Abstract
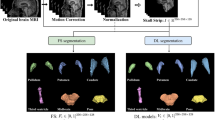
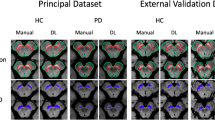
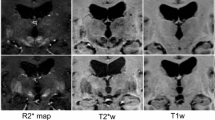
Automatic segmentation of substantia nigra (SN), which is Parkinson’s disease-related tissue, is an important step toward accurate computer-aided diagnosis systems. Conventional methods for SN segmentation depend heavily on limited magnetic resonance imaging (MRI) modalities such as neuromelanin and quantitative susceptibility mapping, which require longer imaging times and are rare in public datasets. To enable a multi-modal investigation for SN anatomic alterations based on medical bigdata researches, the need for automated SN segmentation arises from commonly investigated T2-weighted MRIs. To improve the performance of the automated SN segmentation from a T2-weighted MRI and enhance the model generalization for cross-center researches, this paper proposes a novel test-time normalization (TTN) method to increase the geometric and intensity similarity between the query data and the model’s trained data. Our proposed method requires no additional training procedure or extra annotation for the unseen data. Our results showed that our proposed TTN achieved a mean Dice score of 71.08% in comparison with the baseline model’s 69.87% score with in-house dataset. Additionally, improved SN segmentation performance was observed from the unseen and unlabeled datasets.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Fearnley, J.M., Lees, A.J.: Ageing and Parkinson’s disease: substantia Nigra regional selectivity. Brain 114(5), 2283–2301 (1991)
Ogisu, K., et al.: 3D neuromelanin-sensitive magnetic resonance imaging with semi-automated volume measurement of the substantia nigra pars compacta for diagnosis of Parkinson’s disease. Neuroradiology 55(6), 719–724 (2013)
Hatano, T., et al.: Neuromelanin MRI is useful for monitoring motor complications in Parkinson’s and PARK2 disease. J. Neural Transm. 124(4), 407–415 (2017)
Castellanos, G., et al.: Automated neuromelanin imaging as a diagnostic biomarker for Parkinson’s Disease. Mov. Disord. 30(7), 945–952 (2015)
Zhao, X., Wu, Y., Song, G., Li, Z., Zhang, Y., Fan, Y.: A deep learning model integrating FCNNs and CRFs for brain tumor segmentation. Med. Image Anal. 43, 98–111 (2018)
Shen, C., et al.: Spatial information-embedded fully convolutional networks for multi-organ segmentation with improved data augmentation and instance normalization. In: Medical Imaging 2020: Image Processing, pp. 261–267. SPIE, Houston (2020)
Hu, T., et al.: Aorta-aware GAN for non-contrast to artery contrasted CT translation and its application to abdominal aortic aneurysm detection. Int. J. Comput. Assist. Radiol. Surg. 17(1), 97–105 (2021). https://doi.org/10.1007/s11548-021-02492-0
Le Berre, A., et al.: Convolutional neural network-based segmentation can help in assessing the substantia Nigra in neuromelanin MRI. Neuroradiology 61(12), 1387–1395 (2019). https://doi.org/10.1007/s00234-019-02279-w
Ronneberger, O., Fischer, P., Brox, T.: U-net: convolutional networks for biomedical image segmentation. In: Navab, N., Hornegger, J., Wells, W.M., Frangi, A.F. (eds.) MICCAI 2015. LNCS, vol. 9351, pp. 234–241. Springer, Cham (2015). https://doi.org/10.1007/978-3-319-24574-4_28
Hashemi, S.R., Salehi, S.S., Erdogmus, D., Prabhu, S.P., Warfield, S.K., Gholipour, A.: Asymmetric loss functions and deep densely-connected networks for highly-imbalanced medical image segmentation: Application to multiple sclerosis lesion detection. IEEE Access 7, 1721–1735 (2018)
Wang, G., Li, W., Aertsen, M., Deprest, J., Ourselin, S., Vercauteren, T.: Aleatoric uncertainty estimation with test-time augmentation for medical image segmentation with convolutional neural networks. Neurocomputing 338, 34–45 (2019)
Bourke, P.: Histogram Matching. http://paulbourke.net/miscellaneous/equalisation/. (2011)
Milletari, F., Navab, N., Ahmadi, S.A.: V-Net: Fully convolutional neural networks for volumetric medical image segmentation. In: 2016 fourth international conference on 3D vision (3DV), pp. 565–571. IEEE, Stanford (2016)
Acknowledgements
We appreciate the help and advice from the members of the Mori laboratory. A part of this research was supported by the AMED Grant Numbers 22dm0307101h0004 and JSPS KAKENHI 21k19898, 17K20099.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Hu, T. et al. (2022). Enhancing Model Generalization for Substantia Nigra Segmentation Using a Test-time Normalization-Based Method. In: Wang, L., Dou, Q., Fletcher, P.T., Speidel, S., Li, S. (eds) Medical Image Computing and Computer Assisted Intervention – MICCAI 2022. MICCAI 2022. Lecture Notes in Computer Science, vol 13437. Springer, Cham. https://doi.org/10.1007/978-3-031-16449-1_70
Download citation
DOI: https://doi.org/10.1007/978-3-031-16449-1_70
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-16448-4
Online ISBN: 978-3-031-16449-1
eBook Packages: Computer ScienceComputer Science (R0)