Abstract
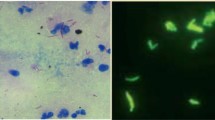
Tuberculosis is one of the most serious infectious diseases, and its treatment is highly dependent on early detection. Microscopy-based analysis of sputum images for bacilli identification is a common technique used for both diagnosis and treatment monitoring. However, it a challenging process since sputum analysis requires time and highly trained experts to avoid potentially fatal mistakes. Capturing fields of view (FOVs) from high resolution whole slide images is a laborious procedure, since they are manually localized and then examined to determine the presence of bacteria. In the present paper we propose a method that automates the process, thus greatly reducing the amount of human labour. In particular, we (i) describe an image processing based method for the extraction of a FOV representation which emphasises salient, bacterial content, while suppressing confounding visual information, and (ii) introduce a novel deep learning based architecture which learns from coarsely labelled FOV images and the corresponding binary masks, and then classifies novel FOV images as salient (bacteria containing) or not. Using a real-world data corpus, the proposed method is shown to outperform 12 state of the art methods in the literature, achieving (i) an approximately 10% lower overall error rate than the next best model and (ii) perfect sensitivity (7% higher than the next best model).
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Arandjelović, O., Cipolla, R.: A new look at filtering techniques for illumination invariance in automatic face recognition, pp. 449–454 (2006)
Baron, V.O., et al.: Label-free optical vibrational spectroscopy to detect the metabolic state of M. tuberculosis cells at the site of disease. Sci. Rep. 7(1), 1–9 (2017)
Costa Filho, C.F.F., Costa, M.G.F., Júnior, A.K.: Autofocus functions for tuberculosis diagnosis with conventional sputum smear microscopy. In: Current Microscopy Contributions to Advances in Science and Technology, pp. 13–20 (2012)
Daniel, J., Kapoor, N., Sirakova, T., Sinha, R., Kolattukudy, P.: The perilipin-like PPE15 protein in Mycobacterium tuberculosis is required for triacylglycerol accumulation under dormancy-inducing conditions. Mol. Microbiol. 101(5), 784–794 (2016)
Deb, C., et al.: A novel in vitro multiple-stress dormancy model for mycobacterium tuberculosis generates a lipid-loaded, drug-tolerant, dormant pathogen. PLoS ONE 4(6), e6077 (2009)
Forero, M.G., Sroubek, F., Cristóbal, G.: Identification of tuberculosis bacteria based on shape and color. Real-Time Imaging 10(4), 251–262 (2004)
Gele, A.A., Bjune, G., Abebe, F.: Pastoralism and delay in diagnosis of TB in Ethiopia. BMC Public Health 9(1), 1–7 (2009)
Greenspan, P., Fowler, S.D.: Spectrofluorometric studies of the lipid probe, nile red. J. Lipid Res. 26(7), 781–789 (1985)
He, D., Xia, Y., Qin, T., Wang, L., Yu, N., Liu, T.Y., Ma, W.Y.: Dual learning for machine translation. Adv. Neural Inf. Process. Syst. 29, 820–828 (2016)
Holmes, C.B., Hausler, H., Nunn, P.: A review of sex differences in the epidemiology of tuberculosis. Int. J. Tuberc. Lung Dis. 2(2), 96–104 (1998)
Huang, G., Liu, Z., Van Der Maaten, L., Weinberger, K.Q.: Densely connected convolutional networks. In: IEEE Conference on Computer Vision and Pattern Recognition, pp. 4700–4708 (2017)
Kant, S., Srivastava, M.M.: Towards automated tuberculosis detection using deep learning. In: IEEE Symposium Series on Computational Intelligence, pp. 1250–1253 (2019)
Kayigire, X.A., Friedrich, S.O., van der Merwe, L., Donald, P.R., Diacon, A.H.: Simultaneous staining of sputum smears for acid-fast and lipid-containing Myobacterium tuberculosis can enhance the clinical evaluation of antituberculosis treatments. Tuberculosis 95(6), 770–779 (2015)
Kennedy, J.A., Baron, V., Hammond, R.J.H., Sloan, D.J., Gillespie, S.H.: Centrifugation and decontamination procedures selectively impair recovery of important populations in Mycobacterium smegmatis. Tuberculosis 112, 79–82 (2018)
Kumar, N.C.S., Radhika, Y.: Optimized maximum principal curvatures based segmentation of blood vessels from retinal images. Biomed. Res. 30(2) (2019)
Lomacenkova, A., Arandjelović, O.: Whole slide pathology image patch based deep classification: an investigation of the effects of the latent autoencoder representation and the loss function form. In: Proceedings of IEEE International Conference on Biomedical and Health Informatics (2021). https://doi.org/10.1109/BHI50953.2021.9508577
Mehta, P.K., Raj, A., Singh, N., Khuller, G.K.: Diagnosis of extrapulmonary tuberculosis by PCR. FEMS Immunol. Med. Microbiol. 66(1), 20–36 (2012)
Merchant, F.A., Castleman, K.R.: Computer-assisted microscopy. In: The Essential Guide to Image Processing, pp. 777–831 (2009)
Panicker, R.O., Kalmady, K.S., Rajan, J., Sabu, M.K.: Automatic detection of tuberculosis bacilli from microscopic sputum smear images using deep learning methods. Biocybernetics Biomed. Eng. 38(3), 691–699 (2018)
Peter, J.G., van Zyl-Smit, R.N., Denkinger, C.M., Pai, M.: Diagnosis of TB: state of the art. Eur. Respir. Monograph 58, 123–143 (2012)
Phillips, P.P.J., et al.: Limited role of culture conversion for decision-making in individual patient care and for advancing novel regimens to confirmatory clinical trials. BMC Med. 14(1), 1–11 (2016)
Rieder, H.L., et al.: Priorities for Tuberculosis Bacteriology Services in Low-Income Countries. International Union Against Tuberculosis and Lung Disease (2007)
Rumin, J., Bonnefond, H., Saint-Jean, B., Rouxel, C., Sciandra, A., Bernard, O., Cadoret, J.P., Bougaran, G.: The use of fluorescent Nile red and BODIPY for lipid measurement in microalgae. Biotechnol. Biofuels 8(1), 1–16 (2015)
Sadaphal, P., Rao, J., Comstock, G.W., Beg, M.F.: Image processing techniques for identifying Mycobacterium tuberculosis in Ziehl-Neelsen stains. Int. J. Tuberc. Lung Dis. 12(5), 579–582 (2008)
Shea, Y.R., et al.: High sensitivity and specificity of acid-fast microscopy for diagnosis of pulmonary tuberculosis in an African population with a high prevalence of human immunodeficiency virus. J. Clin. Microbiol. 47(5), 1553–1555 (2009)
Simonyan, K., Zisserman, A.: Very deep convolutional networks for large-scale image recognition. arXiv preprint arXiv:1409.1556 (2014)
Sloan, D.J., et al.: Pharmacodynamic modeling of bacillary elimination rates and detection of bacterial lipid bodies in sputum to predict and understand outcomes in treatment of pulmonary tuberculosis. Clin. Infect. Dis. 61(1), 1–8 (2015)
Smith, L.N.: Cyclical learning rates for training neural networks. In: IEEE Winter Conference on Applications of Computer Vision, pp. 464–472 (2017)
Sotaquira, M., Rueda, L., Narvaez, R.: Detection and quantification of bacilli and clusters present in sputum smear samples: a novel algorithm for pulmonary tuberculosis diagnosis. In: International Conference on Digital Image Processing, pp. 117–121 (2009)
Spence, D.P., Hotchkiss, J., Williams, C.S., Davies, P.D.: Tuberculosis and poverty. Br. Med. J. 307(6907), 759–761 (1993)
Steingart, K.R., et al.: A systematic review of commercial serological antibody detection tests for the diagnosis of extrapulmonary tuberculosis. Postgrad. Med. J. 83(985), 705–712 (2007)
Steingart, K.R., et al.: Fluorescence versus conventional sputum smear microscopy for tuberculosis: a systematic review. Lancet Infect. Dis. 9(6), 570–581 (2006)
Szegedy, C., et al.: Going deeper with convolutions. In: IEEE Computer Society Conference on Computer Vision and Pattern Recognition, pp. 1–9 (2015)
Toman, K.: Toman’s Tuberculosis: Case Detection, Treatment and Monitoring: Questions and Answers. World Health Organization (2004)
Valsson, S., Arandjelović, O.: Nuances of interpreting X-ray analysis by deep learning and lessons for reporting experimental findings. Sci 4(1), 1–13 (2022)
Vente, D., Arandjelović, O., Baron, V., Dombay, E., Gillespie, S.: Using machine learning for automatic counting of lipid-rich tuberculosis cells in fluorescence microscopy images. In: Proceedings of AAAI Conference on Artificial Intelligence Workshop on Health Intelligence, pp. 57–68 (2019)
Veropoulos, K., Learmonth, G., Campbell, C., Knight, B., Simpson, J.: Automated identification of tubercle bacilli in sputum: a preliminary investigation. Anal. Quant. Cytol. Histol. 21(4), 277–282 (1999)
World Health Organisation: Global Tuberculosis Report. Technical report (2018). https://apps.who.int/iris/bitstream/handle/10665/274453/9789241565646-eng.pdf
Zachariou, M., Arandjelović, O., Sloan, S., Sabiiti, W., Mtafya, B.: Tuberculosis bacteria detection and counting in fluorescence microscopy images using a multi-stage deep learning pipeline. Information 13(2), 96 (2022)
Zagoruyko, S., Komodakis, N.: Wide residual networks. arXiv preprint arXiv:1605.07146 (2016)
Zhai, Y., Liu, Y., Zhou, D., Liu, S.: Automatic identification of mycobacterium tuberculosis from ZN-stained sputum smear: algorithm and system design. In: IEEE International Conference on Robotics and Biomimetics, pp. 41–46 (2010)
Acknowledgements
We will like to express our appreciation to the McKenzie Institute for providing the necessary funding to complete this work.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 Springer Nature Switzerland AG
About this paper
Cite this paper
Zachariou, M., Arandjelović, O., Dombay, E., Sabiiti, W., Mtafya, B., Sloan, D. (2022). Extracting and Classifying Salient Fields of View from Microscopy Slides of Tuberculosis Bacteria. In: El Yacoubi, M., Granger, E., Yuen, P.C., Pal, U., Vincent, N. (eds) Pattern Recognition and Artificial Intelligence. ICPRAI 2022. Lecture Notes in Computer Science, vol 13363. Springer, Cham. https://doi.org/10.1007/978-3-031-09037-0_13
Download citation
DOI: https://doi.org/10.1007/978-3-031-09037-0_13
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-09036-3
Online ISBN: 978-3-031-09037-0
eBook Packages: Computer ScienceComputer Science (R0)