutic target in pancreatic CAFs and human pancreatic stellate cells (hPSCs), as its inhibition led to inhibition of TGF-β-induced differentiation of pancreatic CAFs and reduced expression of myofibroblast markers during the differentiation of hPSCs to myofibroblasts [105]. Furthermore, a role of tumor-secreted miR-214 in the conversion of CD4+ T cells into immune-suppressive regulatory T cells, promoting tumor immune escape has been described [106]. Future independent studies remain to be performed to better understand the functional impact of the endoglin-induced dysregulated microRNAs in melanoma cells and their microenvironment, as well as the possible mechanisms involved.
Along with the hsa-mir-214 and hsa-mir-370 cargos, EVs derived from endoglin-enriched melanoma cells, also contain the protein endoglin, in agreement with previous reports in EVs from endoglin-expressing endothelial cells or primary hepatic stellate cells [49,50,51]. Although endoglin+ EVs have been proposed as biomarkers for metastatic breast cancer [10], the putative functional role of this endoglin cargo in cancer remains to be elucidated. It is well established that endoglin specifically binds integrins [22, 37, 46] and tumor cell-derived EVs contain integrins involved the generation of pre-metastatic niches in specific tissues promoting organ-specific metastases of several types of cancer including melanoma [47, 84, 107]. Accordingly, it is tempting to speculate that by interacting with integrins, endoglin could be involved in these malignant processes. Further investigations are needed to better understand the role of endoglin in melanoma development.
Abbreviations
- AKT:
-
Serine/threonine-specific protein kinase B (PKB)
- ALIX:
-
ALG-2-interacting protein X
- BMP:
-
Bone morphogenetic protein
- BSA:
-
Bovine serum albumin
- CAFs:
-
Cancer-associated fibroblasts
- circRNA:
-
Circular RNA
- CMV:
-
Citomegalovirus
- CYLD:
-
Cylindromatosis
- DMEM:
-
Dulbecco’s modified Eagle medium
- EMT:
-
Epithelial-mesenchymal transition
- EVs:
-
Small extracellular vesicles
- FBS:
-
Fetal bovine serum
- HA:
-
Hemagglutinin
- HEPES:
-
4-[2-Hydroxyethyl]-1-piperazineethanesulfonic acid
- HHT:
-
Hereditary hemorrhagic telangiectasia
- hPSCs:
-
Human pancreatic stellate cells
- HRP:
-
Horseradish peroxidase
- hsa-mir:
-
Homo sapiens microRNA
- MAFs:
-
Melanoma-associated fibroblasts
- MEK:
-
Mitogen-activated ERK kinase
- mir:
-
MicroRNA
- miRNA:
-
MicroRNA
- NP-40:
-
Nonidet P-40
- PAGE:
-
Poly-acrylamide gel electrophoresis
- PBS:
-
Phosphate-buffered saline
- PMEL:
-
Premelanosome protein
- qRT-PCR:
-
Real-time quantitative reverse transcription PCR
- SDS:
-
Sodium dodecyl sulfate
- TAMs:
-
Tumor-associated macrophages
- TBS:
-
Tris-buffered saline
- TGF-β:
-
Transforming growth factor beta
- TYR2:
-
Tyrosinase 2
References
Motwani J, Eccles MR (2021) Genetic and genomic pathways of melanoma development, invasion and metastasis. Genes (Basel) 12(10):1543. https://doi.org/10.3390/genes12101543
Cherepakhin OS, Argenyi ZB, Moshiri AS (2021) Genomic and transcriptomic underpinnings of melanoma genesis, progression, and metastasis. Cancers (Basel) 14(1):123. https://doi.org/10.3390/cancers14010123
Liu Y, Paauwe M, Nixon AB, Hawinkels LJAC (2020) Endoglin targeting: lessons learned and questions that remain. Int J Mol Sci 22(1):147. https://doi.org/10.3390/ijms22010147
González Muñoz T, Amaral AT, Puerto-Camacho P, Peinado H, de Álava E (2021) Endoglin in the spotlight to treat cancer. Int J Mol Sci 22(6):3186. https://doi.org/10.3390/ijms22063186
Gougos A, Letarte M (1990) Primary structure of endoglin, an RGD-containing glycoprotein of human endothelial cells. J Biol Chem 265:8361–8364
López-Novoa JM, Bernabeu C (2010) The physiological role of endoglin in the cardiovascular system. Am J Physiol Heart Circ Physiol 299(4):H959–H974. https://doi.org/10.1152/ajpheart.01251.2009
Bernabeu C, Lopez-Novoa JM, Quintanilla M (2009) The emerging role of TGF-beta superfamily coreceptors in cancer. Biochim Biophys Acta 1792(10):954–973. https://doi.org/10.1016/j.bbadis.2009.07.003
Pardali E, van der Schaft DW, Wiercinska E, Gorter A, Hogendoorn PC, Griffioen AW, ten Dijke P (2011) Critical role of endoglin in tumor cell plasticity of Ewing sarcoma and melanoma. Oncogene 30(3):334–345. https://doi.org/10.1038/onc.2010.418
Liu P, Sun YL, Du J, Hou XS, Meng H (2012) CD105/Ki67 coexpression correlates with tumor progression and poor prognosis in epithelial ovarian cancer. Int J Gynecol Cancer 22:586–592
Douglas SR, Yeung KT, Yang J, Blair SL, Cohen O, Eliceiri BP (2021) Identification of CD105+ extracellular vesicles as a candidate biomarker for metastatic breast cancer. J Surg Res 268:168–173. https://doi.org/10.1016/j.jss.2021.06.050
Laukhtina E, Schuettfort VM, D’Andrea D, Pradere B, Mori K, Quhal F, Sari Motlagh R, Mostafaei H, Katayama S, Grossmann NC, Rajwa P, Zeinler F, Abufaraj M, Moschini M, Zimmermann K, Karakiewicz PI, Fajkovic H, Scherr D, Compérat E, Nyirady P, Rink M, Enikeev D, Shariat SF (2022) Preoperative plasma level of endoglin as a predictor for disease outcomes after radical cystectomy for nonmetastatic urothelial carcinoma of the bladder. Mol Carcinog 61(1):5–18. https://doi.org/10.1002/mc.23355
Greiner SM, Märklin M, Holzmayer S, Kaban K, Meyer S, Hinterleitner C, Tandler C, Hagelstein I, Jung G, Salih HR, Heitmann JS, Kauer J (2022) Identification of CD105 (endoglin) as novel risk marker in CLL. Ann Hematol. https://doi.org/10.1007/s00277-022-04756-4
Puerto-Camacho P, Díaz-Martín J, Olmedo-Pelayo J, Bolado-Carrancio A, Salguero-Aranda C, Jordán-Pérez C, Esteban-Medina M, Álamo-Álvarez I, Delgado-Bellido D, Lobo-Selma L, Dopazo J, Sastre A, Alonso J, Grünewald TGP, Bernabeu C, Byron A, Brunton VG, Amaral AT, de Álava E (2022) Endoglin and MMP14 contribute to Ewing sarcoma spreading by modulation of cell-matrix interactions. Int J Mol Sci 23(15):8657. https://doi.org/10.3390/ijms23158657
Quintanilla M, Ramirez JR, Pérez-Gómez E, Romero D, Velasco B, Letarte M, López-Novoa JM, Bernabéu C (2003) Expression of the TGF-beta coreceptor endoglin in epidermal keratinocytes and its dual role in multistage mouse skin carcinogenesis. Oncogene 22(38):5976–5985. https://doi.org/10.1038/sj.onc.1206841
Pérez-Gómez E, Villa-Morales M, Santos J, Fernández-Piqueras J, Gamallo C, Dotor J, Bernabéu C, Quintanilla M (2007) A role for endoglin as a suppressor of malignancy during mouse skin carcinogenesis. Cancer Res 67(21):10268–10277. https://doi.org/10.1158/0008-5472.CAN-07-1348
Oxmann D, Held-Feindt J, Stark AM, Hattermann K, Yoneda T, Mentlein R (2008) Endoglin expression in metastatic breast cancer cells enhances their invasive phenotype. Oncogene 27(25):3567–3575. https://doi.org/10.1038/sj.onc.1211025
Romero D, Terzic A, Conley BA, Craft CS, Jovanovic B, Bergan RC, Vary CP (2010) Endoglin phosphorylation by ALK2 contributes to the regulation of prostate cancer cell migration. Carcinogenesis 31(3):359–366. https://doi.org/10.1093/carcin/bgp217
Santibanez JF, Pérez-Gómez E, Fernandez-L A, Garrido-Martin EM, Carnero A, Malumbres M, Vary CP, Quintanilla M, Bernabéu C (2010) The TGF-beta co-receptor endoglin modulates the expression and transforming potential of H-Ras. Carcinogenesis 31(12):2145–2154. https://doi.org/10.1093/carcin/bgq199
Lakshman M, Huang X, Ananthanarayanan V, Jovanovic B, Liu Y, Craft CS, Romero D, Vary CP, Bergan RC (2011) Endoglin suppresses human prostate cancer metastasis. Clin Exp Metastasis 28(1):39–53. https://doi.org/10.1007/s10585-010-9356-6
Henry LA, Johnson DA, Sarrió D, Lee S, Quinlan PR, Crook T, Thompson AM, Reis-Filho JS, Isacke CM (2011) Endoglin expression in breast tumor cells suppresses invasion and metastasis and correlates with improved clinical outcome. Oncogene 30(9):1046–1058. https://doi.org/10.1038/onc.2010.488
Romero D, O’Neill C, Terzic A, Contois L, Young K, Conley BA, Bergan RC, Brooks PC, Vary CP (2011) Endoglin regulates cancer-stromal cell interactions in prostate tumors. Cancer Res 71(10):3482–3493. https://doi.org/10.1158/0008-5472.CAN-10-2665
Rossi E, Sanz-Rodriguez F, Eleno N, Düwell A, Blanco FJ, Langa C, Botella LM, Cabañas C, Lopez-Novoa JM, Bernabeu C (2013) Endothelial endoglin is involved in inflammation: role in leukocyte adhesion and transmigration. Blood 121(2):403–415. https://doi.org/10.1182/blood-2012-06-435347
Anderberg C, Cunha SI, Zhai Z, Cortez E, Pardali E, Johnson JR, Franco M, Páez-Ribes M, Cordiner R, Fuxe J, Johansson BR, Goumans MJ, Casanovas O, ten Dijke P, Arthur HM, Pietras K (2013) Deficiency for endoglin in tumor vasculature weakens the endothelial barrier to metastatic dissemination. J Exp Med 210(3):563–579. https://doi.org/10.1084/jem.20120662
Tesic N, Kamensek U, Sersa G, Kranjc S, Stimac M, Lampreht U, Preat V, Vandermeulen G, Butinar M, Turk B, Cemazar M (2015) Endoglin (CD105) silencing mediated by shRNA under the control of endothelin-1 promoter for targeted gene therapy of melanoma. Mol Ther Nucleic Acids 5(4):e239. https://doi.org/10.1038/mtna.2015.12
Li Y, Yang X, Lu X, Peng Z, Lai C, Xie S, Wei S, Yao H, Ding Z, Zhao X, Liu A, Hou X, Mo F (2020) Recombinant endoglin-single-chain variable fragment/induced protein 10 fusion protein potently boosts the anti-tumor efficacy of adoptively transferred TRP2-specific CD8+ CD28+ cytotoxic T lymphocytes in mice. J Biomed Nanotechnol 16(7):1119–1134. https://doi.org/10.1166/jbn.2020.2949
Savarin M, Kamensek U, Znidar K, Todorovic V, Sersa G, Cemazar M (2021) Evaluation of a novel plasmid for simultaneous gene electrotransfer-mediated silencing of CD105 and CD146 in combination with irradiation. Int J Mol Sci 22(6):3069. https://doi.org/10.3390/ijms22063069
Altomonte M, Montagner R, Fonsatti E, Colizzi F, Cattarossi I, Brasoveanu LI, Nicotra MR, Cattelan A, Natali PG, Maio M (1996) Expression and structural features of endoglin (CD105), a transforming growth factor beta1 and beta3 binding protein, in human melanoma. Br J Cancer 74(10):1586–1591
Salgado KB, Toscani NV, Silva LL, Hilbig A, Barbosa-Coutinho LM (2007) Immunoexpression of endoglin in brain metastasis secondary to malignant melanoma: evaluation of angiogenesis and comparison with brain metastasis secondary to breast and lung carcinomas. Clin Exp Metastasis 24(6):403–410
Muñoz R, Arias Y, Ferreras JM, Jiménez P, Langa C, Rojo MA, Gayoso MJ, Córdoba-Díaz D, Bernabéu C, Girbés T (2013) In vitro and in vivo effects of an anti-mouse endoglin (CD105)-immunotoxin on the early stages of mouse B16MEL4A5 melanoma tumours. Cancer Immunol Immunother 62(3):541–551. https://doi.org/10.1007/s00262-012-1357-7
Dolinsek T, Sersa G, Prosen L, Bosnjak M, Stimac M, Razborsek U, Cemazar M (2015) Electrotransfer of plasmid DNA encoding an anti-mouse endoglin (CD105) shRNA to B16 melanoma tumors with low and high metastatic potential results in pronounced anti-tumor effects. Cancers (Basel) 8(1):3. https://doi.org/10.3390/cancers8010003
Schadendorf D, van Akkooi ACJ, Berking C, Griewank KG, Gutzmer R, Hauschild A, Stang A, Roesch A, Ugurel S (2018) Melanoma. Lancet 392(10151):971–984. https://doi.org/10.1016/S0140-6736(18)31559-9
Albittar AA, Alhalabi O, Glitza Oliva IC (2020) Immunotherapy for melanoma. Adv Exp Med Biol 1244:51–68. https://doi.org/10.1007/978-3-030-41008-7_3
Bernabeu C, Conley BA, Vary CP (2007) Novel biochemical pathways of endoglin in vascular cell physiology. J Cell Biochem 102(6):1375–1388. https://doi.org/10.1002/jcb.21594
Llorca O, Trujillo A, Blanco FJ, Bernabeu C (2007) Structural model of human endoglin, a transmembrane receptor responsible for hereditary hemorrhagic telangiectasia. J Mol Biol 365(3):694–705. https://doi.org/10.1016/j.jmb.2006.10.015
Saito T, Bokhove M, Croci R, Zamora-Caballero S, Han L, Letarte M, de Sanctis D, Jovine L (2017) Structural basis of the human endoglin-BMP9 interaction: insights into BMP signaling and HHT1. Cell Rep 19(9):1917–1928. https://doi.org/10.1016/j.celrep.2017.05.011
Alt A, Miguel-Romero L, Donderis J, Aristorena M, Blanco FJ, Round A, Rubio V, Bernabeu C, Marina A (2012) Structural and functional insights into endoglin ligand recognition and binding. PLoS ONE 7(2):e29948. https://doi.org/10.1371/journal.pone.0029948
Rossi E, Smadja DM, Boscolo E, Langa C, Arevalo MA, Pericacho M, Gamella-Pozuelo L, Kauskot A, Botella LM, Gaussem P, Bischoff J, Lopez-Novoa JM, Bernabeu C (2016) Endoglin regulates mural cell adhesion in the circulatory system. Cell Mol Life Sci 73(8):1715–1739. https://doi.org/10.1007/s00018-015-2099-4
Ruiz-Llorente L, Gallardo-Vara E, Rossi E, Smadja DM, Botella LM, Bernabeu C (2017) Endoglin and alk1 as therapeutic targets for hereditary hemorrhagic telangiectasia. Expert Opin Ther Targets 21(10):933–947. https://doi.org/10.1080/14728222.2017.1365839
Bernabeu C, Bayrak-Toydemir P, McDonald J, Letarte M (2020) Potential second-hits in hereditary hemorrhagic telangiectasia. J Clin Med 9(11):3571. https://doi.org/10.3390/jcm9113571
Guerrero-Esteo M, Sanchez-Elsner T, Letamendia A, Bernabeu C (2002) Extracellular and cytoplasmic domains of endoglin interact with the transforming growth factor-beta receptors I and II. J Biol Chem 277(32):29197–29209. https://doi.org/10.1074/jbc.M111991200
Blanco FJ, Santibanez JF, Guerrero-Esteo M, Langa C, Vary CP, Bernabeu C (2005) Interaction and functional interplay between endoglin and ALK-1, two components of the endothelial transforming growth factor-beta receptor complex. J Cell Physiol 204(2):574–584. https://doi.org/10.1002/jcp.20311
Hodgson J, Ruiz-Llorente L, McDonald J, Quarrell O, Ugonna K, Bentham J, Mason R, Martin J, Moore D, Bergstrom K, Bayrak-Toydemir P, Wooderchak-Donahue W, Morrell NW, Condliffe R, Bernabeu C, Upton PD (2021) Homozygous GDF2 nonsense mutations result in a loss of circulating BMP9 and BMP10 and are associated with either PAH or an “HHT-like” syndrome in children. Mol Genet Genomic Med 9(12):e1685. https://doi.org/10.1002/mgg3.1685
Cummings SD, Ryu B, Samuels MA, Yu X, Meeker AK, Healey MA, Alani RM (2008) Id1 delays senescence of primary human melanocytes. Mol Carcinog 47(9):653–659. https://doi.org/10.1002/mc.20422
Hawinkels LJ, de Vinuesa AG, Paauwe M, Kruithof-de Julio M, Wiercinska E, Pardali E, Mezzanotte L, Keereweer S, Braumuller TM, Heijkants RC, Jonkers J, Löwik CW, Goumans MJ, ten Hagen TL, ten Dijke P (2016) Activin receptor-like kinase 1 ligand trap reduces microvascular density and improves chemotherapy efficiency to various solid tumors. Clin Cancer Res 22(1):96–106. https://doi.org/10.1158/1078-0432.CCR-15-0743
Peres J, Damerell V, Chauhan J, Popovic A, Desprez PY, Galibert MD, Goding CR, Prince S (2021) TBX3 promotes melanoma migration by transcriptional activation of ID1, which prevents activation of E-cadherin by MITF. J Invest Dermatol 141(9):2250-2260.e2. https://doi.org/10.1016/j.jid.2021.02.740
Rossi E, Pericacho M, Bachelot-Loza C, Pidard D, Gaussem P, Poirault-Chassac S, Blanco FJ, Langa C, González-Manchón C, Novoa JML, Smadja DM, Bernabeu C (2018) Human endoglin as a potential new partner involved in platelet-endothelium interactions. Cell Mol Life Sci 75(7):1269–1284. https://doi.org/10.1007/s00018-017-2694-7
Huang R, Rofstad EK (2018) Integrins as therapeutic targets in the organ-specific metastasis of human malignant melanoma. J Exp Clin Cancer Res 37(1):92. https://doi.org/10.1186/s13046-018-0763-x
Arias-Mejias SM, Warda KY, Quattrocchi E, Alonso-Quinones H, Sominidi-Damodaran S, Meves A (2020) The role of integrins in melanoma: a review. Int J Dermatol 59(5):525–534. https://doi.org/10.1111/ijd.14850
Belik D, Tsang H, Wharton J, Howard L, Bernabeu C, Wojciak-Stothard B (2016) Endothelium-derived microparticles from chronically thromboembolic pulmonary hypertensive patients facilitate endothelial angiogenesis. J Biomed Sci 19(23):4. https://doi.org/10.1186/s12929-016-0224-9
Ermini L, Ausman J, Melland-Smith M, Yeganeh B, Rolfo A, Litvack ML, Todros T, Letarte M, Post M, Caniggia I (2017) A Single sphingomyelin species promotes exosomal release of endoglin into the maternal circulation in preeclampsia. Sci Rep 7(1):12172. https://doi.org/10.1038/s41598-017-12491-4
Meurer S, Wimmer AE, Leur EV, Weiskirchen R (2019) Endoglin trafficking/exosomal targeting in liver cells depends on N-glycosylation. Cells 8(9):997. https://doi.org/10.3390/cells8090997
Vicen M, Igreja Sá IC, Tripská K, Vitverová B, Najmanová I, Eissazadeh S, Micuda S, Nachtigal P (2021) Membrane and soluble endoglin role in cardiovascular and metabolic disorders related to metabolic syndrome. Cell Mol Life Sci 78(6):2405–2418. https://doi.org/10.1007/s00018-020-03701-w
Willms E, Cabañas C, Mäger I, Wood MJA, Vader P (2018) Extracellular vesicle heterogeneity: subpopulations, isolation techniques, and diverse functions in cancer progression. Front Immunol 30(9):738. https://doi.org/10.3389/fimmu.2018.00738
Zeng EZ, Chen I, Chen X, Yuan X (2022) Exosomal microRNAs as novel cell-free therapeutics in tissue engineering and regenerative medicine. Biomedicines 10(10):2485. https://doi.org/10.3390/biomedicines10102485
Han C, Yang J, Sun J, Qin G (2022) Extracellular vesicles in cardiovascular disease: biological functions and therapeutic implications. Pharmacol Ther 233:108025. https://doi.org/10.1016/j.pharmthera.2021.108025
Peng Y, Croce CM (2016) The role of microRNAs in human cancer. Signal Transduct Target Ther 28(1):15004. https://doi.org/10.1038/sigtrans.2015.4
Sun Z, Shi K, Yang S, Liu J, Zhou Q, Wang G, Song J, Li Z, Zhang Z, Yuan W (2018) Effect of exosomal miRNA on cancer biology and clinical applications. Mol Cancer 17(1):147. https://doi.org/10.1186/s12943-018-0897-7
Hill M, Tran N (2021) miRNA interplay: mechanisms and consequences in cancer. Dis Model Mech 14(4):dmm047662. https://doi.org/10.1242/dmm.047662
Menon A, Abd-Aziz N, Khalid K, Poh CL, Naidu R (2022) miRNA: a promising therapeutic target in cancer. Int J Mol Sci 23(19):11502. https://doi.org/10.3390/ijms231911502
Thyagarajan A, Shaban A, Sahu RP (2018) MicroRNA-directed cancer therapies: implications in melanoma intervention. J Pharmacol Exp Ther 364(1):1–12. https://doi.org/10.1124/jpet.117.242636
Durante G, Broseghini E, Comito F, Naddeo M, Milani M, Salamon I, Campione E, Dika E, Ferracin M (2022) Circulating microRNA biomarkers in melanoma and non-melanoma skin cancer. Expert Rev Mol Diagn 22(3):305–318. https://doi.org/10.1080/14737159.2022.2049243
Huang N, Lee KJ, Stark MS (2022) Current trends in circulating biomarkers for melanoma detection. Front Med (Lausanne) 5(9):873728. https://doi.org/10.3389/fmed.2022.873728
Chen XP, Chen YG, Lan JY, Shen ZJ (2014) MicroRNA-370 suppresses proliferation and promotes endometrioid ovarian cancer chemosensitivity to cDDP by negatively regulating ENG. Cancer Lett 353:201–210. https://doi.org/10.1016/j.canlet.2014.07.026
Ruiz-Llorente L, Albiñana V, Botella LM, Bernabeu C (2020) Differential expression of circulating plasma miRNA-370 and miRNA-10a from patients with hereditary hemorrhagic telangiectasia. J Clin Med 9(9):2855. https://doi.org/10.3390/jcm9092855
Neagu M, Constantin C, Cretoiu SM, Zurac S (2020) miRNAs in the diagnosis and prognosis of skin cancer. Front Cell Dev Biol 28(8):71. https://doi.org/10.3389/fcell.2020.00071
Penna E, Orso F, Taverna D (2015) miR-214 as a key hub that controls cancer networks: small player, multiple functions. J Invest Dermatol 135(4):960–969. https://doi.org/10.1038/jid.2014.479
Sim J, Ahn H, Abdul R, Kim H, Yi KJ, Chung YM, Chung MS, Paik SS, Song YS, Jang K (2015) High microRNA-370 expression correlates with tumor progression and poor prognosis in breast cancer. J Breast Cancer 18(4):323–328. https://doi.org/10.4048/jbc.2015.18.4.323
Wu Z, Sun H, Zeng W, He J, Mao X (2012) Upregulation of mircoRNA-370 induces proliferation in human prostate cancer cells by downregulating the transcription factor FOXO1. PLoS ONE 7(9):e45825. https://doi.org/10.1371/journal.pone.0045825
Zeng Y, Fu M, Wu GW, Zhang AZ, Chen JP, Lin HY, Fu YA, Jia J, Cai ZD, Wu XJ, Lan P (2016) Upregulation of microRNA-370 promotes cell apoptosis and inhibits proliferation by targeting PTEN in human gastric cancer. Int J Oncol 49(4):1589–1599. https://doi.org/10.3892/ijo.2016.3642
Chen M, Xia Y, Tan Y, Jiang G, Jin H, Chen Y (2018) Downregulation of microRNA-370 in esophageal squamous-cell carcinoma is associated with cancer progression and promotes cancer cell proliferation via upregulating PIN1. Gene 30(661):68–77. https://doi.org/10.1016/j.gene.2018.03.090
Pan XP, Wang HX, Tong DM, Li Y, Huang LH, Wang C (2017) miRNA-370 acts as a tumor suppressor via the downregulation of PIM1 in hepatocellular carcinoma. Eur Rev Med Pharmacol Sci 21(6):1254–1263
Wei S, Ma W (2017) MiR-370 functions as oncogene in melanoma by direct targeting pyruvate dehydrogenase B. Biomed Pharmacother 90:278–286. https://doi.org/10.1016/j.biopha.2017.03.068
Muqaku B, Eisinger M, Meier SM, Tahir A, Pukrop T, Haferkamp S, Slany A, Reichle A, Gerner C (2017) Multi-omics analysis of serum samples demonstrates reprogramming of organ functions via systemic calcium mobilization and platelet activation in metastatic melanoma. Mol Cell Proteomics 16(1):86–99. https://doi.org/10.1074/mcp.M116.063313
Zhang S, Chen K, Liu H, Jing C, Zhang X, Qu C, Yu S (2021) PMEL as a prognostic biomarker and negatively associated with immune infiltration in skin cutaneous melanoma (SKCM). J Immunother 44(6):214–223. https://doi.org/10.1097/CJI.0000000000000374
Sendoel A, Kohler I, Fellmann C, Lowe SW, Hengartner MO (2010) HIF-1 antagonizes p53-mediated apoptosis through a secreted neuronal tyrosinase. Nature 465(7298):577–583. https://doi.org/10.1038/nature09141
Dass SE, Huizenga T, Farshchian M, Mehregan DR (2021) Comparison of SOX-10, HMB-45, and melan-A in benign melanocytic lesions. Clin Cosmet Investig Dermatol 5(14):1419–1425. https://doi.org/10.2147/CCID.S333376
Vincek E, Rudnick E (2022) Melanocytic marker melan-A detects molluscum contagiosum bodies. J Histotechnol 45(1):36–38. https://doi.org/10.1080/01478885.2021.1964872
Abrahamian C, Grimm C (2021) Endolysosomal cation channels and MITF in melanocytes and melanoma. Biomolecules 11(7):1021. https://doi.org/10.3390/biom11071021
Goding CR, Arnheiter H (2019) MITF-the first 25 years. Genes Dev 33(15–16):983–1007. https://doi.org/10.1101/gad.324657.119
AlAbdi L, He M, Yang Q, Norvil AB, Gowher H (2018) The transcription factor Vezf1 represses the expression of the antiangiogenic factor Cited2 in endothelial cells. J Biol Chem 293(28):11109–11118. https://doi.org/10.1074/jbc.RA118.002911
Shi X, Zhao P, Zhao G (2022) VEZF1, destabilized by STUB1, affects cellular growth and metastasis of hepatocellular carcinoma by transcriptionally regulating PAQR4. Cancer Gene Ther. https://doi.org/10.1038/s41417-022-00540-8
Wong VC, Chan PL, Bernabeu C, Law S, Wang LD, Li JL, Tsao SW, Srivastava G, Lung ML (2008) Identification of an invasion and tumor-suppressing gene, endoglin (ENG), silenced by both epigenetic inactivation and allelic loss in esophageal squamous cell carcinoma. Int J Cancer 123(12):2816–2823. https://doi.org/10.1002/ijc.23882
O’Leary K, Shia A, Cavicchioli F, Haley V, Comino A, Merlano M, Mauri F, Walter K, Lackner M, Wischnewsky MB, Crook T, Lo Nigro C, Schmid P (2015) Identification of Endoglin as an epigenetically regulated tumour-suppressor gene in lung cancer. Br J Cancer 113(6):970–978. https://doi.org/10.1038/bjc.2015.302
Peinado H, Zhang H, Matei IR, Costa-Silva B, Hoshino A, Rodrigues G, Psaila B, Kaplan RN, Bromberg JF, Kang Y, Bissell MJ, Cox TR, Giaccia AJ, Erler JT, Hiratsuka S, Ghajar CM, Lyden D (2017) Pre-metastatic niches: organ-specific homes for metastases. Nat Rev Cancer 17(5):302–317. https://doi.org/10.1038/nrc.2017.6
Nogués L, Benito-Martin A, Hergueta-Redondo M, Peinado H (2018) The influence of tumour-derived extracellular vesicles on local and distal metastatic dissemination. Mol Aspects Med 60:15–26. https://doi.org/10.1016/j.mam.2017.11.012
Cardeñes B, Clares I, Bezos T, Toribio V, López-Martín S, Rocha A, Peinado H, Yáñez-Mó M, Cabañas C (2022) ALCAM/CD166 is involved in the binding and uptake of cancer-derived extracellular vesicles. Int J Mol Sci 23(10):5753. https://doi.org/10.3390/ijms23105753
Bhatta B, Cooks T (2020) Reshaping the tumor microenvironment: extracellular vesicles as messengers of cancer cells. Carcinogenesis 41(11):1461–1470. https://doi.org/10.1093/carcin/bgaa107
Gunaydin G (2021) CAFs interacting with TAMs in tumor microenvironment to enhance tumorigenesis and immune evasion. Front Oncol 14(11):668349. https://doi.org/10.3389/fonc.2021.668349
Su T, Zhang P, Zhao F, Zhang S (2021) Exosomal microRNAs mediating crosstalk between cancer cells with cancer-associated fibroblasts and tumor-associated macrophages in the tumor microenvironment. Front Oncol 1(11):631703. https://doi.org/10.3389/fonc.2021.631703
Laplagne C, Domagala M, Le Naour A, Quemerais C, Hamel D, Fournié JJ, Couderc B, Bousquet C, Ferrand A, Poupot M (2019) Latest advances in targeting the tumor microenvironment for tumor suppression. Int J Mol Sci 20(19):4719. https://doi.org/10.3390/ijms20194719
Cao H, Gao S, Jogani R, Sugimura R (2022) The tumor microenvironment reprograms immune cells. Cell Reprogram. https://doi.org/10.1089/cell.2022.0047
Xiao Y, Yu D (2021) Tumor microenvironment as a therapeutic target in cancer. Pharmacol Ther 221:107753. https://doi.org/10.1016/j.pharmthera.2020.107753
Pernot S, Evrard S, Khatib AM (2022) The give-and-take interaction between the tumor microenvironment and immune cells regulating tumor progression and repression. Front Immunol 13(13):850856. https://doi.org/10.3389/fimmu.2022.850856
Bellei B, Migliano E, Picardo M (2020) A framework of major tumor-promoting signal transduction pathways implicated in melanoma-fibroblast dialogue. Cancers (Basel) 12(11):3400. https://doi.org/10.3390/cancers12113400
Romano V, Belviso I, Venuta A, Ruocco MR, Masone S, Aliotta F, Fiume G, Montagnani S, Avagliano A, Arcucci A (2021) Influence of tumor microenvironment and fibroblast population plasticity on melanoma growth, therapy resistance and immunoescape. Int J Mol Sci 22(10):5283. https://doi.org/10.3390/ijms22105283
Érsek B, Silló P, Cakir U, Molnár V, Bencsik A, Mayer B, Mezey E, Kárpáti S, Pós Z, Németh K (2021) Melanoma-associated fibroblasts impair CD8+ T cell function and modify expression of immune checkpoint regulators via increased arginase activity. Cell Mol Life Sci 78(2):661–673. https://doi.org/10.1007/s00018-020-03517-8
Zhou J, Yang Y, Wang W, Zhang Y, Chen Z, Hao C, Zhang J (2018) Melanoma-released exosomes directly activate the mitochondrial apoptotic pathway of CD4+ T cells through their microRNA cargo. Exp Cell Res 371(2):364–371. https://doi.org/10.1016/j.yexcr.2018.08.030
Simiczyjew A, Dratkiewicz E, Mazurkiewicz J, Ziętek M, Matkowski R, Nowak D (2020) The influence of tumor microenvironment on immune escape of melanoma. Int J Mol Sci 21(21):8359. https://doi.org/10.3390/ijms21218359
Gerloff D, Lützkendorf J, Moritz RKC, Wersig T, Mäder K, Müller LP, Sunderkötter C (2020) Melanoma-derived exosomal miR-125b-5p educates tumor associated macrophages (TAMs) by targeting lysosomal acid lipase A (LIPA). Cancers (Basel) 12(2):464. https://doi.org/10.3390/cancers12020464
Ren Z, Lv M, Yu Q, Bao J, Lou K, Li X (2021) MicroRNA-370-3p shuttled by breast cancer cell-derived extracellular vesicles induces fibroblast activation through the CYLD/Nf-κB axis to promote breast cancer progression. FASEB J 35(3):e21383. https://doi.org/10.1096/fj.202001430RR
Paauwe M, Schoonderwoerd MJA, Helderman RFCP, Harryvan TJ, Groenewoud A, van Pelt GW, Bor R, Hemmer DM, Versteeg HH, Snaar-Jagalska BE, Theuer CP, Hardwick JCH, Sier CFM, Ten Dijke P, Hawinkels LJAC (2018) Endoglin expression on cancer-associated fibroblasts regulates invasion and stimulates colorectal cancer metastasis. Clin Cancer Res 24(24):6331–6344. https://doi.org/10.1158/1078-0432.CCR-18-0329
Wei CY, Zhu MX, Lu NH, Liu JQ, Yang YW, Zhang Y, Shi YD, Feng ZH, Li JX, Qi FZ, Gu JY (2020) Circular RNA circ_0020710 drives tumor progression and immune evasion by regulating the miR-370-3p/CXCL12 axis in melanoma. Mol Cancer 19(1):84. https://doi.org/10.1186/s12943-020-01191-9
McConnell AT, Ellis R, Pathy B, Plummer R, Lovat PE, O’Boyle G (2016) The prognostic significance and impact of the CXCR4-CXCR7-CXCL12 axis in primary cutaneous melanoma. Br J Dermatol 175(6):1210–1220. https://doi.org/10.1111/bjd.14720
Wang R, Sun Y, Yu W, Yan Y, Qiao M, Jiang R, Guan W, Wang L (2019) Downregulation of miRNA-214 in cancer-associated fibroblasts contributes to migration and invasion of gastric cancer cells through targeting FGF9 and inducing EMT. J Exp Clin Cancer Res 38(1):20. https://doi.org/10.1186/s13046-018-0995-9
Kuninty PR, Bojmar L, Tjomsland V, Larsson M, Storm G, Östman A, Sandström P, Prakash J (2016) MicroRNA-199a and -214 as potential therapeutic targets in pancreatic stellate cells in pancreatic tumor. Oncotarget 7(13):16396–16408. https://doi.org/10.18632/oncotarget.7651
Yin Y, Cai X, Chen X, Liang H, Zhang Y, Li J, Wang Z, Chen X, Zhang W, Yokoyama S, Wang C, Li L, Li L, Hou D, Dong L, Xu T, Hiroi T, Yang F, Ji H, Zhang J, Zen K, Zhang CY (2014) Tumor-secreted miR-214 induces regulatory T cells: a major link between immune evasion and tumor growth. Cell Res 24(10):1164–1180. https://doi.org/10.1038/cr.2014.121
Aladowicz E, Lanfrancone L (2014) Investigating the metastatic niche in melanoma: a new therapeutic opportunity? Future Oncol 10(5):699–701. https://doi.org/10.2217/fon.14.26
Statements and Declarations
Funding
This work was supported by grants from Ministerio de Ciencia, Innovación y Universidades (SAF2013-43421-R to CB), Consejo Superior de Investigaciones Científicas (201920E022 to CB), Fondo de Investigación Sanitaria (FIS) de la Seguridad Social (PI-20/01553 to JLR-P), and Centro de Investigación Biomédica en Red de Enfermedades Raras (CIBERER; ISCIII-CB06/07/0038 to CB and contract CNV-234-PRF-360 to LR-L) of Spain. CIBERER is an initiative of the Instituto de Salud Carlos III (ISCIII) of Spain supported by FEDER funds. The CNIO, certified as a Severo Ochoa Excellence Centre, is supported by the Spanish government through the ISCIII.
Disclosure of Interests
All authors declare they have no conflict of interest.
Ethical Approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed Consent
Informed consent for participation and publication was obtained from all individual participants included in the study.
Acknowledgements
The authors would like to thank Carmen Langa and Elena de Blas for technical support, Dr. María S. Soengas (Spanish National Cancer Research Centre [CNIO], Madrid, Spain) for melanoma cell lines, Prof. Peter ten Dijke (LUMC, Leiden, The Netherlands) for lentiviral constructs expressing human endoglin, and Prof. Dr. César Menor-Salván for support and advice with BioRender software.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2023 The Author(s)
About this chapter
Cite this chapter
Ruiz-Llorente, L. et al. (2023). Correlation Between Endoglin and Malignant Phenotype in Human Melanoma Cells: Analysis of hsa-mir-214 and hsa-mir-370 in Cells and Their Extracellular Vesicles. In: Simon, F., Bernabeu, C. (eds) Advances in Molecular Pathology. Advances in Experimental Medicine and Biology, vol 1408. Springer, Cham. https://doi.org/10.1007/978-3-031-26163-3_14
Download citation
DOI: https://doi.org/10.1007/978-3-031-26163-3_14
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-26162-6
Online ISBN: 978-3-031-26163-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)