Abstract
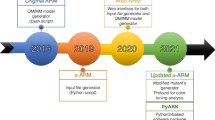
In recent years, photoactive proteins such as rhodopsins have become a common target for cutting-edge research in the field of optogenetics. Alongside wet-lab research, computational methods are also developing rapidly to provide the necessary tools to analyze and rationalize experimental results and, most of all, drive the design of novel systems. The Automatic Rhodopsin Modeling (ARM) protocol is focused on providing exactly the necessary computational tools to study rhodopsins, those being either natural or resulting from mutations. The code has evolved along the years to finally provide results that are reproducible by any user, accurate and reliable so as to replicate experimental trends. Furthermore, the code is efficient in terms of necessary computing resources and time, and scalable in terms of both number of concurrent calculations as well as features. In this review, we will show how the code underlying ARM achieved each of these properties.
Chapter PDF
Similar content being viewed by others
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2022 The Author(s)
About this chapter
Cite this chapter
González, L.P., Barneschi, L., Padula, D., Vico, L.D., Olivucci, M. (2022). Evolution of the Automatic Rhodopsin Modeling (ARM) Protocol. In: Filatov, M., Choi, C.H., Olivucci, M. (eds) New Horizons in Computational Chemistry Software. Topics in Current Chemistry Collections. Springer, Cham. https://doi.org/10.1007/978-3-031-07658-9_5
Download citation
DOI: https://doi.org/10.1007/978-3-031-07658-9_5
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-07657-2
Online ISBN: 978-3-031-07658-9
eBook Packages: Chemistry and Materials ScienceChemistry and Material Science (R0)